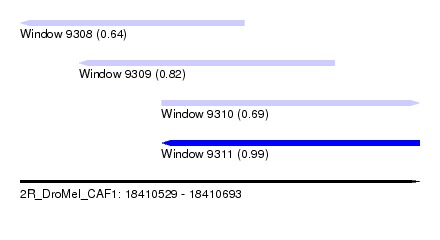

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,410,529 – 18,410,693 |
| Length | 164 |
| Max. P | 0.993393 |

| Location | 18,410,529 – 18,410,621 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 97.46 |
| Mean single sequence MFE | -26.66 |
| Consensus MFE | -25.36 |
| Energy contribution | -25.36 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.637038 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18410529 92 - 20766785 GCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAGCGGCACAAUGGUAACAACAUCAGC ((((((((((((((.((....------)).))))))))))).....((((((....)).))))............)))....((((......)))).. ( -26.50) >DroSec_CAF1 13519 92 - 1 GCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAGCGGCACAAUGGUAACAACAUCAGC ((((((((((((((.((....------)).))))))))))).....((((((....)).))))............)))....((((......)))).. ( -26.50) >DroSim_CAF1 13615 92 - 1 GCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAGCGGCACAAUGGUAACAACAUCAGC ((((((((((((((.((....------)).))))))))))).....((((((....)).))))............)))....((((......)))).. ( -26.50) >DroEre_CAF1 13579 92 - 1 GCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAGCGGCACAAUGGUAACAACAUCAGC ((((((((((((((.((....------)).))))))))))).....((((((....)).))))............)))....((((......)))).. ( -26.50) >DroYak_CAF1 13740 98 - 1 GCCUCGUUUGGUUUUGAUUACUCGUACUCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAGCGGCACAAUGGUAACAACAUCAGC ((((((((((((((.((.((....)).)).))))))))))).....((((((....)).))))............)))....((((......)))).. ( -27.30) >consensus GCCUCGUUUGGUUUUGAUUAC______UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAGCGGCACAAUGGUAACAACAUCAGC ((((((((((((((.((..........)).))))))))))).....((((((....)).))))............)))....((((......)))).. (-25.36 = -25.36 + -0.00)

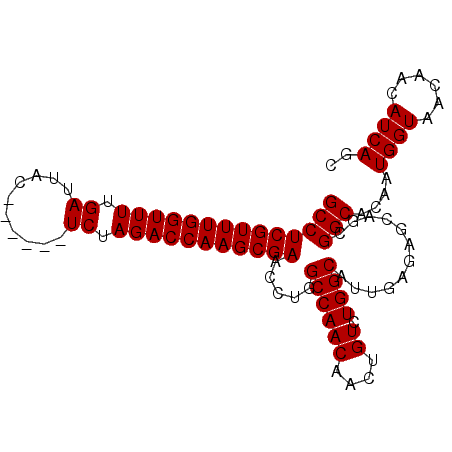
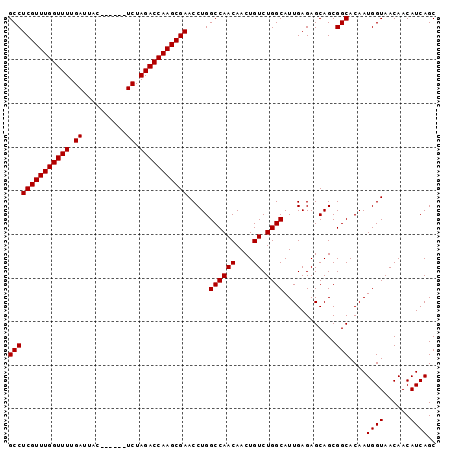
| Location | 18,410,553 – 18,410,658 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 96.28 |
| Mean single sequence MFE | -37.68 |
| Consensus MFE | -34.78 |
| Energy contribution | -35.18 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.821226 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18410553 105 - 20766785 ACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAG ..(.((..((((((((((.(..(....)..).)).)))))))(((((((((.((....------)).))))))))).......((((((....)).))))...)..))).. ( -38.60) >DroSec_CAF1 13543 105 - 1 ACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAG ..(.((..((((((((((.(..(....)..).)).)))))))(((((((((.((....------)).))))))))).......((((((....)).))))...)..))).. ( -38.60) >DroSim_CAF1 13639 105 - 1 ACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAG ..(.((..((((((((((.(..(....)..).)).)))))))(((((((((.((....------)).))))))))).......((((((....)).))))...)..))).. ( -38.60) >DroEre_CAF1 13603 102 - 1 --GACUUCCGACGC-GCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAG --(.((..(((.((-((((.((.......((....))((((((((((((((.((....------)).)))))))))))....)))...)).))))...)).)))..))).. ( -33.20) >DroYak_CAF1 13764 111 - 1 ACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUACUCGUACUCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAG ..(.((..((((((((((.(..(....)..).)).)))))))(((((((((.((.((....)).)).))))))))).......((((((....)).))))...)..))).. ( -39.40) >consensus ACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC______UCUAGACCAAGCGAACCUGGCCAACAACUGUCUGGCAUUGAGAGCAG ..(.((..((((((((((.(..(....)..).)).)))))))(((((((((.((..........)).))))))))).......((((((....)).))))...)..))).. (-34.78 = -35.18 + 0.40)


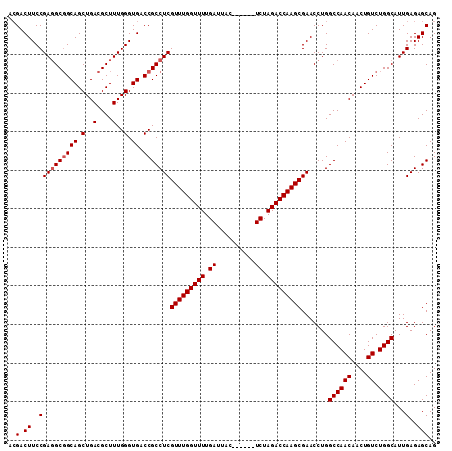
| Location | 18,410,587 – 18,410,693 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 93.48 |
| Mean single sequence MFE | -32.21 |
| Consensus MFE | -25.77 |
| Energy contribution | -26.57 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685784 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18410587 106 + 20766785 CGCUUGGUCUAGA------GUAAUCAAAACCAAACGAGGCGGUCACCCAAAGCGUCAGCUGCCGCCUCGGAAGUCGUUGCCGCACCGUUAUGGCUUAUAAAAAUGUUUAUAU----- ...(((((...((------....))...))))).(((((((((.......(((....)))))))))))).(((((((.((......)).)))))))................----- ( -32.41) >DroSec_CAF1 13577 106 + 1 CGCUUGGUCUAGA------GUAAUCAAAACCAAACGAGGCGGUCACCCAAAGCGUCAGCUGCCGCCUCGGAAGUCGUUGCGGCACCGUUAUGGCUUAUAAAAAUGUUUAUAU----- ...(((((...((------....))...))))).(((((((((.......(((....)))))))))))).(((((((.(((....))).)))))))................----- ( -36.61) >DroSim_CAF1 13673 106 + 1 CGCUUGGUCUAGA------GUAAUCAAAACCAAACGAGGCGGUCACCCAAAGCGUCAGCUGCCGCCUCGGAAGUCGUUGCCGCACCGUUAUGGCUUAUAAAAAUGUUUAUAU----- ...(((((...((------....))...))))).(((((((((.......(((....)))))))))))).(((((((.((......)).)))))))................----- ( -32.41) >DroEre_CAF1 13637 101 + 1 CGCUUGGUCUAGA------GUAAUCAAAACCAAACGAGGCGGUCACCCAAAGCGUCAGCUGC-GCGUCGGAAGUC---GCCGCACCGUUAUGGCUUAUAAAAAUGUUUAUA------ .(((((((...((------....))...))))((((..(((((.((((...((((.....))-))...))..)).---)))))..))))..))).................------ ( -28.40) >DroYak_CAF1 13798 117 + 1 CGCUUGGUCUAGAGUACGAGUAAUCAAAACCAAACGAGGCGGUCACCCAAAGCGUCAGCUGCCGCCUCGGAAGUCGUUGCCGCACCGUUAUGGCUUAUAAAAAUGUUUAUAUAUAUA .(((((..........))))).....((((....(((((((((.......(((....)))))))))))).(((((((.((......)).)))))))........))))......... ( -31.21) >consensus CGCUUGGUCUAGA______GUAAUCAAAACCAAACGAGGCGGUCACCCAAAGCGUCAGCUGCCGCCUCGGAAGUCGUUGCCGCACCGUUAUGGCUUAUAAAAAUGUUUAUAU_____ ...(((((...((..........))...))))).(((((((((.......(((....)))))))))))).(((((((.((......)).)))))))..................... (-25.77 = -26.57 + 0.80)

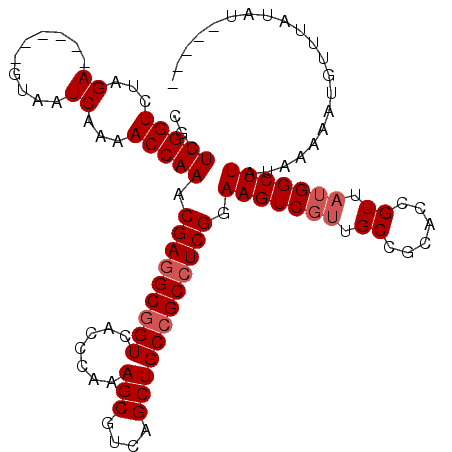
| Location | 18,410,587 – 18,410,693 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 93.48 |
| Mean single sequence MFE | -35.84 |
| Consensus MFE | -30.56 |
| Energy contribution | -31.36 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.85 |
| SVM decision value | 2.39 |
| SVM RNA-class probability | 0.993393 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18410587 106 - 20766785 -----AUAUAAACAUUUUUAUAAGCCAUAACGGUGCGGCAACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCG -----.((((((....)))))).........((..((....))....))(((((((((.(..(....)..).)).)))))))(((((((((.((....------)).))))))))). ( -37.70) >DroSec_CAF1 13577 106 - 1 -----AUAUAAACAUUUUUAUAAGCCAUAACGGUGCCGCAACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCG -----.((((((....)))))).((.....(((((((((..((.....))..)))))).))).......((....))))..((((((((((.((....------)).)))))))))) ( -34.80) >DroSim_CAF1 13673 106 - 1 -----AUAUAAACAUUUUUAUAAGCCAUAACGGUGCGGCAACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCG -----.((((((....)))))).........((..((....))....))(((((((((.(..(....)..).)).)))))))(((((((((.((....------)).))))))))). ( -37.70) >DroEre_CAF1 13637 101 - 1 ------UAUAAACAUUUUUAUAAGCCAUAACGGUGCGGC---GACUUCCGACGC-GCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC------UCUAGACCAAGCG ------((((((....)))))).(((.....)))..(((---(...(((((.((-((....).))).)))))....)))).((((((((((.((....------)).)))))))))) ( -30.50) >DroYak_CAF1 13798 117 - 1 UAUAUAUAUAAACAUUUUUAUAAGCCAUAACGGUGCGGCAACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUACUCGUACUCUAGACCAAGCG ......((((((....)))))).........((..((....))....))(((((((((.(..(....)..).)).)))))))(((((((((.((.((....)).)).))))))))). ( -38.50) >consensus _____AUAUAAACAUUUUUAUAAGCCAUAACGGUGCGGCAACGACUUCCGAGGCGGCAGCUGACGCUUUGGGUGACCGCCUCGUUUGGUUUUGAUUAC______UCUAGACCAAGCG ......((((((....)))))).........((..((....))....))(((((((((.(..(....)..).)).)))))))(((((((((.((..........)).))))))))). (-30.56 = -31.36 + 0.80)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:55:04 2006