
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,478,376 – 18,478,474 |
| Length | 98 |
| Max. P | 0.968161 |

| Location | 18,478,376 – 18,478,474 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 93.37 |
| Mean single sequence MFE | -28.52 |
| Consensus MFE | -24.85 |
| Energy contribution | -24.60 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.968161 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
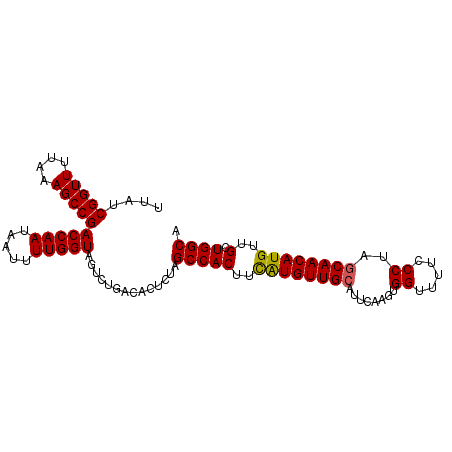
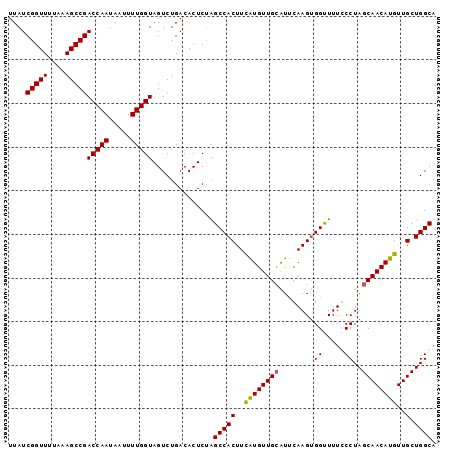
>2R_DroMel_CAF1 18478376 98 + 20766785 UUAUCGGUUUUAAAGCCGACCAAUAAUUUUGGUAGUCUGACACUCUAGCCACUUUAUGUUGCGUGCAAGUGGUUUUGCCUAGCAACAUGUUGCUGGCU ....(((((....)))))(((((.....))))).............((((((..((((((((..(((((....)))))...))))))))..).))))) ( -32.30) >DroSec_CAF1 35412 98 + 1 UUAUCGGUUUUAAAGCCGACCAAUAAUUUUGGUAGUCUGACACUCUAGCCACUUCGUGUUGGAUUCAAGUGGUUUUCCCUAGCAACAUGUUGCUGGCA .....((....((((((((((((.....)))))((((..((((............))))..))))....)))))))))(((((((....))))))).. ( -27.50) >DroSim_CAF1 35711 98 + 1 UUAUCGGUUUUAAAGCCGACCAAUAAUUUUGGUAGUCUGACACUCUAGCCACUUCGUGUUGCAUUCAAGCGGUUUUGCCUAGCAACAUGUUGCUGGCA ....(((((....)))))(((((.....)))))..............(((((..((((((((...((((....))))....))))))))..).)))). ( -26.80) >DroEre_CAF1 35524 98 + 1 UUAUCGGUUUUAUAGCCGACCAAUAAUUUUGGUAGUCUGACACUCCUGCCACUUUAUGUUGCAUUCAAGUGGCUUUCCCUAGCAACAUGUUGCUGGCA ....(((((....)))))(((((.....)))))..............(((((((.(((...)))..))))))).....(((((((....))))))).. ( -27.50) >consensus UUAUCGGUUUUAAAGCCGACCAAUAAUUUUGGUAGUCUGACACUCUAGCCACUUCAUGUUGCAUUCAAGUGGUUUUCCCUAGCAACAUGUUGCUGGCA ....(((((....)))))(((((.....)))))..............(((((..((((((((........((.....))..))))))))..).)))). (-24.85 = -24.60 + -0.25)

| Location | 18,478,376 – 18,478,474 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 93.37 |
| Mean single sequence MFE | -24.32 |
| Consensus MFE | -20.51 |
| Energy contribution | -20.82 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.665061 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
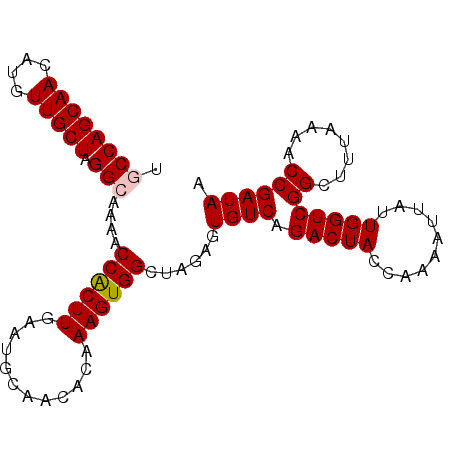
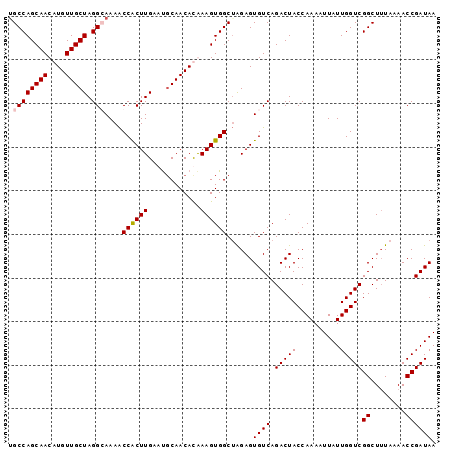
>2R_DroMel_CAF1 18478376 98 - 20766785 AGCCAGCAACAUGUUGCUAGGCAAAACCACUUGCACGCAACAUAAAGUGGCUAGAGUGUCAGACUACCAAAAUUAUUGGUCGGCUUUAAAACCGAUAA (((((.(...(((((((...((((......))))..)))))))...))))))....((((.(((((..........)))))((........)))))). ( -25.70) >DroSec_CAF1 35412 98 - 1 UGCCAGCAACAUGUUGCUAGGGAAAACCACUUGAAUCCAACACGAAGUGGCUAGAGUGUCAGACUACCAAAAUUAUUGGUCGGCUUUAAAACCGAUAA ..(((((((....))))).))......(((((....(((........)))...)))))........((((.....))))((((........))))... ( -19.90) >DroSim_CAF1 35711 98 - 1 UGCCAGCAACAUGUUGCUAGGCAAAACCGCUUGAAUGCAACACGAAGUGGCUAGAGUGUCAGACUACCAAAAUUAUUGGUCGGCUUUAAAACCGAUAA (((((((((....))))).))))....(((((....((.((.....)).))..)))))........((((.....))))((((........))))... ( -24.50) >DroEre_CAF1 35524 98 - 1 UGCCAGCAACAUGUUGCUAGGGAAAGCCACUUGAAUGCAACAUAAAGUGGCAGGAGUGUCAGACUACCAAAAUUAUUGGUCGGCUAUAAAACCGAUAA ..(((((((....))))).))....(((((((..(((...))).))))))).(((((.....))).))..........(((((........))))).. ( -27.20) >consensus UGCCAGCAACAUGUUGCUAGGCAAAACCACUUGAAUGCAACACAAAGUGGCUAGAGUGUCAGACUACCAAAAUUAUUGGUCGGCUUUAAAACCGAUAA .((((((((....))))).)))....((((((............))))))......((((.(((((..........)))))((........)))))). (-20.51 = -20.82 + 0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:55:34 2006