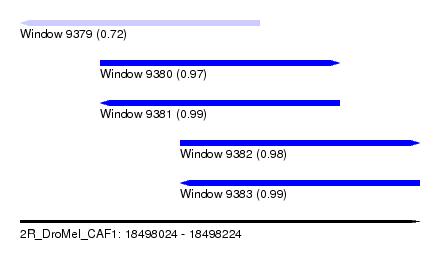

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,498,024 – 18,498,224 |
| Length | 200 |
| Max. P | 0.987430 |

| Location | 18,498,024 – 18,498,144 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 99.44 |
| Mean single sequence MFE | -44.13 |
| Consensus MFE | -43.30 |
| Energy contribution | -43.30 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724111 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
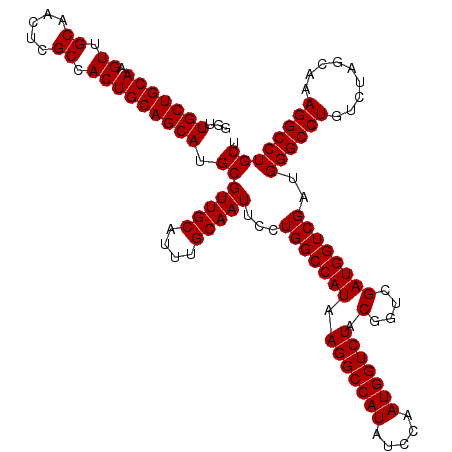
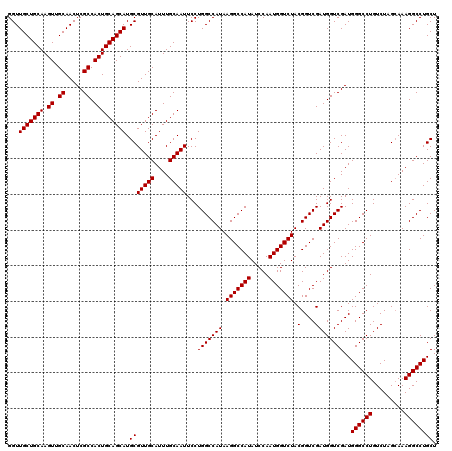
>2R_DroMel_CAF1 18498024 120 - 20766785 GGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUGCAUUUGCAAUUCCUGGCCAUAAGGCCAUAUCCAAUGGUCUACGGUCGAUGGUCGAUGGGCCUGUCUAGGAAAGGCCUGCU .(((((((((.((.((.....)).))))))))).))...(((...((..(((((((((...(((((((.....)))))))..))))(((((((....)))).))).)))))..)).))). ( -44.80) >DroSec_CAF1 16683 120 - 1 GGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUGCAUUUGCAAUUCCUGGCCAUAAGGCCAUAUCCAAUGGUCUACGGUCGAUGGUCGAUGGGCCUGUCUAGCAAAGGCCUGCU ((((((((((.((.((.....)).)))))))).((((((((....)))))....((((...(((((((.....)))))))...((((.....)))).))))......)))..)))).... ( -43.80) >DroSim_CAF1 5820 120 - 1 GGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUGCAUUUGCAAUUCCUGGCCAUAAGGCCAUAUCCAAUGGUCUACGGUCGAUGGUCGAUGGGCCUGUCUAGCAAAGGCCUGCU ((((((((((.((.((.....)).)))))))).((((((((....)))))....((((...(((((((.....)))))))...((((.....)))).))))......)))..)))).... ( -43.80) >consensus GGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUGCAUUUGCAAUUCCUGGCCAUAAGGCCAUAUCCAAUGGUCUACGGUCGAUGGUCGAUGGGCCUGUCUAGCAAAGGCCUGCU ...(((((((.((.((.....)).))))))))).(((((((....)))))...(((((((.(((((((.....))))))).(....))))))))..((((((.........)))))))). (-43.30 = -43.30 + 0.00)

| Location | 18,498,064 – 18,498,184 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 100.00 |
| Mean single sequence MFE | -38.60 |
| Consensus MFE | -38.60 |
| Energy contribution | -38.60 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.44 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966781 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
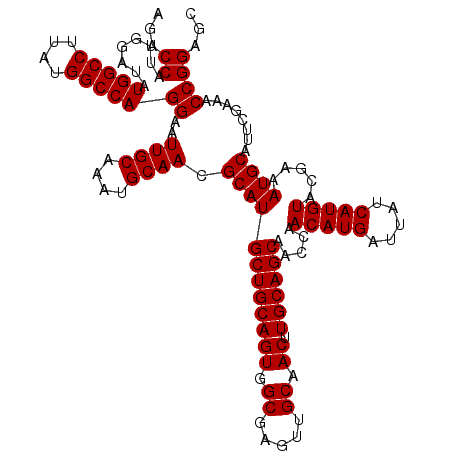
>2R_DroMel_CAF1 18498064 120 + 20766785 AGACCAUUGGAUAUGGCCUUAUGGCCAGGAAUUGCAAAUGCAACGCAUGCUGCAGUGGCGAGUUGCAACUUGCAGCAACCAACAUGAUUAUCAUGUACGAAAUGCAUUCGAAACCGGAGC ...((........(((((....)))))((..((((....)))).((((((((((((.((.....)).)).)))))).....(((((.....))))).....))))........))))... ( -38.60) >DroSec_CAF1 16723 120 + 1 AGACCAUUGGAUAUGGCCUUAUGGCCAGGAAUUGCAAAUGCAACGCAUGCUGCAGUGGCGAGUUGCAACUUGCAGCAACCAACAUGAUUAUCAUGUACGAAAUGCAUUCGAAACCGGAGC ...((........(((((....)))))((..((((....)))).((((((((((((.((.....)).)).)))))).....(((((.....))))).....))))........))))... ( -38.60) >DroSim_CAF1 5860 120 + 1 AGACCAUUGGAUAUGGCCUUAUGGCCAGGAAUUGCAAAUGCAACGCAUGCUGCAGUGGCGAGUUGCAACUUGCAGCAACCAACAUGAUUAUCAUGUACGAAAUGCAUUCGAAACCGGAGC ...((........(((((....)))))((..((((....)))).((((((((((((.((.....)).)).)))))).....(((((.....))))).....))))........))))... ( -38.60) >consensus AGACCAUUGGAUAUGGCCUUAUGGCCAGGAAUUGCAAAUGCAACGCAUGCUGCAGUGGCGAGUUGCAACUUGCAGCAACCAACAUGAUUAUCAUGUACGAAAUGCAUUCGAAACCGGAGC ...((........(((((....)))))((..((((....)))).((((((((((((.((.....)).)).)))))).....(((((.....))))).....))))........))))... (-38.60 = -38.60 + 0.00)


| Location | 18,498,064 – 18,498,184 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 100.00 |
| Mean single sequence MFE | -40.60 |
| Consensus MFE | -40.60 |
| Energy contribution | -40.60 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.01 |
| Structure conservation index | 1.00 |
| SVM decision value | 2.08 |
| SVM RNA-class probability | 0.987430 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
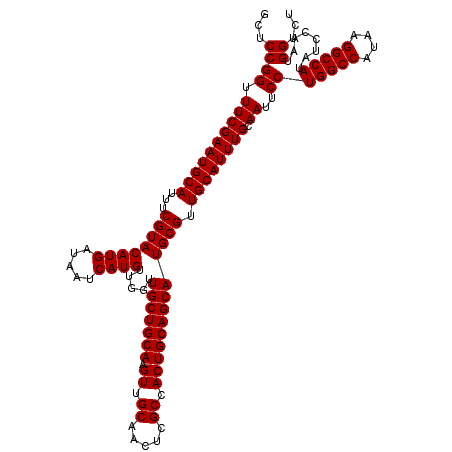
>2R_DroMel_CAF1 18498064 120 - 20766785 GCUCCGGUUUCGAAUGCAUUUCGUACAUGAUAAUCAUGUUGGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUGCAUUUGCAAUUCCUGGCCAUAAGGCCAUAUCCAAUGGUCU ...((((.((((((((((...((((((((.....)))).....(((((((.((.((.....)).))))))))))))).)))))))).))..))(((((....)))))........))... ( -40.60) >DroSec_CAF1 16723 120 - 1 GCUCCGGUUUCGAAUGCAUUUCGUACAUGAUAAUCAUGUUGGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUGCAUUUGCAAUUCCUGGCCAUAAGGCCAUAUCCAAUGGUCU ...((((.((((((((((...((((((((.....)))).....(((((((.((.((.....)).))))))))))))).)))))))).))..))(((((....)))))........))... ( -40.60) >DroSim_CAF1 5860 120 - 1 GCUCCGGUUUCGAAUGCAUUUCGUACAUGAUAAUCAUGUUGGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUGCAUUUGCAAUUCCUGGCCAUAAGGCCAUAUCCAAUGGUCU ...((((.((((((((((...((((((((.....)))).....(((((((.((.((.....)).))))))))))))).)))))))).))..))(((((....)))))........))... ( -40.60) >consensus GCUCCGGUUUCGAAUGCAUUUCGUACAUGAUAAUCAUGUUGGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUGCAUUUGCAAUUCCUGGCCAUAAGGCCAUAUCCAAUGGUCU ...((((.((((((((((...((((((((.....)))).....(((((((.((.((.....)).))))))))))))).)))))))).))..))(((((....)))))........))... (-40.60 = -40.60 + 0.00)

| Location | 18,498,104 – 18,498,224 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.78 |
| Mean single sequence MFE | -42.07 |
| Consensus MFE | -41.60 |
| Energy contribution | -41.60 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.73 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.97 |
| SVM RNA-class probability | 0.984356 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
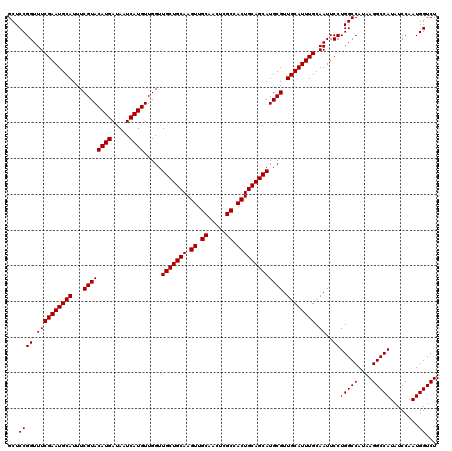
>2R_DroMel_CAF1 18498104 120 + 20766785 CAACGCAUGCUGCAGUGGCGAGUUGCAACUUGCAGCAACCAACAUGAUUAUCAUGUACGAAAUGCAUUCGAAACCGGAGCCAGCUGGCCAACAGCUGACGAAGCUGGACAAAACCGGCCG ....((.(((((((((.((.....)).)).))))))).((.(((((.....))))).((((.....)))).....)).))((((((.....)))))).....(((((......))))).. ( -41.60) >DroSec_CAF1 16763 120 + 1 CAACGCAUGCUGCAGUGGCGAGUUGCAACUUGCAGCAACCAACAUGAUUAUCAUGUACGAAAUGCAUUCGAAACCGGAGCCAGCUGGCCAACAGCUGACGUAGCUGGCUAAAACCGGCCU ....((((((((((((.((.....)).)).)))))).....(((((.....))))).....))))........((((((((((((((.((.....)).).)))))))))....))))... ( -42.30) >DroSim_CAF1 5900 120 + 1 CAACGCAUGCUGCAGUGGCGAGUUGCAACUUGCAGCAACCAACAUGAUUAUCAUGUACGAAAUGCAUUCGAAACCGGAGCCAGCUGGCCAACAGCUGACGUAGCUGGCUAAAACCGGCCU ....((((((((((((.((.....)).)).)))))).....(((((.....))))).....))))........((((((((((((((.((.....)).).)))))))))....))))... ( -42.30) >consensus CAACGCAUGCUGCAGUGGCGAGUUGCAACUUGCAGCAACCAACAUGAUUAUCAUGUACGAAAUGCAUUCGAAACCGGAGCCAGCUGGCCAACAGCUGACGUAGCUGGCUAAAACCGGCCU ....((.(((((((((.((.....)).)).))))))).((.(((((.....))))).((((.....)))).....)).))((((((.....)))))).....(((((......))))).. (-41.60 = -41.60 + 0.00)


| Location | 18,498,104 – 18,498,224 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.78 |
| Mean single sequence MFE | -47.37 |
| Consensus MFE | -46.45 |
| Energy contribution | -46.57 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.02 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.98 |
| SVM decision value | 2.04 |
| SVM RNA-class probability | 0.986447 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
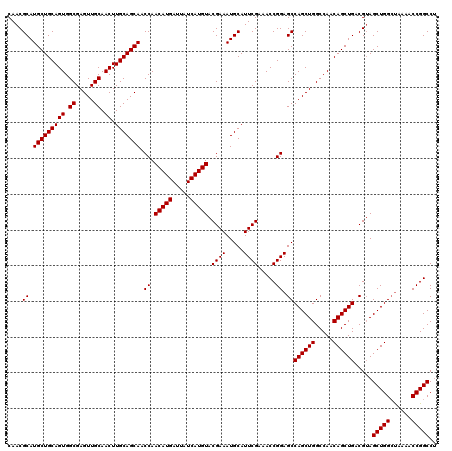
>2R_DroMel_CAF1 18498104 120 - 20766785 CGGCCGGUUUUGUCCAGCUUCGUCAGCUGUUGGCCAGCUGGCUCCGGUUUCGAAUGCAUUUCGUACAUGAUAAUCAUGUUGGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUG .((((((....(.((((((..(((((...))))).))))))).))))))...((((((..(((.(((((.....)))))))).(((((((.((.((.....)).))))))))))))))). ( -44.90) >DroSec_CAF1 16763 120 - 1 AGGCCGGUUUUAGCCAGCUACGUCAGCUGUUGGCCAGCUGGCUCCGGUUUCGAAUGCAUUUCGUACAUGAUAAUCAUGUUGGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUG (((((((....((((((((..(((((...))))).)))))))))))))))..((((((..(((.(((((.....)))))))).(((((((.((.((.....)).))))))))))))))). ( -48.60) >DroSim_CAF1 5900 120 - 1 AGGCCGGUUUUAGCCAGCUACGUCAGCUGUUGGCCAGCUGGCUCCGGUUUCGAAUGCAUUUCGUACAUGAUAAUCAUGUUGGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUG (((((((....((((((((..(((((...))))).)))))))))))))))..((((((..(((.(((((.....)))))))).(((((((.((.((.....)).))))))))))))))). ( -48.60) >consensus AGGCCGGUUUUAGCCAGCUACGUCAGCUGUUGGCCAGCUGGCUCCGGUUUCGAAUGCAUUUCGUACAUGAUAAUCAUGUUGGUUGCUGCAAGUUGCAACUCGCCACUGCAGCAUGCGUUG .((((((....((((((((..(((((...))))).))))))))))))))...((((((..(((.(((((.....)))))))).(((((((.((.((.....)).))))))))))))))). (-46.45 = -46.57 + 0.11)

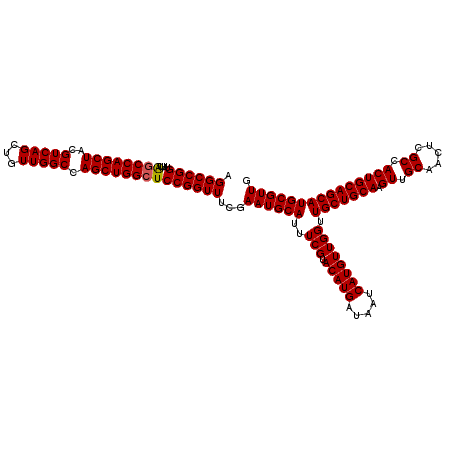

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:13 2006