
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,115,276 – 18,115,382 |
| Length | 106 |
| Max. P | 0.577790 |

| Location | 18,115,276 – 18,115,382 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 79.33 |
| Mean single sequence MFE | -22.00 |
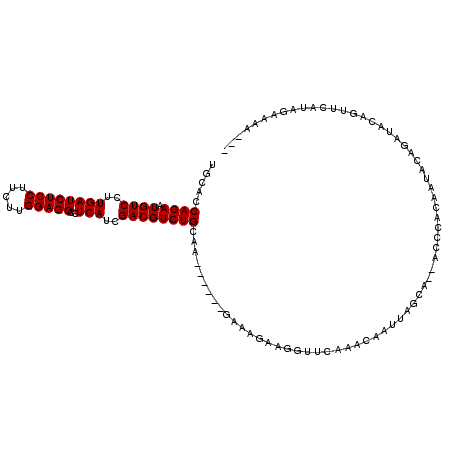
| Consensus MFE | -16.20 |
| Energy contribution | -16.20 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.577790 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
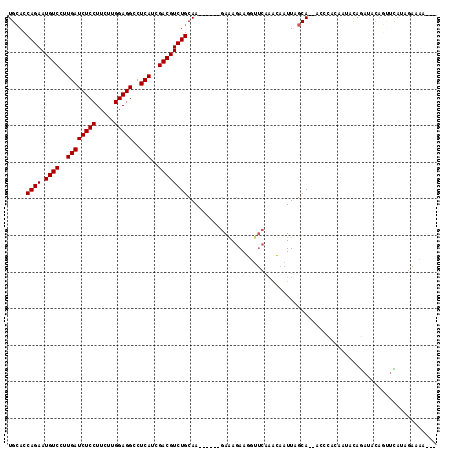
>2R_DroMel_CAF1 18115276 106 - 20766785 UGCACCAGAAUGUCCUUGAUCUCCUUCUUGGAGGCCUCAUCGACGUCUGCAA------GAAAGAAGUUUCAAACAAUUAGCA--ACCCACAACACAGAUACAGUUCAUAGAAAA--- (((..((((.((((..((((((((.....)))))..)))..))))))))...------((((....)))).........)))--..............................--- ( -20.00) >DroGri_CAF1 819 102 - 1 UGGACCAGAAUGUCCUUGAUCUCCUUCUUGGAGGCCUCAUCGACGUCUGCAAUGGGAUAAAAUAAG-U----ACAAUUAGCACGAGUUACUUUCUAAAUUCAUCUUA---------- .....((((.((((..((((((((.....)))))..)))..))))))))...((((((.......(-(----.......))..(((((........)))))))))))---------- ( -20.20) >DroSec_CAF1 1208 106 - 1 UGCACCAGAAUGUCCUUGAUCUCCUUCUUGGAGGCCUCAUCGACGUCUGCAA------GAAAGAAGUUUCAAACAAUUAGCA--ACCCACAAUGCAGAUACAGUUCAUAGAAAA--- .......(((((((..((((((((.....)))))..)))..)))(((((((.------((((....)))).........(..--.....)..)))))))...))))........--- ( -21.90) >DroSim_CAF1 904 106 - 1 UGCACCAGAAUGUCCUUGAUCUCCUUCUUGGAGGCCUCAUCGACGUCUGCAA------GAAAGAAGAUUCAAACAAUUAGCA--ACCUACAAUACAGAUACAGUAUAUAGAAAA--- (((..((((.((((..((((((((.....)))))..)))..))))))))...------(((......))).........)))--..(((..((((.......)))).)))....--- ( -20.80) >DroEre_CAF1 1270 109 - 1 UGCACCAGAAUGUCCUUGAUCUCCUUCUUGGAGGCCUCAUCGACGUCUGCAA------GAAAGAAUGAUCAAGGAAUUAGCA--ACCCGCAAUUUAGAAACAGUUCAUAGAAUACAU .......((((.(((((((((((.((((((.((((.((...)).)))).)))------))).))..)))))))))....((.--....))............))))........... ( -30.00) >DroYak_CAF1 1226 91 - 1 UGCACCAGAAUGUCCUUGAUCUCCUUCUUGGAGGCCUCAUCGACGUCUGCAA------GAGAGAAUGUUCAAAGAAUUAGCA--A-------UAUAG--------CAUAGAACG--- (((..((((.((((..((((((((.....)))))..)))..))))))))...------(((......))).........)))--.-------.....--------.........--- ( -19.10) >consensus UGCACCAGAAUGUCCUUGAUCUCCUUCUUGGAGGCCUCAUCGACGUCUGCAA______GAAAGAAGGUUCAAACAAUUAGCA__ACCCACAAUACAGAUACAGUUCAUAGAAAA___ .....((((.((((..((((((((.....)))))..)))..)))))))).................................................................... (-16.20 = -16.20 + -0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:52:36 2006