| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,200,682 – 18,200,774 |
| Length | 92 |
| Max. P | 0.992439 |

| Location | 18,200,682 – 18,200,774 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 80.04 |
| Mean single sequence MFE | -26.86 |
| Consensus MFE | -14.83 |
| Energy contribution | -16.71 |
| Covariance contribution | 1.88 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.62 |
| Structure conservation index | 0.55 |
| SVM decision value | 1.69 |
| SVM RNA-class probability | 0.972363 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
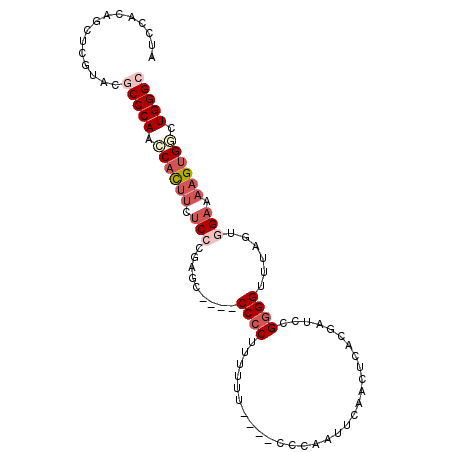
>2R_DroMel_CAF1 18200682 92 + 20766785 AUCCACAGCUCGUAUGCCCAACCACUUCUCGCGAGC----CCCCUUUUUU----CCCAAUUCAACUCACGAUCCGGGGUUUAGUGGAAAAGUGUCUGGGC ...............(((((..(((((.((((((((----(((.......----....................))))))).))))..)))))..))))) ( -25.53) >DroSec_CAF1 25780 91 + 1 AUCCACAGCUCGUACGCCCAACCACU-CUCCCGAGC----CCCCUGUUUU----CCCAAUUCAACUCACGAUCCGGGGUUUAGUGGAAAAGUGGCUGGGC ...............(((((.(((((-.((((((((----(((..(((..----...............)))..))))))).).)))..))))).))))) ( -28.13) >DroSim_CAF1 27500 92 + 1 AUCCACAGCUCGUACGCCCAACCACUUCUCCCGAGC----CCCCUUUUUU----UCCAAUUCAACUCACGAUCCGGGGUUUAGUGGAAAAGUGGCUGGGC ...............(((((.((((((.((((((((----(((.......----....................))))))).).))).)))))).))))) ( -30.63) >DroEre_CAF1 26444 96 + 1 AUCCACAGCUCGUACUCCCAAGCACUUCUCCCUAUCCAUUCCCCCUUUUU----CCGGAUUCAACUCACGAUCCGUGGUUUAGUGGAAAAGUGCCUGGGC ................((((.((((((.((((((.(((............----.((((((........)))))))))..))).))).)))))).)))). ( -28.41) >DroYak_CAF1 27251 83 + 1 -------------ACUCCCAAGCUUAUAACACUAUA----CCCCUUCUUUCUAUUCCGAUUCAACUCACGAUCCGGGGUUUAGUGGAAAAGUGGAUGGGC -------------...((((.((((....(((((.(----((((.....((......)).((.......))...))))).)))))...))))...)))). ( -21.60) >consensus AUCCACAGCUCGUACGCCCAACCACUUCUCCCGAGC____CCCCUUUUUU____CCCAAUUCAACUCACGAUCCGGGGUUUAGUGGAAAAGUGGCUGGGC ...............(((((.((((((.(((.........((((..............................))))......))).)))))).))))) (-14.83 = -16.71 + 1.88)

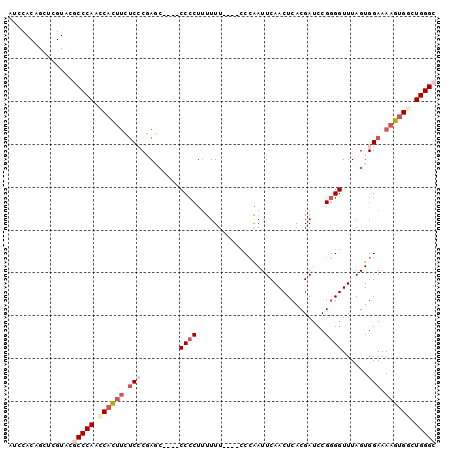
| Location | 18,200,682 – 18,200,774 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 80.04 |
| Mean single sequence MFE | -33.64 |
| Consensus MFE | -22.81 |
| Energy contribution | -25.45 |
| Covariance contribution | 2.64 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.68 |
| SVM decision value | 2.33 |
| SVM RNA-class probability | 0.992439 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
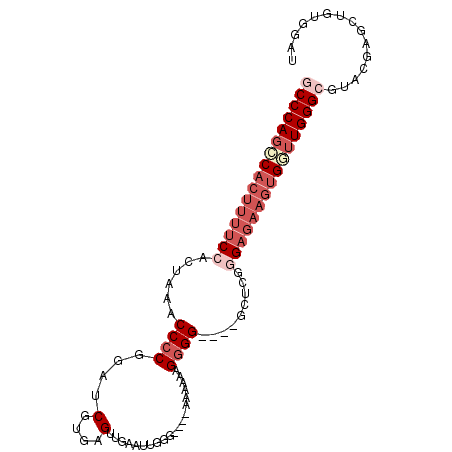
>2R_DroMel_CAF1 18200682 92 - 20766785 GCCCAGACACUUUUCCACUAAACCCCGGAUCGUGAGUUGAAUUGGG----AAAAAAGGGG----GCUCGCGAGAAGUGGUUGGGCAUACGAGCUGUGGAU ((((((.((((((((.......((((...((.(((......))).)----).....))))----......)))))))).))))))............... ( -29.82) >DroSec_CAF1 25780 91 - 1 GCCCAGCCACUUUUCCACUAAACCCCGGAUCGUGAGUUGAAUUGGG----AAAACAGGGG----GCUCGGGAG-AGUGGUUGGGCGUACGAGCUGUGGAU ((((((((((((((((......((((...((.(((......))).)----).....))))----....)))))-)))))))))))............... ( -39.30) >DroSim_CAF1 27500 92 - 1 GCCCAGCCACUUUUCCACUAAACCCCGGAUCGUGAGUUGAAUUGGA----AAAAAAGGGG----GCUCGGGAGAAGUGGUUGGGCGUACGAGCUGUGGAU ((((((((((((((((......((((....(....).....((...----.))...))))----.....))))))))))))))))............... ( -40.50) >DroEre_CAF1 26444 96 - 1 GCCCAGGCACUUUUCCACUAAACCACGGAUCGUGAGUUGAAUCCGG----AAAAAGGGGGAAUGGAUAGGGAGAAGUGCUUGGGAGUACGAGCUGUGGAU .(((((((((((((((.((((((((((...)))).)))...(((..----.....)))........))))))))))))))))))(((....)))...... ( -35.80) >DroYak_CAF1 27251 83 - 1 GCCCAUCCACUUUUCCACUAAACCCCGGAUCGUGAGUUGAAUCGGAAUAGAAAGAAGGGG----UAUAGUGUUAUAAGCUUGGGAGU------------- .((((....(((...(((((.(((((....(....).....((......)).....))))----).)))))....)))..))))...------------- ( -22.80) >consensus GCCCAGCCACUUUUCCACUAAACCCCGGAUCGUGAGUUGAAUUGGG____AAAAAAGGGG____GCUCGGGAGAAGUGGUUGGGCGUACGAGCUGUGGAU ((((((((((((((((......((((....(....)....................)))).........))))))))))))))))............... (-22.81 = -25.45 + 2.64)

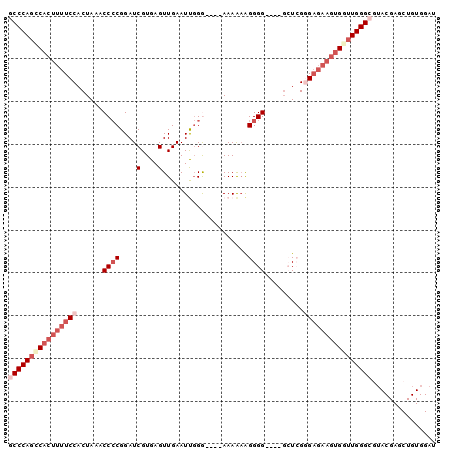
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:53:21 2006