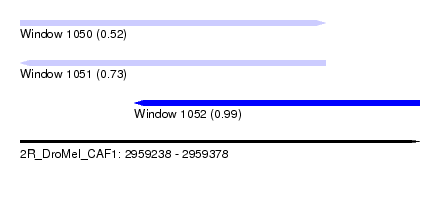

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 2,959,238 – 2,959,378 |
| Length | 140 |
| Max. P | 0.989442 |

| Location | 2,959,238 – 2,959,345 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 79.20 |
| Mean single sequence MFE | -28.28 |
| Consensus MFE | -18.59 |
| Energy contribution | -19.15 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.521705 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
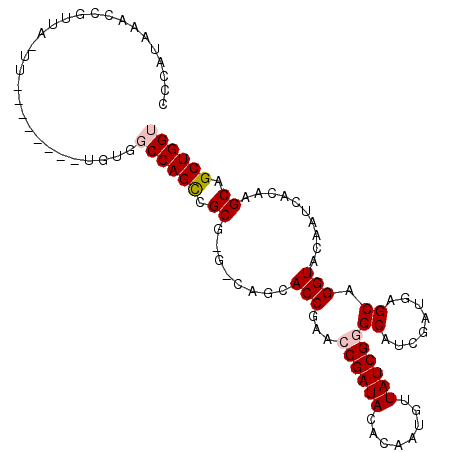
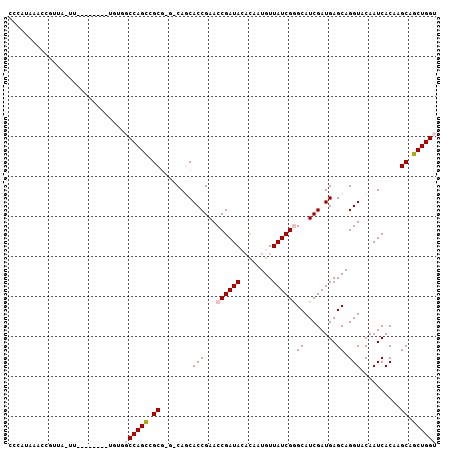
>2R_DroMel_CAF1 2959238 107 + 20766785 CCCAUAAACCGUUAGUUGGCUCGAAUGUGGCCAGCCGCGGGUCAGCACCGAACCGAUAUAUAAUGUUAUCGGGCAUCGAUGAGCAGGUACAAUCACAAGCAGCUGGU ........((((..(((((((.......))))))).)))).(((((..(((.((((((........))))))...)))....((..((......))..)).))))). ( -33.50) >DroSec_CAF1 2351 85 + 1 CAAAUUAACCG-------------------CCAGUCGCG---CAUCACCGAACCGAUACACAAUGUUAUCGGGCAUCGAUGAGCAGGUACAAUCACAAGCAGCUGGU ..........(-------------------(((((.(((---(.(((.(((.((((((........))))))...))).)))))..((......))..)).)))))) ( -25.40) >DroSim_CAF1 1035 85 + 1 CACAUAAACCG-------------------CCAGCCGCG---CAUCACCGAACCGAUAGACAAUGUUAUCGGGCAUCGAUGAGCAGGUACAAUCACAAGCAGCUGGU ..........(-------------------(((((.(((---(.(((.(((.(((((((......)))))))...))).)))))..((......))..)).)))))) ( -27.90) >DroEre_CAF1 970 107 + 1 UCCAUAAACCGUUAAUUCAAUAUGGUGUGGCCAGUCGCAAGGCAGCACCGAACCGAUACUCAAUGUUAUCGAGCAUCGAUGAGCAGGUACAAUCACAAGCAGCUGGU .((((..(((((.........)))))))))(((((.((..((.....))(.(((....(((((((((....)))))...))))..))).)........)).))))). ( -25.50) >DroYak_CAF1 940 107 + 1 CCCAUAAACCGUUAAUUCAACAUGGUGUGGCCAGCCGCAAGGCAGCACCGAACCGAUAUAUACUGUUAUCGAGCAUCGAUGAGCAGGUACAAUCACAAGCAGCUGGU .((((..(((((.........)))))))))(((((.((..((.....)).....(((.....(((((((((.....)))))).))).....)))....)).))))). ( -29.10) >consensus CCCAUAAACCGUUA_UU________UGUGGCCAGCCGCG_G_CAGCACCGAACCGAUACACAAUGUUAUCGGGCAUCGAUGAGCAGGUACAAUCACAAGCAGCUGGU .............................((((((.((........(((...((((((........))))))((........)).)))..........)).)))))) (-18.59 = -19.15 + 0.56)

| Location | 2,959,238 – 2,959,345 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 79.20 |
| Mean single sequence MFE | -32.06 |
| Consensus MFE | -23.78 |
| Energy contribution | -23.94 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.734189 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
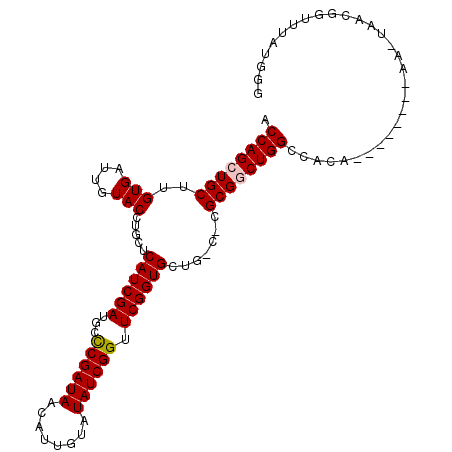
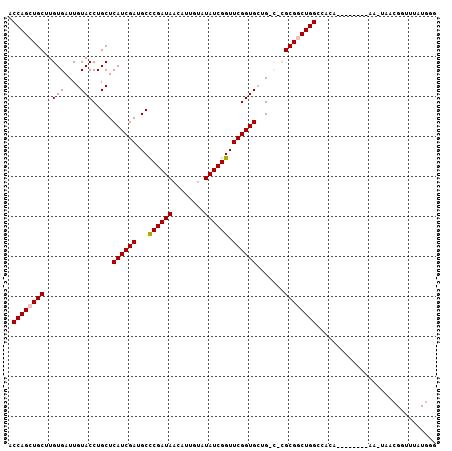
>2R_DroMel_CAF1 2959238 107 - 20766785 ACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCCCGAUAACAUUAUAUAUCGGUUCGGUGCUGACCCGCGGCUGGCCACAUUCGAGCCAACUAACGGUUUAUGGG .((((((((..((.(..(((((.((........))((((((........))))))...)))))).))..))))))))((.(((..(((((.......)))))))))) ( -34.90) >DroSec_CAF1 2351 85 - 1 ACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCCCGAUAACAUUGUGUAUCGGUUCGGUGAUG---CGCGACUGG-------------------CGGUUAAUUUG ...(((((((...(.((((....(((((((((...((((((........)))))).))))))).)---)))))).))-------------------)))))...... ( -26.50) >DroSim_CAF1 1035 85 - 1 ACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCCCGAUAACAUUGUCUAUCGGUUCGGUGAUG---CGCGGCUGG-------------------CGGUUUAUGUG .((((((((..(((....)))..(((((((((...((((((........)))))).))))))).)---)))))))))-------------------........... ( -32.60) >DroEre_CAF1 970 107 - 1 ACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCUCGAUAACAUUGAGUAUCGGUUCGGUGCUGCCUUGCGACUGGCCACACCAUAUUGAAUUAACGGUUUAUGGA .((((.(((..(.(...(((((.....(((((((((((((...)))))))))))))..)))))..))..))).)))).....(((((..............))))). ( -34.14) >DroYak_CAF1 940 107 - 1 ACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCUCGAUAACAGUAUAUAUCGGUUCGGUGCUGCCUUGCGGCUGGCCACACCAUGUUGAAUUAACGGUUUAUGGG .((((((((..(.(...(((((.((........))((((((........))))))...)))))..))..)))))))).....(((((..............))))). ( -32.14) >consensus ACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCCCGAUAACAUUGUAUAUCGGUUCGGUGCUG_C_CGCGGCUGGCCACA________AA_UAACGGUUUAUGGG .((((((((..(((....))).....((((((...((((((........)))))).)))))).......)))))))).............................. (-23.78 = -23.94 + 0.16)

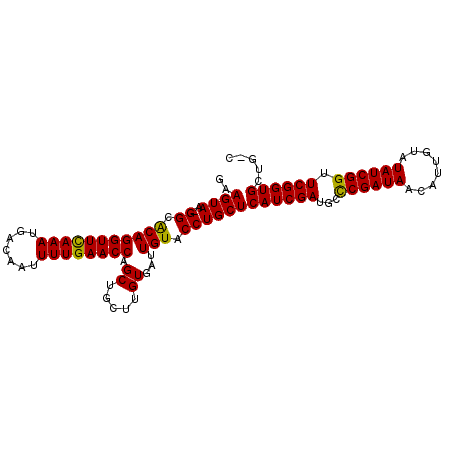
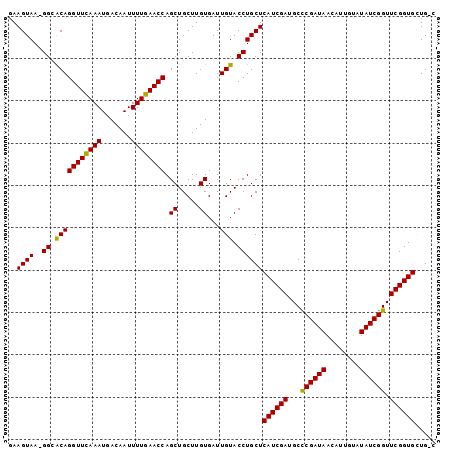
| Location | 2,959,278 – 2,959,378 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 92.81 |
| Mean single sequence MFE | -30.16 |
| Consensus MFE | -26.10 |
| Energy contribution | -25.54 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.87 |
| SVM decision value | 2.16 |
| SVM RNA-class probability | 0.989442 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 2959278 100 - 20766785 GAAGUAA-GGCACAGGUUCAAAUGACAAUUUUGAACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCCCGAUAACAUUAUAUAUCGGUUCGGUGCUGAC ..((((.-((.(((((((((((.......)))))))).((.....))...))).))))))((((((...((((((........)))))).))))))..... ( -28.20) >DroSec_CAF1 2371 98 - 1 GAAGUAA-GGCACAGGUUUAAAUGACAAUUUUGAACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCCCGAUAACAUUGUGUAUCGGUUCGGUGAUG-- ...(((.-((.(((((((((((.......)))))))).((.....))...))).)))))(((((((...((((((........)))))).)))))))..-- ( -27.50) >DroSim_CAF1 1055 98 - 1 GAAGUAA-GGCACAGGUUCAAAUGACAAUUUUGAACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCCCGAUAACAUUGUCUAUCGGUUCGGUGAUG-- ...(((.-((.(((((((((((.......)))))))).((.....))...))).)))))(((((((...((((((........)))))).)))))))..-- ( -29.90) >DroEre_CAF1 1010 100 - 1 AAAGUAA-GGGGCAGGUUCAAAUGACAAUUUUGAACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCUCGAUAACAUUGAGUAUCGGUUCGGUGCUGCC ....(((-(.(((.((((((((.......)))))))).))).)))).....(((((.....(((((((((((((...)))))))))))))..))))).... ( -37.80) >DroYak_CAF1 980 101 - 1 AAAGUAAAGGGACAGGUUCAAAUGACAAUUUUGAACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCUCGAUAACAGUAUAUAUCGGUUCGGUGCUGCC .......(((.(((((((((((.......)))))))).((.....))...))).)))((.((((((.((.(((((........)))))))))))))..)). ( -27.40) >consensus GAAGUAA_GGCACAGGUUCAAAUGACAAUUUUGAACCAGCUGCUUGUGAUUGUACCUGCUCAUCGAUGCCCGAUAACAUUGUAUAUCGGUUCGGUGCUG_C ..((((..((.(((((((((((.......)))))))).((.....))...))).))))))((((((...((((((........)))))).))))))..... (-26.10 = -25.54 + -0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:42:00 2006