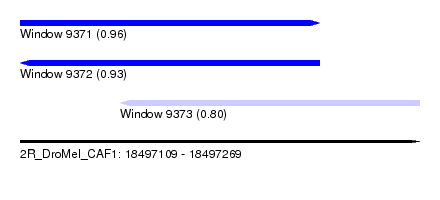

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,497,109 – 18,497,269 |
| Length | 160 |
| Max. P | 0.957612 |

| Location | 18,497,109 – 18,497,229 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.75 |
| Mean single sequence MFE | -39.38 |
| Consensus MFE | -38.51 |
| Energy contribution | -38.33 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.13 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.48 |
| SVM RNA-class probability | 0.957612 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18497109 120 + 20766785 UUAAAAAGCGCCUACACGCCUGACACGCCCCCUCAAGUGUUGGCAGUCGCUGCUGUUGACAAACAAACAGCAAAAGGCCACAAAAGACACAAAGGGGGCUGUGUUCCAACACUUGAGUUC .......(((......)))..((((((((((((...(((((.(..(((..(((((((........)))))))...)))..)....)))))..))))))).)))))............... ( -40.10) >DroSec_CAF1 15739 119 + 1 UUAAAAAGCGCCUACACGCCUGACACGCC-CCUCAAGUGUUGGCAGUCGCUGCUGUUGACAAACAAACAGCAAAAGGCCACAAAAGACACAAAGGGGGCUGCGUUCCAACACUUGAGUUC .......(((......)))..........-.((((((((((((.((.((((((((((........)))))))...((((.(............)..)))))))))))))))))))))... ( -38.80) >DroSim_CAF1 4889 120 + 1 UUAAAAAGCGCCUACACGCCUGACACGCCCCCUCAAGUGUUGGCAGUCGCUGCUGUUGACAAACAAACAGCAAAAGGCCACAAAAGACACAAAGGGGGCUGCGUUCCAACACUUGAGUUC .......(((......)))............((((((((((((.((.((((((((((........)))))))...((((.(............)..)))))))))))))))))))))... ( -38.80) >DroEre_CAF1 21305 120 + 1 UUAAAAAGCGCCUACACGCCUGACACGCCCCCUCAAGUGUUGGCAGCCGCUGCUGUUGACAAACAAACAGCAAAAGGCCACAAAAGACACAAAGGGGGCUGCGUUCCAACACUUGAGUUC .......(((......)))............(((((((((((((((((.((((((((........))))))....(....)............)).))))))....)))))))))))... ( -39.80) >consensus UUAAAAAGCGCCUACACGCCUGACACGCCCCCUCAAGUGUUGGCAGUCGCUGCUGUUGACAAACAAACAGCAAAAGGCCACAAAAGACACAAAGGGGGCUGCGUUCCAACACUUGAGUUC .......(((......)))............((((((((((((.((.((((((((((........)))))))...((((.(............)..)))))))))))))))))))))... (-38.51 = -38.33 + -0.19)

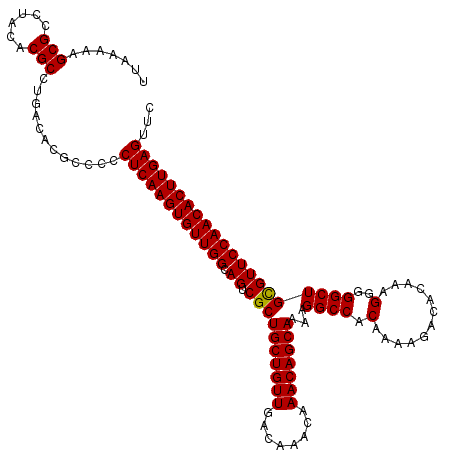
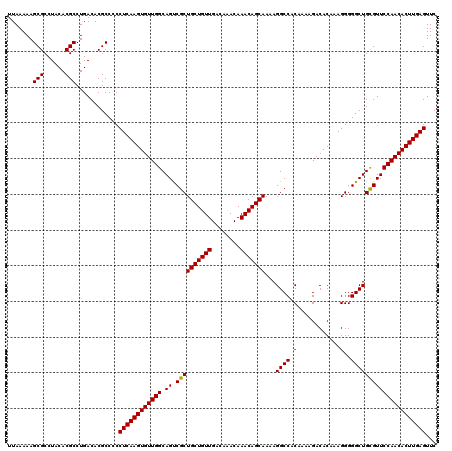
| Location | 18,497,109 – 18,497,229 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.75 |
| Mean single sequence MFE | -46.73 |
| Consensus MFE | -44.09 |
| Energy contribution | -44.40 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.84 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925162 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18497109 120 - 20766785 GAACUCAAGUGUUGGAACACAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGACUGCCAACACUUGAGGGGGCGUGUCAGGCGUGUAGGCGCUUUUUAA ......(((((((...((((.((((((((.((((......((((...((((((((.(((....)))))))))))..))))))))..))))))))((.....)))))).)))))))..... ( -46.00) >DroSec_CAF1 15739 119 - 1 GAACUCAAGUGUUGGAACGCAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGACUGCCAACACUUGAGG-GGCGUGUCAGGCGUGUAGGCGCUUUUUAA ......(((((((...((((.((((((((.((((......((((...((((((((.(((....)))))))))))..))))))))..))))-)).)).....))))...)))))))..... ( -43.60) >DroSim_CAF1 4889 120 - 1 GAACUCAAGUGUUGGAACGCAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGACUGCCAACACUUGAGGGGGCGUGUCAGGCGUGUAGGCGCUUUUUAA ......(((((((...((((.((((((((.((((......((((...((((((((.(((....)))))))))))..))))))))..)))))))).......))))...)))))))..... ( -47.70) >DroEre_CAF1 21305 120 - 1 GAACUCAAGUGUUGGAACGCAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGGCUGCCAACACUUGAGGGGGCGUGUCAGGCGUGUAGGCGCUUUUUAA ......(((((((...((((.((((((((.((((.....(..(((..((((((((........)))))))).)))..)..))))..)))))))).......))))...)))))))..... ( -49.60) >consensus GAACUCAAGUGUUGGAACGCAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGACUGCCAACACUUGAGGGGGCGUGUCAGGCGUGUAGGCGCUUUUUAA ......(((((((...((((.((((((((.((((......((((...((((((((.(((....)))))))))))..))))))))..)))))))).......))))...)))))))..... (-44.09 = -44.40 + 0.31)
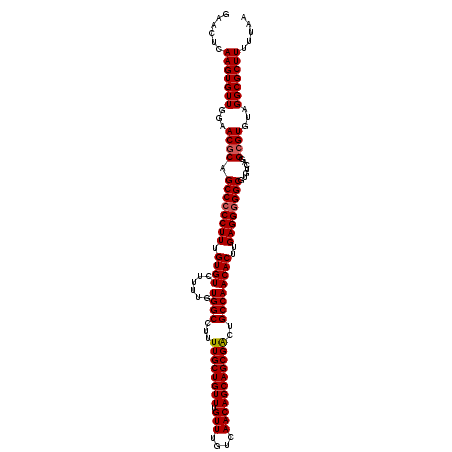


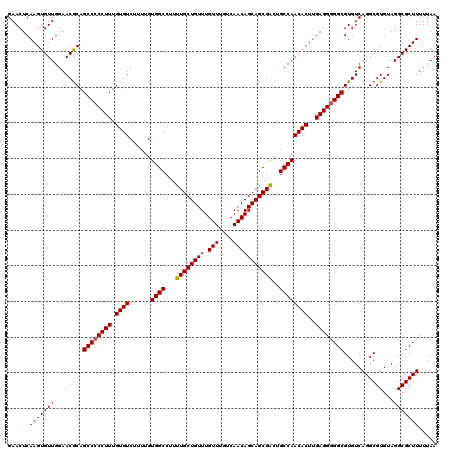
| Location | 18,497,149 – 18,497,269 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.75 |
| Mean single sequence MFE | -41.86 |
| Consensus MFE | -41.71 |
| Energy contribution | -41.59 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.98 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.801616 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18497149 120 - 20766785 GAGCAAAAGAGACGCGACUGUGGCCGACUGCUAAAAAACAGAACUCAAGUGUUGGAACACAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGACUGCCA ..((((..((((((((((((((.(((((.(((...............))))))))..)))))......)))))))))))))(((...((((((((.(((....)))))))))))..))). ( -37.36) >DroSec_CAF1 15778 120 - 1 GAGCAAAAGAGACGCGGCUGUGGCCGACUGCUAAAAAACAGAACUCAAGUGUUGGAACGCAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGACUGCCA ..((((((((.(((.(((((((.(((((.(((...............))))))))..))))))).....))).))))))))(((...((((((((.(((....)))))))))))..))). ( -43.36) >DroSim_CAF1 4929 120 - 1 GAGCAAAAGAGACGCGGCUGUGGCCGACUGCUAAAAAACAGAACUCAAGUGUUGGAACGCAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGACUGCCA ..((((((((.(((.(((((((.(((((.(((...............))))))))..))))))).....))).))))))))(((...((((((((.(((....)))))))))))..))). ( -43.36) >DroEre_CAF1 21345 120 - 1 GAGCAAAAGAGACGCGGCUGUGGCCGACUGCUAAAAAACAGAACUCAAGUGUUGGAACGCAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGGCUGCCA ......((((((((((((((((.(((((.(((...............))))))))..)))))))......)))))))))(..(((..((((((((........)))))))).)))..).. ( -43.36) >consensus GAGCAAAAGAGACGCGGCUGUGGCCGACUGCUAAAAAACAGAACUCAAGUGUUGGAACGCAGCCCCCUUUGUGUCUUUUGUGGCCUUUUGCUGUUUGUUUGUCAACAGCAGCGACUGCCA ..((((((((.(((.(((((((.(((((.(((...............))))))))..))))))).....))).))))))))(((...((((((((.(((....)))))))))))..))). (-41.71 = -41.59 + -0.12)


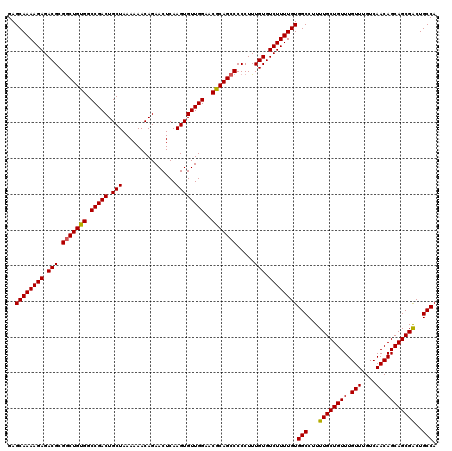
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:04 2006