| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,459,711 – 18,459,820 |
| Length | 109 |
| Max. P | 0.993363 |

| Location | 18,459,711 – 18,459,820 |
|---|---|
| Length | 109 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 71.11 |
| Mean single sequence MFE | -22.53 |
| Consensus MFE | -14.27 |
| Energy contribution | -14.77 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.63 |
| SVM decision value | 2.04 |
| SVM RNA-class probability | 0.986521 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18459711 109 + 20766785 GCCUUGACUGGACUAAAUUGAUGCCUUGUCUUUCUUG-UUUUUGAAAGUUGCAUCUGUUGCACAAAUUGGUGC-UGCAGUUGCAGCUAAAUAUUGGUCUAAUUUU---------UUUUUC ........((((((((...............(((...-.....)))(((((((.((((.((((......))))-.)))).))))))).....)))))))).....---------...... ( -29.90) >DroSec_CAF1 21757 85 + 1 G----------------------CCUUGUCUUUCUCG-UUUU-AAAAG-UGCAUCUGUUGCACAAAUUGGUGC-UGCAGUUGCAGCUAAAUAUUGGUCUUAUUUU---------UUUUUU (----------------------((............-....-....(-((((.....)))))..(((...((-(((....)))))..)))...)))........---------...... ( -17.70) >DroSim_CAF1 21958 106 + 1 GCCUUGACUGGAAUAAAUUGAUGCCUUGUCUUUCUCG-UUUU-AAAAGUUGCAUCUGUUGCACAAAUUGGUGC-UGCAGUUGCAGCUAAAUAUUGGUCUUAUUUU-----------UUUU .....(((..(........(((.....))).......-....-...(((((((.((((.((((......))))-.)))).))))))).....)..))).......-----------.... ( -27.00) >DroEre_CAF1 21666 117 + 1 UUUUUGCUUUGACUUAAUUGAUGCCUAUGCUUUCUUG-UUAU-AAAAGUUGCAUCUGUUGCACAAAUUGGUGU-UGCAGUUGCAGCUAAAUAUUGUUAUUAUUUUAACUGCAUAUUUUUU .....((.(..(.....)..).)).(((((.......-....-...(((((((.((((.((((......))))-.)))).))))))).......((((......)))).)))))...... ( -26.60) >DroYak_CAF1 22408 116 + 1 GUUUUGCUUUGACUAAAUUGUUGCCCUUUCUUUCUUG-UUAU-AAAAGUUGCAUCUGUUGCACAAAUUGGUGU-UGCAGUUCCAGCUAAAUAUUGUUAUUAUUUUAACUG-AUUUUUUUU .....((...(((......))))).............-....-...(((((...((((.((((......))))-.))))...)))))((((...((((......))))..-))))..... ( -16.50) >DroAna_CAF1 19836 105 + 1 UGUUUGA---GAGCGA------GUUCUAUCUCUCUUUUUUAG-CAAACUUGCAACUGUUGCACAAAAUAGUGCCUUCUACUGCAACUAAAUAUUGUUAUUUUC---CCUUCAC--UUUUU ((((.((---(((.((------(......))).)))))..))-))...(((((...(..((((......))))...)...)))))..................---.......--..... ( -17.50) >consensus GCUUUGA_U_GACUAAAUUGAUGCCUUGUCUUUCUUG_UUAU_AAAAGUUGCAUCUGUUGCACAAAUUGGUGC_UGCAGUUGCAGCUAAAUAUUGGUAUUAUUUU_________UUUUUU ..............................................(((((((.(((..((((......))))...))).)))))))................................. (-14.27 = -14.77 + 0.50)


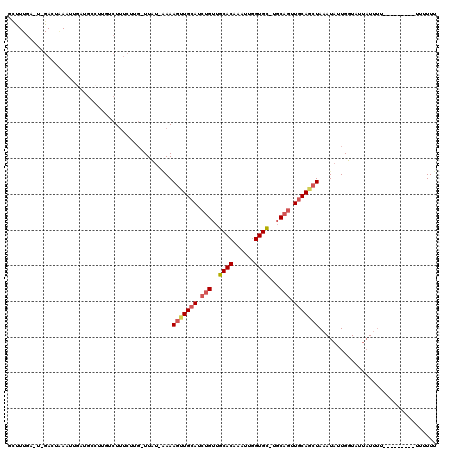
| Location | 18,459,711 – 18,459,820 |
|---|---|
| Length | 109 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.11 |
| Mean single sequence MFE | -19.67 |
| Consensus MFE | -9.39 |
| Energy contribution | -9.75 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.48 |
| SVM decision value | 2.39 |
| SVM RNA-class probability | 0.993363 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18459711 109 - 20766785 GAAAAA---------AAAAUUAGACCAAUAUUUAGCUGCAACUGCA-GCACCAAUUUGUGCAACAGAUGCAACUUUCAAAAA-CAAGAAAGACAAGGCAUCAAUUUAGUCCAGUCAAGGC ......---------.......(((.........(((((....)))-))................(((((..(((((.....-...))))).....)))))...........)))..... ( -22.50) >DroSec_CAF1 21757 85 - 1 AAAAAA---------AAAAUAAGACCAAUAUUUAGCUGCAACUGCA-GCACCAAUUUGUGCAACAGAUGCA-CUUUU-AAAA-CGAGAAAGACAAGG----------------------C ......---------.........((........(((((....)))-))........(((((.....))))-)....-....-............))----------------------. ( -16.30) >DroSim_CAF1 21958 106 - 1 AAAA-----------AAAAUAAGACCAAUAUUUAGCUGCAACUGCA-GCACCAAUUUGUGCAACAGAUGCAACUUUU-AAAA-CGAGAAAGACAAGGCAUCAAUUUAUUCCAGUCAAGGC ....-----------.......(((.........(((((....)))-))................(((((..(((((-....-...))))).....)))))...........)))..... ( -20.10) >DroEre_CAF1 21666 117 - 1 AAAAAAUAUGCAGUUAAAAUAAUAACAAUAUUUAGCUGCAACUGCA-ACACCAAUUUGUGCAACAGAUGCAACUUUU-AUAA-CAAGAAAGCAUAGGCAUCAAUUAAGUCAAAGCAAAAA ........((((((((((............))))))))))..(((.-.(((......))).....(((((..(((((-....-...))))).....)))))............))).... ( -22.00) >DroYak_CAF1 22408 116 - 1 AAAAAAAAU-CAGUUAAAAUAAUAACAAUAUUUAGCUGGAACUGCA-ACACCAAUUUGUGCAACAGAUGCAACUUUU-AUAA-CAAGAAAGAAAGGGCAACAAUUUAGUCAAAGCAAAAC ........(-((((((((............)))))))))..(((..-.(((......)))...))).(((..(((((-....-...)))))....(....)............))).... ( -15.40) >DroAna_CAF1 19836 105 - 1 AAAAA--GUGAAGG---GAAAAUAACAAUAUUUAGUUGCAGUAGAAGGCACUAUUUUGUGCAACAGUUGCAAGUUUG-CUAAAAAAGAGAGAUAGAAC------UCGCUC---UCAAACA ....(--(..((..---..(((((....)))))..((((((..(...((((......))))..)..)))))).))..-))......(((((...(...------.).)))---))..... ( -21.70) >consensus AAAAAA_________AAAAUAAGAACAAUAUUUAGCUGCAACUGCA_GCACCAAUUUGUGCAACAGAUGCAACUUUU_AAAA_CAAGAAAGACAAGGCAUCAAUUUAGUC_A_UCAAAAC ..................................((((((.(((...((((......))))..))).)))..(((((.........)))))....)))...................... ( -9.39 = -9.75 + 0.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:55:23 2006