
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,280,422 – 18,280,513 |
| Length | 91 |
| Max. P | 0.993715 |

| Location | 18,280,422 – 18,280,513 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 81.56 |
| Mean single sequence MFE | -25.27 |
| Consensus MFE | -17.68 |
| Energy contribution | -18.20 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.95 |
| Structure conservation index | 0.70 |
| SVM decision value | 2.42 |
| SVM RNA-class probability | 0.993715 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
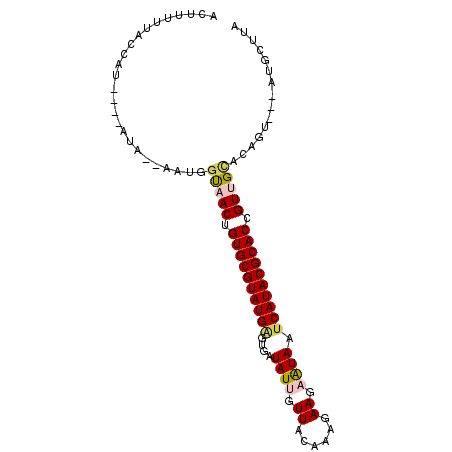
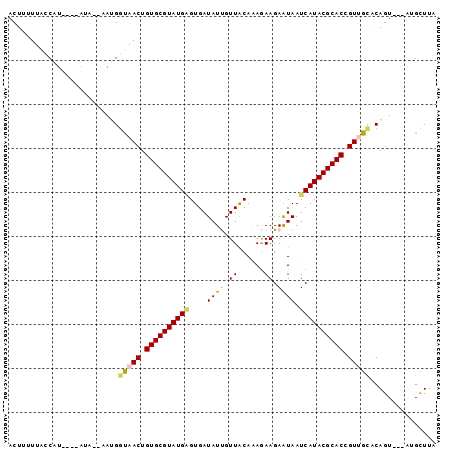
>2R_DroMel_CAF1 18280422 91 + 20766785 ACUUUUUACCAU----AUA--UAUGGUAACUGUGCGUAUGAGUGAUAUUGUUGCAAAGAAGAAUAAUCAUACGCACCGUUGUGCAGUUAAGUGCUUA ......((((((----...--.))))))((.((((((((((....((((.((......)).)))).)))))))))).)).(..(......)..)... ( -24.80) >DroSec_CAF1 78173 88 + 1 ACUUUUUACCAU----AUG--UAUGGUAACUGUGCGUAUGAGUGAUAUUGUUACAAAGAAGAAUAAUCAUACGCACCGUUGCCCAGU---AUGCUUA .........(((----((.--...((((((.((((((((((....((((.((......)).)))).)))))))))).))))))..))---))).... ( -28.90) >DroSim_CAF1 79236 88 + 1 ACUUUUUACCAU----AUG--AAUGGUAACUGUGCGUAUGAGUGAUAUUGUUACAAAGAAGAAUAAUCAUACGCACCGUUGCCCAGU---AUGCUUA .........(((----((.--...((((((.((((((((((....((((.((......)).)))).)))))))))).))))))..))---))).... ( -28.90) >DroEre_CAF1 81713 92 + 1 ACUUUU--ACAUUGAGAAAGUAAGGGCAACUGUGCGUAUGCGUGAUAUCGUUACACAGAAGAAUAAUCAUACGCACUGUGGUACAGU---AAGCUUA ((((((--.......))))))..((((.(((((((....(((......)))..(((((.................))))))))))))---..)))). ( -19.83) >DroYak_CAF1 79189 90 + 1 ACUUUUUAACAU----GAAUUGGGGACAACUGUGCGUAUGGGUGAUAUCGUUACACAAAAGAGUAAUCAUACGCACUGUGGCACAGU---AAGUUUA .......(((..----....((....))(((((((.((((((((.(((.(((((........))))).)))))).))))))))))))---..))).. ( -23.90) >consensus ACUUUUUACCAU____AUA__AAUGGUAACUGUGCGUAUGAGUGAUAUUGUUACAAAGAAGAAUAAUCAUACGCACCGUUGCACAGU___AUGCUUA .........................(((((.((((((((((....((((.((......)).)))).)))))))))).)))))............... (-17.68 = -18.20 + 0.52)

| Location | 18,280,422 – 18,280,513 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 81.56 |
| Mean single sequence MFE | -19.77 |
| Consensus MFE | -12.83 |
| Energy contribution | -12.83 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873235 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
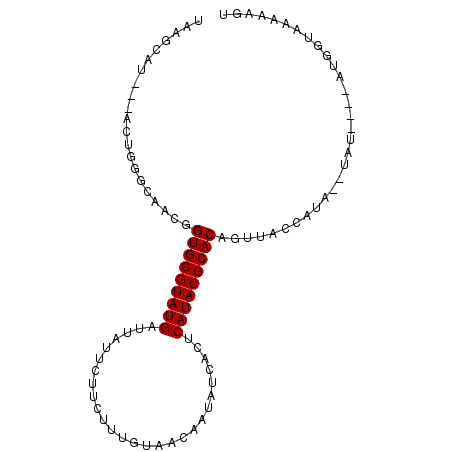
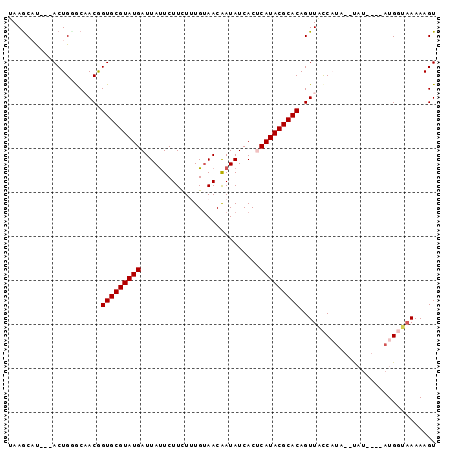
>2R_DroMel_CAF1 18280422 91 - 20766785 UAAGCACUUAACUGCACAACGGUGCGUAUGAUUAUUCUUCUUUGCAACAAUAUCACUCAUACGCACAGUUACCAUA--UAU----AUGGUAAAAAGU ...(((......)))......((((((((((.((((.((......)).))))....))))))))))..(((((((.--...----)))))))..... ( -22.60) >DroSec_CAF1 78173 88 - 1 UAAGCAU---ACUGGGCAACGGUGCGUAUGAUUAUUCUUCUUUGUAACAAUAUCACUCAUACGCACAGUUACCAUA--CAU----AUGGUAAAAAGU .......---....(....).((((((((((.((((.((......)).))))....))))))))))..(((((((.--...----)))))))..... ( -21.30) >DroSim_CAF1 79236 88 - 1 UAAGCAU---ACUGGGCAACGGUGCGUAUGAUUAUUCUUCUUUGUAACAAUAUCACUCAUACGCACAGUUACCAUU--CAU----AUGGUAAAAAGU .......---....(....).((((((((((.((((.((......)).))))....))))))))))..(((((((.--...----)))))))..... ( -21.60) >DroEre_CAF1 81713 92 - 1 UAAGCUU---ACUGUACCACAGUGCGUAUGAUUAUUCUUCUGUGUAACGAUAUCACGCAUACGCACAGUUGCCCUUACUUUCUCAAUGU--AAAAGU ...((((---...........(((((((((........((........)).......)))))))))........((((.........))--)))))) ( -16.16) >DroYak_CAF1 79189 90 - 1 UAAACUU---ACUGUGCCACAGUGCGUAUGAUUACUCUUUUGUGUAACGAUAUCACCCAUACGCACAGUUGUCCCCAAUUC----AUGUUAAAAAGU .......---(((.....((((((((((((.((((.(....).)))).(....)...)))))))))(((((....))))).----.))).....))) ( -17.20) >consensus UAAGCAU___ACUGGGCAACGGUGCGUAUGAUUAUUCUUCUUUGUAACAAUAUCACUCAUACGCACAGUUACCAUA__UAU____AUGGUAAAAAGU .....................(((((((((...........................)))))))))............................... (-12.83 = -12.83 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:53:53 2006