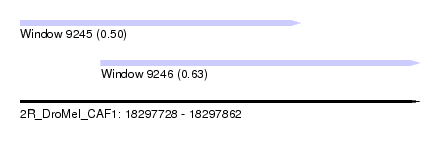
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,297,728 – 18,297,862 |
| Length | 134 |
| Max. P | 0.631696 |

| Location | 18,297,728 – 18,297,822 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 87.45 |
| Mean single sequence MFE | -22.42 |
| Consensus MFE | -16.46 |
| Energy contribution | -18.26 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.73 |
| SVM decision value | -0.08 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18297728 94 + 20766785 ACCGGCGUAAGUAUUCACAAAGGAAUGCGUUAAGUAAUAGCAUUUAAGUAGUGCAAAUGGGAAAGUGUUGUGUUUAUCAAUGCACUUUAUGCAA ....((((((((((((......))))))....(((....(((((.....)))))..........((((((.......))))))))))))))).. ( -20.60) >DroSec_CAF1 95008 94 + 1 UCCGGCGUAAGUAUUCACAAAGGAAUGCGUUAAGUAAUAGCAUUUAAGGAGUGCAAAUGGGAAAGCGUUAUGUUUGUCAUUGCACUUUAUGCAA ....(((((.((((((......))))))((((.....)))).......((((((((.(((..((((.....)))).)))))))))))))))).. ( -23.80) >DroSim_CAF1 95769 94 + 1 UCCGGCGUAAGUAUUCACAAAGGAAUGCGUUAAGUAAUAGCAUUUAAGGAGUGCAAAUGGGAAAGCGUUAUGUUUGUCAUUGCACUUUAUGCAA ....(((((.((((((......))))))((((.....)))).......((((((((.(((..((((.....)))).)))))))))))))))).. ( -23.80) >DroEre_CAF1 98749 78 + 1 UCCGGCGUAAGUAUUCACAAAGGAAUGCGUUA----AUAGCAUUUAAGGGGUGC------------UUCAUGUUUGCCAUUGCACUUUAGGCAA ...(((((((((((((......((((((....----...))))))...))))))------------)).)))))((((...........)))). ( -20.00) >DroYak_CAF1 96749 94 + 1 UCCGGCGUAAGUAUUCACAAAGGAAUGCGUUAAGUAAUAGCAUUUAAGGAGUGCAAACGGAGAAGCGUUAUGUUUGUCAUUGCACUUUAGGCAA ....((.((.((((((......))))))((((.....)))).......((((((((...((.((((.....)))).)).)))))))))).)).. ( -23.90) >consensus UCCGGCGUAAGUAUUCACAAAGGAAUGCGUUAAGUAAUAGCAUUUAAGGAGUGCAAAUGGGAAAGCGUUAUGUUUGUCAUUGCACUUUAUGCAA ....((((((((((((......))))))((((.....))))........(((((((...((.((((.....)))).)).))))))))))))).. (-16.46 = -18.26 + 1.80)

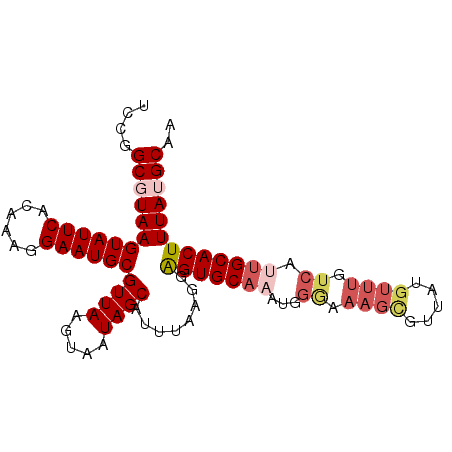
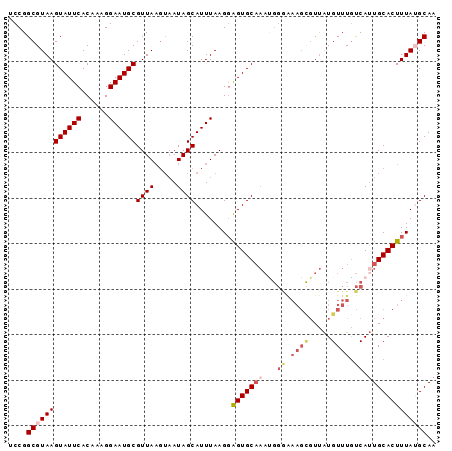
| Location | 18,297,755 – 18,297,862 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 80.60 |
| Mean single sequence MFE | -24.62 |
| Consensus MFE | -13.61 |
| Energy contribution | -14.22 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.631696 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18297755 107 + 20766785 CGUUAAGUAAUAGCAUUUAAGUAGUGCAAAUGGGAAAGUGUUGUGUUUAUCAAUGCACUUUAUGCAAAUGUGCAAAUAUUUCUUUAAUAUCUUUAAAAGUGCAACCA ............(((((.....)))))...(((......((((.......))))(((((((.((((....)))).(((((.....))))).....)))))))..))) ( -19.00) >DroSec_CAF1 95035 107 + 1 CGUUAAGUAAUAGCAUUUAAGGAGUGCAAAUGGGAAAGCGUUAUGUUUGUCAUUGCACUUUAUGCAAAUGUGCAAAUAUUUCUUUAAAAUCUUUAAAUGUGCAAGAU ............(((((((((((((((((.(((..((((.....)))).)))))))))....((((....))))...............)))))))))))....... ( -24.40) >DroSim_CAF1 95796 107 + 1 CGUUAAGUAAUAGCAUUUAAGGAGUGCAAAUGGGAAAGCGUUAUGUUUGUCAUUGCACUUUAUGCAAAUGUGCAAAUAUUUCUUUAAAAUCUUUAAAUGGGCAACCC .((((.....))))(((((((((((((((.(((..((((.....)))).)))))))))....((((....))))...............)))))))))((....)). ( -24.30) >DroEre_CAF1 98776 88 + 1 CGUUA----AUAGCAUUUAAGGGGUGC------------UUCAUGUUUGCCAUUGCACUUUAGGCAAAUGUGCAAAUAUUU---AAGGAUCCUUAAAAGUGCUCCCA .....----..(((((((.((((((.(------------((.((((((((((((((.(....))).)))).))))))))..---))).))))))..))))))).... ( -27.30) >DroYak_CAF1 96776 104 + 1 CGUUAAGUAAUAGCAUUUAAGGAGUGCAAACGGAGAAGCGUUAUGUUUGUCAUUGCACUUUAGGCAAAUGGGCAAAUAUUU---AAGGUUCUUUAUAAGUGUACCCA .((((.....))))......((.(((((.(..(((((.(...(((((((((.((((.(....)))))...)))))))))..---..).)))))..)...))))))). ( -28.10) >consensus CGUUAAGUAAUAGCAUUUAAGGAGUGCAAAUGGGAAAGCGUUAUGUUUGUCAUUGCACUUUAUGCAAAUGUGCAAAUAUUUCUUUAAAAUCUUUAAAAGUGCAACCA ............(((((((((((.....((((......))))((((((((((((((.......)).)))).))))))))..........)))))))..))))..... (-13.61 = -14.22 + 0.61)

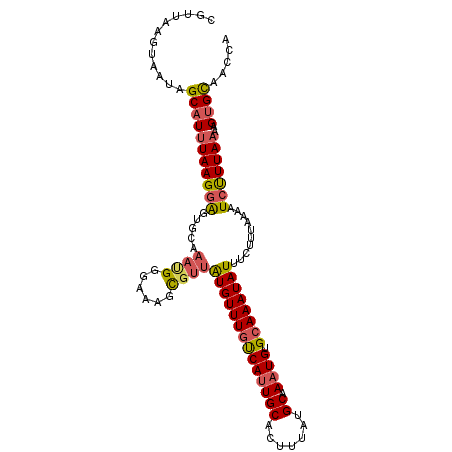
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:54:01 2006