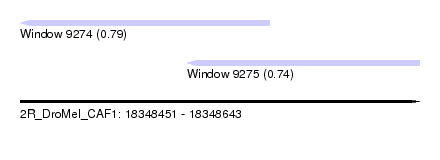
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,348,451 – 18,348,643 |
| Length | 192 |
| Max. P | 0.790619 |

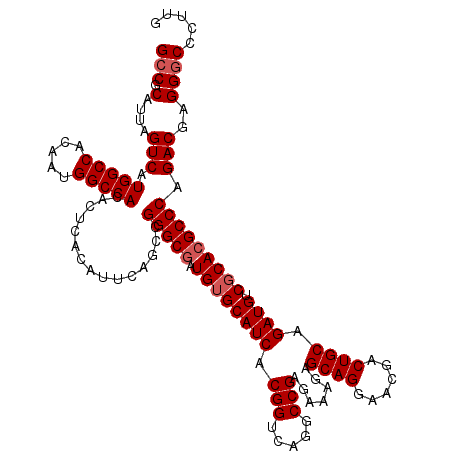
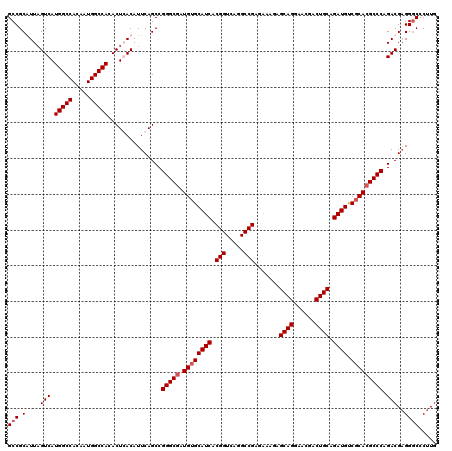
| Location | 18,348,451 – 18,348,571 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.56 |
| Mean single sequence MFE | -48.95 |
| Consensus MFE | -43.03 |
| Energy contribution | -43.77 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.790619 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18348451 120 - 20766785 GCCGCGUUAGUCAUGGCCACAAUGGCCACACUGACAUUCAGCCGGGCAAUGUGCAUCACGGUCAGGCCGAGAAAGAGCAGGAACGACUGCAGAUGUCGCACGCCCAGACGAGGGCCCUUG (((.((((.(((((((((.....)))))...))))........((((..((((((((.(((.....))).......((((......)))).)))).)))).)))).))))..)))..... ( -49.00) >DroSim_CAF1 208 120 - 1 GCCGCGUUAGUCAUGGCCACAAUGGCCACACUGACAUUCAGCCGGGCGAUGUGCAUCACGGUCAGGCCGAGAAAGAGCAGGAACGACUGCAGAUGUCUCACGCCCAGACGAGGGCCCUUG (((.((((.(((((((((.....)))))...))))........(((((.((.(((((.(((.....))).......((((......)))).)))))..))))))).))))..)))..... ( -49.80) >DroEre_CAF1 30 120 - 1 GCCGCAUUAGUCAUGGCCACAAUGGCCACACUCACAUUCAGCCGGGCGAUGUGCAUCACGGUCAGGCCGAGAAAGAGCAGGAACGACUGCAGAUGUCGCACGCCCAGACGAGGCCCCUUG (((......(((.(((((.....)))))...............(((((.((((((((.(((.....))).......((((......)))).)))).))))))))).)))..)))...... ( -47.60) >DroYak_CAF1 348 120 - 1 GCCGCAUUUGUCAUGGCCACAAUGGCCACACUCACAUUCAGCCGGGCGAUGUGCAUCACGGUGAGGCCGAGGAAGAGCAGGAACGACUGCAGAUGCCGCACGCCCAGACGUGGGCCCUUG (((.(((..(((.(((((.....)))))...............(((((.((((((((.(((.....))).......((((......)))).)))).))))))))).)))))))))..... ( -49.40) >consensus GCCGCAUUAGUCAUGGCCACAAUGGCCACACUCACAUUCAGCCGGGCGAUGUGCAUCACGGUCAGGCCGAGAAAGAGCAGGAACGACUGCAGAUGUCGCACGCCCAGACGAGGGCCCUUG (((.(....(((.(((((.....)))))...............(((((.((((((((.(((.....))).......((((......)))).)))).))))))))).)))..))))..... (-43.03 = -43.77 + 0.75)

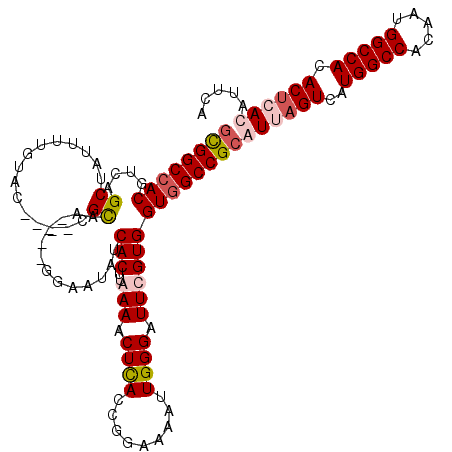
| Location | 18,348,531 – 18,348,643 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.80 |
| Mean single sequence MFE | -36.50 |
| Consensus MFE | -26.17 |
| Energy contribution | -27.73 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.744928 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18348531 112 - 20766785 GCGGCCACGUCAGCUAUUUCCAAC----CGCAA----GGAAUAUCACUUAAAACUCACCGGAAAAUUGGGAUUCGUGGUGGCCGCGUUAGUCAUGGCCACAAUGGCCACACUGACAUUCA ((((((((....((...(((((((----(....----))...............((....))...))))))...)).))))))))((((((..(((((.....))))).))))))..... ( -42.70) >DroSim_CAF1 288 112 - 1 GCGGCCACGUCAGCUAUUUUGUAC----CGUAA----GGAAUAUCACUUAAAACUCACCGGAAAGUUGGGAUUCGUGGUAGCCGCGUUAGUCAUGGCCACAAUGGCCACACUGACAUUCA (((((((((((((((.((..((..----..(((----(........))))......))..)).))))))....))))...)))))((((((..(((((.....))))).))))))..... ( -36.60) >DroEre_CAF1 110 116 - 1 AUGGCCACGUCAGCUAGGUUGUACCUUGCGCAG----AGAAUAUCACUUGAAGCUUACCGGAAAAUUGGGAUUCGUGGUGGCCGCAUUAGUCAUGGCCACAAUGGCCACACUCACAUUCA ..((((((....((.(((.....))).))....----.......(((..(((.....((((....))))..)))))))))))).....(((..(((((.....))))).)))........ ( -31.10) >DroYak_CAF1 428 116 - 1 GUGGCCACGUCAGC----UUGUGCCUUGUGCAGUUUGGUAAUAUCACUUGAAACUUACCGGAAAAUUGGGAUUCGUGGUGGCCGCAUUUGUCAUGGCCACAAUGGCCACACUCACAUUCA ((((((((....((----..((.(((.((....((((((((..((....))...))))))))..)).)))))..)).))))))))........(((((.....)))))............ ( -35.60) >consensus GCGGCCACGUCAGCUAUUUUGUAC____CGCAA____GGAAUAUCACUUAAAACUCACCGGAAAAUUGGGAUUCGUGGUGGCCGCAUUAGUCAUGGCCACAAUGGCCACACUCACAUUCA ((((((((....((...............)).............(((..(((.((((.........)))).))))))))))))))((((((..(((((.....))))).))))))..... (-26.17 = -27.73 + 1.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:54:28 2006