
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,178,890 – 18,179,063 |
| Length | 173 |
| Max. P | 0.879709 |

| Location | 18,178,890 – 18,178,994 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 80.39 |
| Mean single sequence MFE | -27.38 |
| Consensus MFE | -15.64 |
| Energy contribution | -16.07 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.879709 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18178890 104 + 20766785 CAUGUUCCGAUUUUCGCUUUCUUGAUCAUGUGUCAUG-GAUCGGUUUGUUUAUCCUUCUACAAGGAUACACA------------CACAUCGGUAUUAACAAUAUUGGAAUAUAUCUG .((((((((((..............(((((...))))-)((((((.(((.(((((((....)))))))...)------------)).)))))).........))))))))))..... ( -24.20) >DroSec_CAF1 4215 104 + 1 CGUGUUCCGGUUUUCGCAUUCUCGAUCAUGUGUCAUG-GAGCGAUUUGUUUAUCCUUCUACAAGGAUACAAA------------UAAAUCAGUAUCAACAAUAUUGGAAUAUAUAUG .((((((((((...(((((........)))))...((-(.(((((((((((((((((....)))))))..))------------)))))).)).))).....))))))))))..... ( -25.70) >DroSim_CAF1 4220 104 + 1 CCUGUUCCGGUUUUCGCAUUCUCGAUCAUGUGUCAUG-GAGCGAUUUGUUUAUCCUUCCACAAGGAUACACA------------CAAAUCAGUAUCAACAAUAUUGGAAUAUAUCUG ((.((((((.....(((((........)))))...))-))))(((((((.(((((((....)))))))...)------------))))))...............)).......... ( -25.40) >DroYak_CAF1 4261 115 + 1 UGUGUUCCGGUUUUCGUAUUAUGGAUGCAG--UUGUGGCAGCAAUUUGUUUAUCCUUCUGCAAGGAUACACACUGAUUUAUGCACACAUUAGCAUCAACAAUAUUGGAAUAUAUGUG (((((((((......(((((...(((((..--.(((((((..(((((((.(((((((....))))))).)))..))))..))).))))...)))))...)))))))))))))).... ( -34.20) >consensus CGUGUUCCGGUUUUCGCAUUCUCGAUCAUGUGUCAUG_GAGCGAUUUGUUUAUCCUUCUACAAGGAUACACA____________CAAAUCAGUAUCAACAAUAUUGGAAUAUAUCUG .((((((((((..((((......(((.....)))......))))..(((.(((((((....))))))).)))..............................))))))))))..... (-15.64 = -16.07 + 0.44)

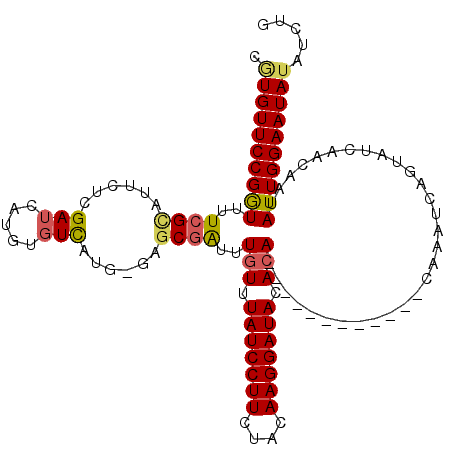

| Location | 18,178,961 – 18,179,063 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 71.26 |
| Mean single sequence MFE | -21.93 |
| Consensus MFE | -9.14 |
| Energy contribution | -11.20 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.42 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 18178961 102 - 20766785 GAUACCAUCGUUGGCCAUCUAAUCAACAGUCAACAGUCUUUUUUUGACUAUUAC---UAU-------CUACAGUUCAACCAGAUAUAUUCCAAUAUUGUUAAUACCGAUGUG------- .....((((((((((.............))))))((((.......)))).....---(((-------..(((((.......((.....))....)))))..)))..))))..------- ( -13.32) >DroSec_CAF1 4286 93 - 1 GAUACCAUCGUUGGCCAUCUAAUU--UGGUCAACAGUCUUUGU-----------------CUAUUGACUACAGUUCAGCCAUAUAUAUUCCAAUAUUGUUGAUACUGAUUUA------- .........((((((((.......--))))))))((((.....-----------------.....)))).(((((((((.((((........)))).))))).)))).....------- ( -24.50) >DroSim_CAF1 4291 107 - 1 GAUACCAUCGUUGGCCAUCUAAUG--UGGUCAACAGUCUUUGUUUGACUAUUAA---UAUCUAUUGACUACAGUUCAACCAGAUAUAUUCCAAUAUUGUUGAUACUGAUUUG------- ((((.....(((((((((.....)--))))))))((((.......)))).....---)))).........(((((((((..(((((......)))))))))).)))).....------- ( -26.70) >DroEre_CAF1 4208 114 - 1 GAUACCAUCGAUGACUAUC--AUUAGAAGCCAACAGUAUUUGUUUGACUAUUACUUAUAUCUAUCAAAUACAAAUCAGCCUGAUAUAUCCCAAU---GCUGAUGCUGAUGUGUGCGUAA (.(((((((((((.....)--)))...(((.....(((((((...((.(((.....)))))...)))))))..((((((.((........))..---))))))))))))).)))).... ( -23.20) >consensus GAUACCAUCGUUGGCCAUCUAAUU__UAGUCAACAGUCUUUGUUUGACUAUUAC___UAUCUAUUGACUACAGUUCAACCAGAUAUAUUCCAAUAUUGUUGAUACUGAUGUG_______ .........(((((((...........)))))))....................................(((((((((..(((((......)))))))))).))))............ ( -9.14 = -11.20 + 2.06)

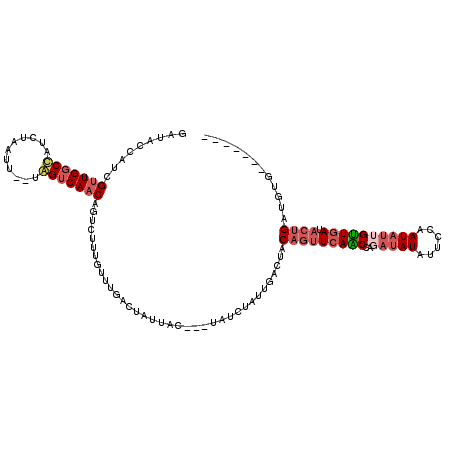
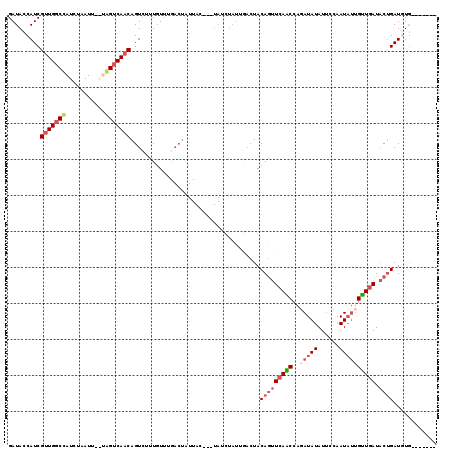
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:53:03 2006