
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,542,646 – 18,542,791 |
| Length | 145 |
| Max. P | 0.846852 |

| Location | 18,542,646 – 18,542,761 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.79 |
| Mean single sequence MFE | -33.08 |
| Consensus MFE | -23.12 |
| Energy contribution | -23.45 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.779264 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18542646 115 + 20766785 UCGCAGU--GCAUAUUUC---AAUGUUGUUGGAGGCGACGCUGGCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUCG ...((((--(....((((---(((...)))))))....)))))((.((((((((((((.........))))))))))))))((....)).((((((....)))))).............. ( -30.80) >DroPse_CAF1 25343 116 + 1 -UGUAGUCGGCGUUGUAC---CUCGUAGUUGGAGGCGACGCUCUCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGCAAACCACAGUUUACAUUCACUCAAUUCG -(((((.((((((((..(---(((.......)))))))))))....((((((((((((.........))))))))))))((((....).)))........).)))))............. ( -34.30) >DroGri_CAF1 24370 111 + 1 ------UGGGAGCUGCAG---UUUGAAGUUGGAGGCGACGCUGCCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUUG ------(((((((.((((---(.....((((....)))))))))..((((((((((((.........)))))))))))))))(....)..((((((....)))))).....))))..... ( -32.20) >DroWil_CAF1 32661 114 + 1 UUGUAGU--G----UCUCGGAAUUGUAGUUGGAGGCGAUGCUGGCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGAUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUCG .......--.----.....((((((.(((.((((((.......)))((((((((((((.........))))))))))))..((....)).((((((....)))))).)))))))))))). ( -33.10) >DroMoj_CAF1 29516 108 + 1 ------UGGGA-CCUGGG-----AGCAGUUGCAGGCGACGCUGCCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUUG ------((((.-...(((-----(((.((((....))))))).)))((((((((((((.........))))))))))))..((....)).((((((....)))))).....))))..... ( -35.20) >DroAna_CAF1 20710 103 + 1 --------------UCUC---AGUGCUGGCAGAGGCGACGCUGCCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUCG --------------((.(---(((...(((((.(....).))))).((((((((((((.........)))))))))))))))))).....((((((....)))))).............. ( -32.90) >consensus ______U__G____UCUC___AUUGUAGUUGGAGGCGACGCUGCCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUCG ...............................(((..((.......(((((((((((((.........))))))))))))).((....)).((((((....))))))..)).)))...... (-23.12 = -23.45 + 0.33)
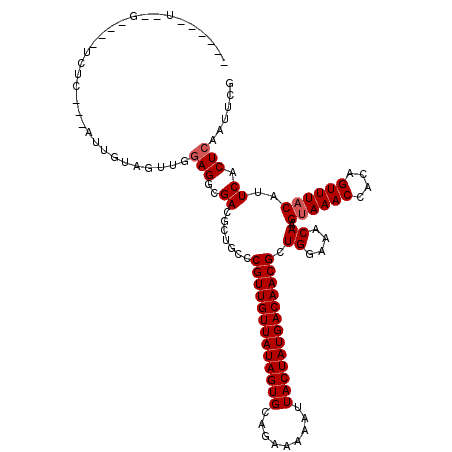


| Location | 18,542,681 – 18,542,791 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.95 |
| Mean single sequence MFE | -35.95 |
| Consensus MFE | -21.00 |
| Energy contribution | -21.50 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.89 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.846852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18542681 110 + 20766785 CUGGCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUCGGUUC--CGGCGAG----GG----UGGCGGGCUACGGCGGC ...(((((((((((((((.........))))))))))))((((....)).((((((....))))))...(((((...(((....--)))...)----))----))))..((....))))) ( -37.20) >DroVir_CAF1 26238 104 + 1 CUGCCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUUGGUGC---GACAAG------------GCAGGC-GAGGAGCA (((((.((((((((((((.........))))))))))))((((....)).((((((....))))))................))---.....)------------))))((-.....)). ( -32.60) >DroPse_CAF1 25379 114 + 1 CUCUCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGCAAACCACAGUUUACAUUCACUCAAUUCGGUGC--UGGAGAAAGACGG----CGACCGGGUGACUCGGA ...(((((((((((((((.........))))))))))))((((....).)))................(((((.....((((((--((........)))----).)))))))))...))) ( -35.50) >DroWil_CAF1 32695 118 + 1 CUGGCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGAUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUCGGUUGGAUGGCCAG--AGGAGGAGCAGCUGAAUGGUGUGGA .....(((((((((((((.........))))))))))))).((....)).((((((....)))))).(((((.(.(((((((((.....((..--....))..))))))))).).))))) ( -38.30) >DroMoj_CAF1 29544 105 + 1 CUGCCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUUGGUGC---GACAAG------------GCAGGCAGCGGGGCA ..((((((((((((((((.........))))))))))))((((.......((((((....))))))...............(((---......------------)))..)))).)))). ( -35.50) >DroPer_CAF1 24371 114 + 1 CUCUCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUCGGUGC--UGGAGAAAGACGG----CGACCGGGUGACUCGGA ...(((((((((((((((.........))))))))))))..((....)).((((((....))))))..(((((.....((((((--((........)))----).)))))))))...))) ( -36.60) >consensus CUGCCCGUUGUUAUAGUGCAGAAAAAUUACUAUGACAACGCUGGAAACAAGUAAACCACAGUUUACAUUCACUCAAUUCGGUGC__UGGCGAG____GG____CGGCAGGCUGAGGCGGA .....(((((((((((((.........))))))))))))).((....)).((((((....))))))...((((......))))..................................... (-21.00 = -21.50 + 0.50)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:47 2006