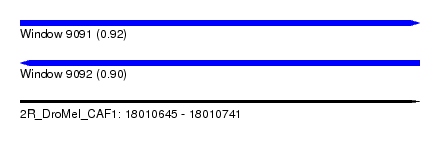
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 18,010,645 – 18,010,741 |
| Length | 96 |
| Max. P | 0.920554 |

| Location | 18,010,645 – 18,010,741 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 86.47 |
| Mean single sequence MFE | -23.20 |
| Consensus MFE | -17.18 |
| Energy contribution | -18.18 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.920554 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
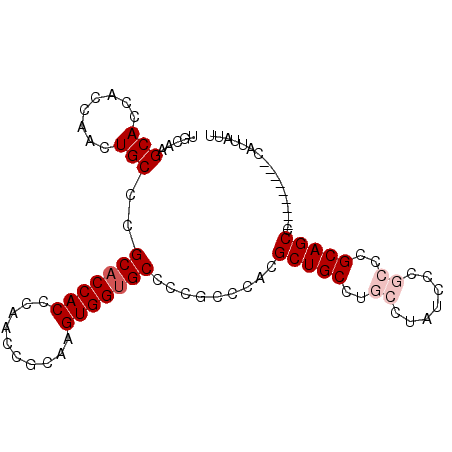
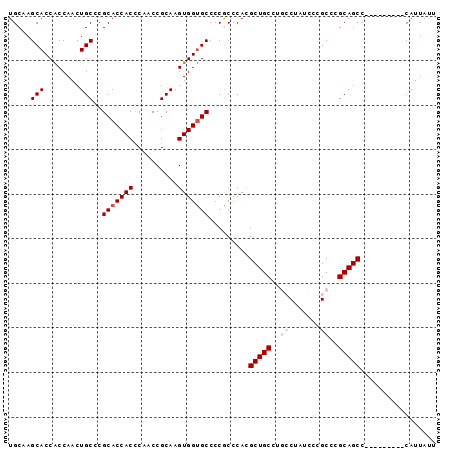
>2R_DroMel_CAF1 18010645 96 + 20766785 UGCCAGCACCACCAACUGCCCGCACCACCCAACCGCAAGUGGUGCCCCGCCCACGCUGCCUGCCUAGCCCGCCCGCAGCCUUAUUGCAGCAUUAUU (((..(((.........((..(((((((..........)))))))...))....(((((..((.......))..))))).....))).)))..... ( -25.30) >DroSec_CAF1 24719 87 + 1 UGCAAGCACCGCCAACUGCCCGCACCACCCAACCGCAAGUGGUGCCCCGCCCACGCUGCCUGCCUAUCCCGCCCGCAGCC---------CAUUAUU .((..(((........)))..(((((((..........)))))))...))....(((((..((.......))..))))).---------....... ( -22.60) >DroSim_CAF1 23688 87 + 1 UGCAAGCACCGCCAACUGCCCGCACCACCCAACCGCAAGUGGCGCCCCGCCCACGCUGCCUGCCUAUCCCGCCCGCAGCC---------CAUUAUU (((..(((........)))..)))..............((((.((...))))))(((((..((.......))..))))).---------....... ( -19.80) >DroEre_CAF1 26094 83 + 1 UGCUAGCACCACCAACUGCACGCACCACUCAACCGCAAGUGGUGCCCCACCCAUGCUGC----CGCUCCCGCCCGCAGCC---------CAUUAUU .....(((........)))..((((((((........)))))))).........(((((----.((....))..))))).---------....... ( -23.60) >DroYak_CAF1 23626 83 + 1 UGCCAGCACCACCAACUGCACGCACCACUCAACCGCAAGUGGUGCCCCACCCAGGCUGC----CUUUUCCGCCCGCAGCC---------CAUUAUU .....(((........)))..((((((((........))))))))........((((((----...........))))))---------....... ( -24.70) >consensus UGCAAGCACCACCAACUGCCCGCACCACCCAACCGCAAGUGGUGCCCCGCCCACGCUGCCUGCCUAUCCCGCCCGCAGCC_________CAUUAUU .....(((........)))..(((((((..........))))))).........(((((..((.......))..)))))................. (-17.18 = -18.18 + 1.00)

| Location | 18,010,645 – 18,010,741 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 86.47 |
| Mean single sequence MFE | -35.78 |
| Consensus MFE | -27.38 |
| Energy contribution | -27.62 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.903316 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
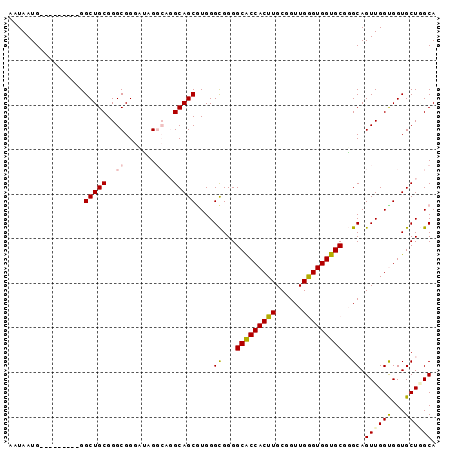
>2R_DroMel_CAF1 18010645 96 - 20766785 AAUAAUGCUGCAAUAAGGCUGCGGGCGGGCUAGGCAGGCAGCGUGGGCGGGGCACCACUUGCGGUUGGGUGGUGCGGGCAGUUGGUGGUGCUGGCA .....(((((((....((((((...(.(.(((.((.....)).))).).).(((((((((......)))))))))..)))))).....))).)))) ( -39.40) >DroSec_CAF1 24719 87 - 1 AAUAAUG---------GGCUGCGGGCGGGAUAGGCAGGCAGCGUGGGCGGGGCACCACUUGCGGUUGGGUGGUGCGGGCAGUUGGCGGUGCUUGCA .......---------..(((....))).....(((((((.(((.(((...(((((((((......))))))))).....))).))).))))))). ( -37.50) >DroSim_CAF1 23688 87 - 1 AAUAAUG---------GGCUGCGGGCGGGAUAGGCAGGCAGCGUGGGCGGGGCGCCACUUGCGGUUGGGUGGUGCGGGCAGUUGGCGGUGCUUGCA .......---------..(((....))).....(((((((.(((.(((...(((((((((......))))))))).....))).))).))))))). ( -37.10) >DroEre_CAF1 26094 83 - 1 AAUAAUG---------GGCUGCGGGCGGGAGCG----GCAGCAUGGGUGGGGCACCACUUGCGGUUGAGUGGUGCGUGCAGUUGGUGGUGCUAGCA .......---------.(((....((....))(----(((.(((.((((.((((((((((......))))))))).).)).)).))).))))))). ( -32.70) >DroYak_CAF1 23626 83 - 1 AAUAAUG---------GGCUGCGGGCGGAAAAG----GCAGCCUGGGUGGGGCACCACUUGCGGUUGAGUGGUGCGUGCAGUUGGUGGUGCUGGCA ......(---------((((((...........----)))))))...((.((((((((((......))))))))).).))((..(.....)..)). ( -32.20) >consensus AAUAAUG_________GGCUGCGGGCGGGAUAGGCAGGCAGCGUGGGCGGGGCACCACUUGCGGUUGGGUGGUGCGGGCAGUUGGUGGUGCUGGCA .................(((((..((.......))..)))))....((...(((((((((......)))))))))..)).((((((...)))))). (-27.38 = -27.62 + 0.24)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:51:34 2006