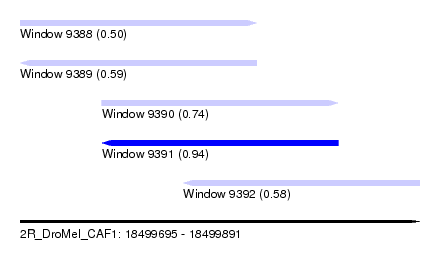

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,499,695 – 18,499,891 |
| Length | 196 |
| Max. P | 0.938959 |

| Location | 18,499,695 – 18,499,811 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 93.16 |
| Mean single sequence MFE | -25.81 |
| Consensus MFE | -21.28 |
| Energy contribution | -22.03 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.82 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18499695 116 + 20766785 GAUUAAUUAAGAACUAAAAAUGAAAUGGCAUAAAACCAAUCAAAUGGCCAAAUGGCCAAACGGCUUUCACCGCAAUUGCGAA-AACGUUCGCUCCAGAGCCGUUUGCUCAAGCUGCA ....................(((..(((.......))).)))..(((((....)))))..((((((...........(((((-....)))))....((((.....)))))))))).. ( -26.80) >DroSec_CAF1 18491 109 + 1 GAUUAAUUAAGAACUAAAAAUUAAAUGGCAUAAAACCAAUCAAAUG--------GCCAAACGGCUUUCACCGCAAUUGCGAAAAACGUUCGCUCCAGAGCCGUUUGCUCAAGCUGCA ..........................(((......(((......))--------).(((((((((((..........(((((.....)))))...))))))))))).....)))... ( -22.32) >DroSim_CAF1 7529 117 + 1 GAUUAAUUAAGAACUAAAAAUGAAAUGGCAUAAAACCAAUCAAAUGGCCAAAUGGCCAAACGGCUUUCACCGCAAUUGCGAAAAACGUUCGCUACAGUGCCGUUUGCUCAAGCUGCA ......................(((((((((.............(((((....)))))..(((......))).....(((((.....)))))....)))))))))((.......)). ( -28.30) >consensus GAUUAAUUAAGAACUAAAAAUGAAAUGGCAUAAAACCAAUCAAAUGGCCAAAUGGCCAAACGGCUUUCACCGCAAUUGCGAAAAACGUUCGCUCCAGAGCCGUUUGCUCAAGCUGCA ......................(((((((...............(((((....)))))..(((......))).....(((((.....)))))......)))))))((.......)). (-21.28 = -22.03 + 0.75)

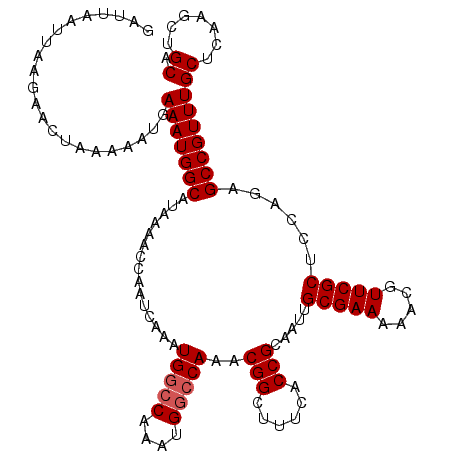
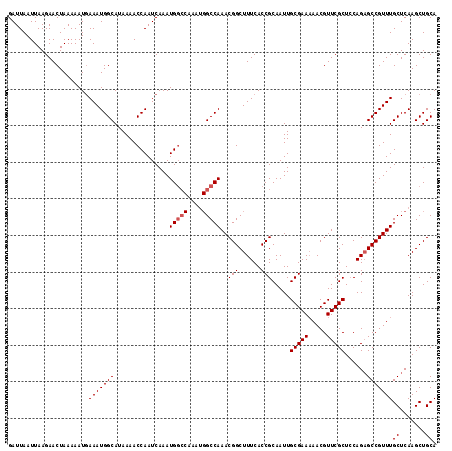
| Location | 18,499,695 – 18,499,811 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 93.16 |
| Mean single sequence MFE | -33.23 |
| Consensus MFE | -26.62 |
| Energy contribution | -27.70 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.589751 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18499695 116 - 20766785 UGCAGCUUGAGCAAACGGCUCUGGAGCGAACGUU-UUCGCAAUUGCGGUGAAAGCCGUUUGGCCAUUUGGCCAUUUGAUUGGUUUUAUGCCAUUUCAUUUUUAGUUCUUAAUUAAUC ....((((((((.....))))..))))(((((((-(((((.......))))))))((..(((((....)))))..))..((((.....))))...........)))).......... ( -34.70) >DroSec_CAF1 18491 109 - 1 UGCAGCUUGAGCAAACGGCUCUGGAGCGAACGUUUUUCGCAAUUGCGGUGAAAGCCGUUUGGC--------CAUUUGAUUGGUUUUAUGCCAUUUAAUUUUUAGUUCUUAAUUAAUC ....((((((((.....))))..))))((((.......((....(((((....)))))...))--------...((((.((((.....)))).))))......)))).......... ( -29.60) >DroSim_CAF1 7529 117 - 1 UGCAGCUUGAGCAAACGGCACUGUAGCGAACGUUUUUCGCAAUUGCGGUGAAAGCCGUUUGGCCAUUUGGCCAUUUGAUUGGUUUUAUGCCAUUUCAUUUUUAGUUCUUAAUUAAUC ........((((((((((((((((((((((.....)))))...))))))....)))))))((((....))))...(((.((((.....))))..)))......)))).......... ( -35.40) >consensus UGCAGCUUGAGCAAACGGCUCUGGAGCGAACGUUUUUCGCAAUUGCGGUGAAAGCCGUUUGGCCAUUUGGCCAUUUGAUUGGUUUUAUGCCAUUUCAUUUUUAGUUCUUAAUUAAUC ....(((.((((.....))))...)))((((.......((....))(((((((((((..(((((....)))))......))))))).))))............)))).......... (-26.62 = -27.70 + 1.08)

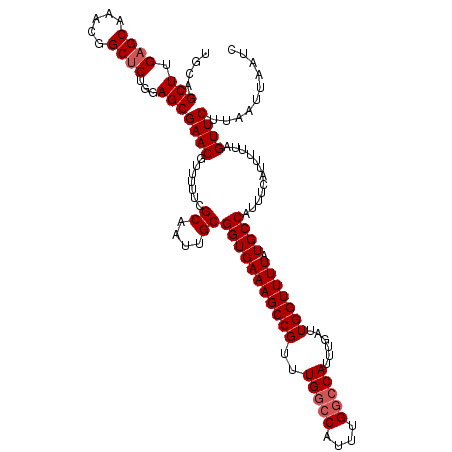
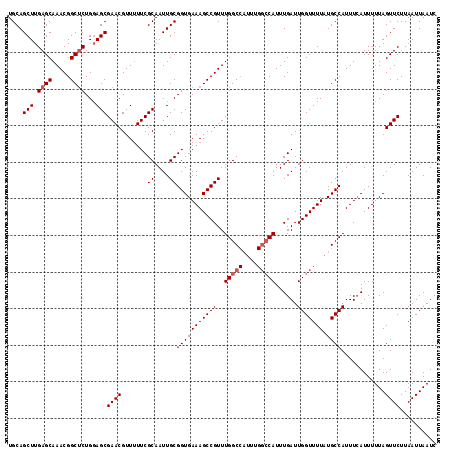
| Location | 18,499,735 – 18,499,851 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 90.78 |
| Mean single sequence MFE | -33.02 |
| Consensus MFE | -27.37 |
| Energy contribution | -28.79 |
| Covariance contribution | 1.42 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.736018 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18499735 116 + 20766785 CAAAUGGCCAAAUGGCCAAACGGCUUUCACCGCAAUUGCGAA-AACGUUCGCUCCAGAGCCGUUUGCUCAAGCUGCAAAAUAUGACUUCAAAUUAAUGCAAAGUGAGGUAGGUGCUC .....((((....))))((((((((((..........(((((-....)))))...)))))))))).((((...((((.(((.((....)).)))..))))...)))).......... ( -31.92) >DroSec_CAF1 18531 105 + 1 CAAAUG--------GCCAAACGGCUUUCACCGCAAUUGCGAAAAACGUUCGCUCCAGAGCCGUUUGCUCAAGCUGCAAAAUAUGACUUCAAAUUAAUGCAGAGUGAG----GUGCUC .....(--------(((((((((((((..........(((((.....)))))...)))))))))))((((..(((((.(((.((....)).)))..)))))..))))----..))). ( -32.72) >DroSim_CAF1 7569 113 + 1 CAAAUGGCCAAAUGGCCAAACGGCUUUCACCGCAAUUGCGAAAAACGUUCGCUACAGUGCCGUUUGCUCAAGCUGCAAAAUAUGACUUCAAAUUAAUGCAGAGUGAG----GUGCUC .....((((....))))(((((((.......((....(((((.....)))))....))))))))).((((..(((((.(((.((....)).)))..)))))..))))----...... ( -34.41) >consensus CAAAUGGCCAAAUGGCCAAACGGCUUUCACCGCAAUUGCGAAAAACGUUCGCUCCAGAGCCGUUUGCUCAAGCUGCAAAAUAUGACUUCAAAUUAAUGCAGAGUGAG____GUGCUC .....((((....))))((((((((((..........(((((.....)))))...)))))))))).((((..(((((.(((.((....)).)))..)))))..)))).......... (-27.37 = -28.79 + 1.42)

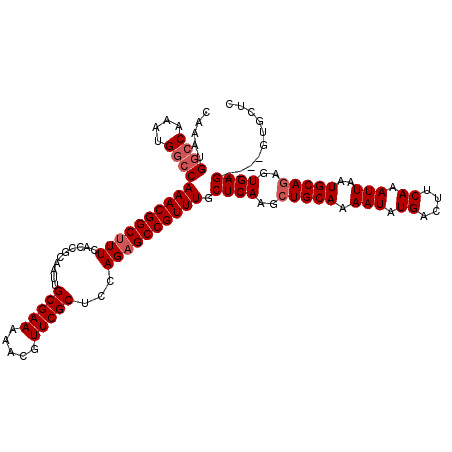
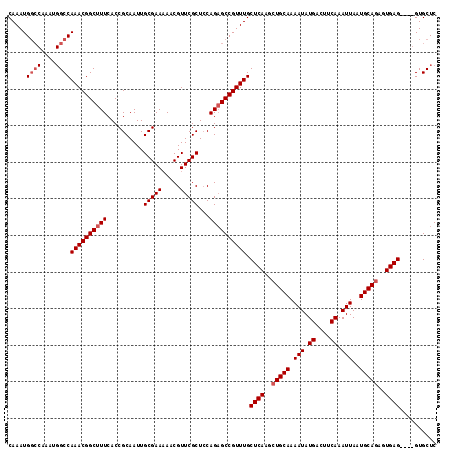
| Location | 18,499,735 – 18,499,851 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 90.78 |
| Mean single sequence MFE | -36.63 |
| Consensus MFE | -33.44 |
| Energy contribution | -34.30 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938959 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
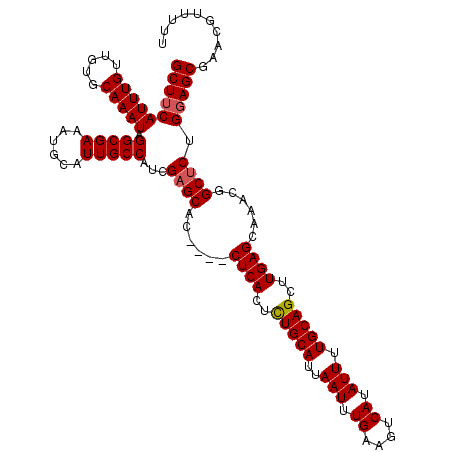
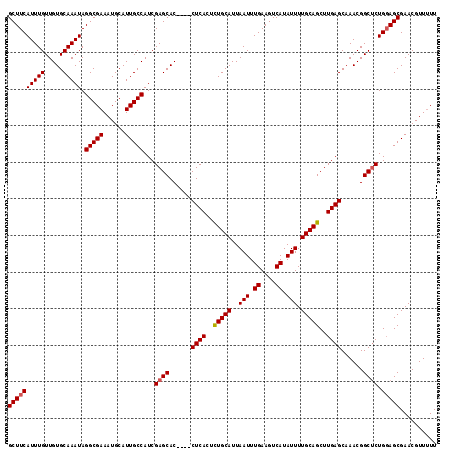
>2R_DroMel_CAF1 18499735 116 - 20766785 GAGCACCUACCUCACUUUGCAUUAAUUUGAAGUCAUAUUUUGCAGCUUGAGCAAACGGCUCUGGAGCGAACGUU-UUCGCAAUUGCGGUGAAAGCCGUUUGGCCAUUUGGCCAUUUG ((((......((((..(((((..(((.((....)).))).)))))..))))......))))....(((((....-)))))....(((((....))))).(((((....))))).... ( -38.40) >DroSec_CAF1 18531 105 - 1 GAGCAC----CUCACUCUGCAUUAAUUUGAAGUCAUAUUUUGCAGCUUGAGCAAACGGCUCUGGAGCGAACGUUUUUCGCAAUUGCGGUGAAAGCCGUUUGGC--------CAUUUG ((((..----((((..(((((..(((.((....)).))).)))))..))))......))))(((..((((((.(((((((.......))))))).)))))).)--------)).... ( -33.10) >DroSim_CAF1 7569 113 - 1 GAGCAC----CUCACUCUGCAUUAAUUUGAAGUCAUAUUUUGCAGCUUGAGCAAACGGCACUGUAGCGAACGUUUUUCGCAAUUGCGGUGAAAGCCGUUUGGCCAUUUGGCCAUUUG ......----((((..(((((..(((.((....)).))).)))))..)))).((((((((((((((((((.....)))))...))))))....)))))))((((....))))..... ( -38.40) >consensus GAGCAC____CUCACUCUGCAUUAAUUUGAAGUCAUAUUUUGCAGCUUGAGCAAACGGCUCUGGAGCGAACGUUUUUCGCAAUUGCGGUGAAAGCCGUUUGGCCAUUUGGCCAUUUG ((((......((((..(((((..(((.((....)).))).)))))..))))......))))....(((((.....)))))....(((((....))))).(((((....))))).... (-33.44 = -34.30 + 0.86)

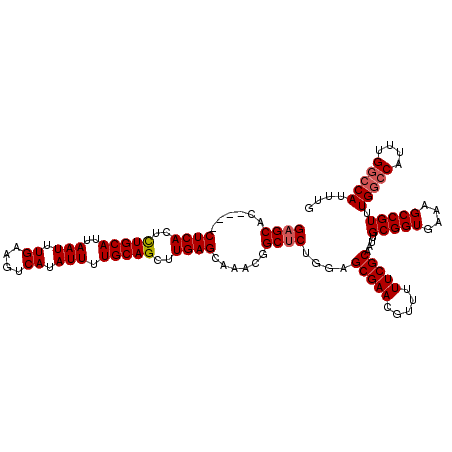
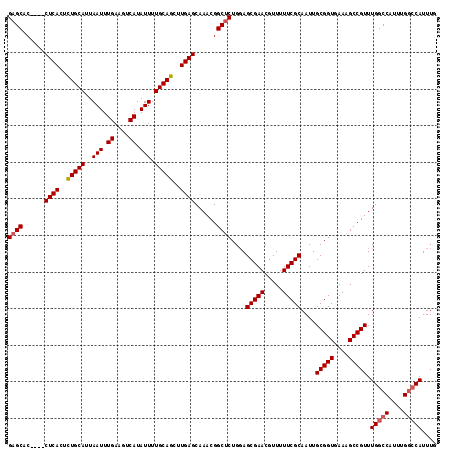
| Location | 18,499,775 – 18,499,891 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 95.39 |
| Mean single sequence MFE | -30.63 |
| Consensus MFE | -30.32 |
| Energy contribution | -30.77 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.583173 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18499775 116 - 20766785 GCUUCAUUUGUUGUGCAAAUAGGCGAAAUGCAUUGCCAUCGAGCACCUACCUCACUUUGCAUUAAUUUGAAGUCAUAUUUUGCAGCUUGAGCAAACGGCUCUGGAGCGAACGUU-UU ((((((((((.....))))).(((((......)))))...((((......((((..(((((..(((.((....)).))).)))))..))))......)))).))))).......-.. ( -31.80) >DroSec_CAF1 18563 113 - 1 GCUUCAUUUGUUGUGCAAAUAGGCGAAAUGCAUUGCCAUCGAGCAC----CUCACUCUGCAUUAAUUUGAAGUCAUAUUUUGCAGCUUGAGCAAACGGCUCUGGAGCGAACGUUUUU ((((((((((.....))))).(((((......)))))...((((..----((((..(((((..(((.((....)).))).)))))..))))......)))).))))).......... ( -32.30) >DroSim_CAF1 7609 113 - 1 GCUUCAUUUGUUGUGCAAAUAGGCGAAAUGCAUUGCCAUCGAGCAC----CUCACUCUGCAUUAAUUUGAAGUCAUAUUUUGCAGCUUGAGCAAACGGCACUGUAGCGAACGUUUUU (((.((..((((((((.....(((((......))))).....))..----((((..(((((..(((.((....)).))).)))))..))))...)))))).)).))).......... ( -27.80) >consensus GCUUCAUUUGUUGUGCAAAUAGGCGAAAUGCAUUGCCAUCGAGCAC____CUCACUCUGCAUUAAUUUGAAGUCAUAUUUUGCAGCUUGAGCAAACGGCUCUGGAGCGAACGUUUUU ((((((((((.....))))).(((((......)))))...((((......((((..(((((..(((.((....)).))).)))))..))))......)))).))))).......... (-30.32 = -30.77 + 0.45)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:21 2006