| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,514,398 – 18,514,488 |
| Length | 90 |
| Max. P | 0.547980 |

| Location | 18,514,398 – 18,514,488 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 86.01 |
| Mean single sequence MFE | -36.82 |
| Consensus MFE | -24.72 |
| Energy contribution | -25.36 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.547980 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
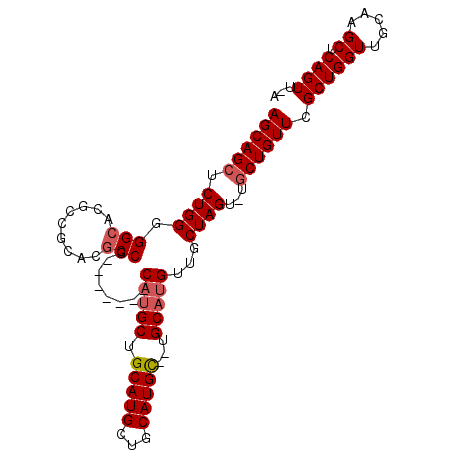
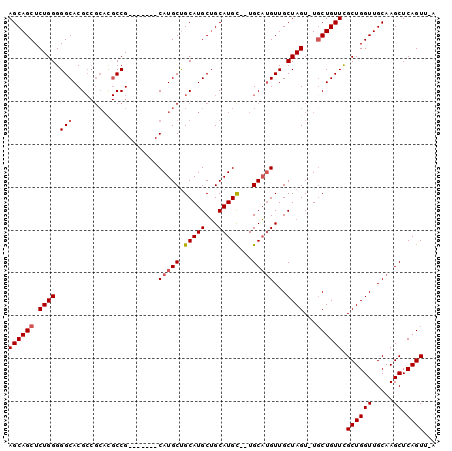
>2R_DroMel_CAF1 18514398 90 + 20766785 AGCAGCUCUGGGGGCACGCCGCACGCCG-------CAUGCUGCAUGCUGCAUGC--UGCAUGUUGCUAGU-UGCUGUUCGCUGGUUGCAAGCUCAGUUCA ((((((.((((.((((.((.(((.((.(-------((((...))))).)).)))--.)).)))).)))).-.)))))).((((((.....)).))))... ( -36.30) >DroSec_CAF1 13400 90 + 1 AGCAGCUCUGGGGGCACGCCGCACGCCG-------CAUGCUGCAUGCUGCAUGC--UGCAUGUUGCUAGU-UGCUGUUCGCUGGUUGCAAGCUCAGUUUA ((((((.((((.((((.((.(((.((.(-------((((...))))).)).)))--.)).)))).)))).-.)))))).((((((.....)).))))... ( -36.30) >DroSim_CAF1 11633 83 + 1 AGCAGCUCUGGGGGCACG-------CCG-------CAUGCUGCAUGCUGCAUGC--UGCAUGUUGCUAGU-UGCUGUUCGCUGGUUGCAAGCUCAGUUCA ((((((.((((.((((.(-------(.(-------(((((........))))))--.)).)))).)))).-.)))))).((((((.....)).))))... ( -34.40) >DroEre_CAF1 16587 96 + 1 AGCAGCUCUGGGGGCACGCCGCACGCCGCAUGCCGCAUGCUGCAUGCAGCAUGUUUUGCUAGUUGCUAGU-UGCUGUUUGCUGGUUGCAAGCUCAGU--- (((((((((((.((((.((.(....).)).))))((((((((....))))))))....)))).....)))-))))((((((.....)))))).....--- ( -39.00) >DroYak_CAF1 19188 95 + 1 AGCAGCUCUGGGGGCACGCCGCAUGCCGCACGCCGCAUGCCGCAUGCUGCAUGC--UGCAUGUUGCUAGUUUUCUGUUCGCUGGUUGCAAGCUCAGU--- .(((((....(.((((.((.((((((.(....).)))))).)).)))).)..))--))).(((.((((((.........)))))).)))........--- ( -38.10) >consensus AGCAGCUCUGGGGGCACGCCGCACGCCG_______CAUGCUGCAUGCUGCAUGC__UGCAUGUUGCUAGU_UGCUGUUCGCUGGUUGCAAGCUCAGUU_A ((((((.((((.(((.........)))........(((((.(((((...)))))...)))))...))))...)))))).((((((.....)).))))... (-24.72 = -25.36 + 0.64)

| Location | 18,514,398 – 18,514,488 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 86.01 |
| Mean single sequence MFE | -32.54 |
| Consensus MFE | -22.68 |
| Energy contribution | -23.28 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.542149 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
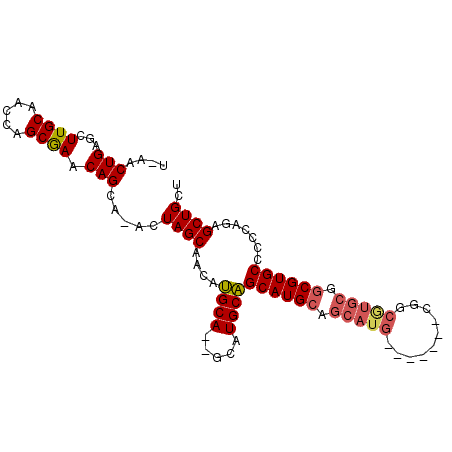
>2R_DroMel_CAF1 18514398 90 - 20766785 UGAACUGAGCUUGCAACCAGCGAACAGCA-ACUAGCAACAUGCA--GCAUGCAGCAUGCAGCAUG-------CGGCGUGCGGCGUGCCCCCAGAGCUGCU .......(((((((.....)).....(((-....((..(((((.--((((((........)))))-------).)))))..)).))).....)))))... ( -31.10) >DroSec_CAF1 13400 90 - 1 UAAACUGAGCUUGCAACCAGCGAACAGCA-ACUAGCAACAUGCA--GCAUGCAGCAUGCAGCAUG-------CGGCGUGCGGCGUGCCCCCAGAGCUGCU .......(((((((.....)).....(((-....((..(((((.--((((((........)))))-------).)))))..)).))).....)))))... ( -31.10) >DroSim_CAF1 11633 83 - 1 UGAACUGAGCUUGCAACCAGCGAACAGCA-ACUAGCAACAUGCA--GCAUGCAGCAUGCAGCAUG-------CGG-------CGUGCCCCCAGAGCUGCU .......(((((((.....)).....(((-....((..(((((.--(((((...))))).)))))-------..)-------).))).....)))))... ( -25.80) >DroEre_CAF1 16587 96 - 1 ---ACUGAGCUUGCAACCAGCAAACAGCA-ACUAGCAACUAGCAAAACAUGCUGCAUGCAGCAUGCGGCAUGCGGCGUGCGGCGUGCCCCCAGAGCUGCU ---.......((((.....)))).((((.-....((.....))....((((((((((((.(((((...))))).))))))))))))........)))).. ( -36.70) >DroYak_CAF1 19188 95 - 1 ---ACUGAGCUUGCAACCAGCGAACAGAAAACUAGCAACAUGCA--GCAUGCAGCAUGCGGCAUGCGGCGUGCGGCAUGCGGCGUGCCCCCAGAGCUGCU ---.(((...((((.....)))).)))..............(((--((.((..((((((.((((((.(....).)))))).))))))...))..))))). ( -38.00) >consensus U_AACUGAGCUUGCAACCAGCGAACAGCA_ACUAGCAACAUGCA__GCAUGCAGCAUGCAGCAUG_______CGGCGUGCGGCGUGCCCCCAGAGCUGCU ....(((...((((.....)))).))).....((((....((((.....))))((((((.(((((..........))))).)))))).......)))).. (-22.68 = -23.28 + 0.60)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:29 2006