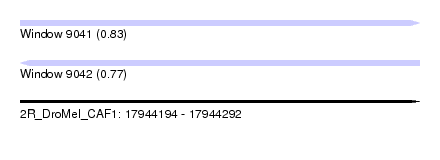

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,944,194 – 17,944,292 |
| Length | 98 |
| Max. P | 0.829718 |

| Location | 17,944,194 – 17,944,292 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 93.56 |
| Mean single sequence MFE | -27.68 |
| Consensus MFE | -23.67 |
| Energy contribution | -23.48 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.829718 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
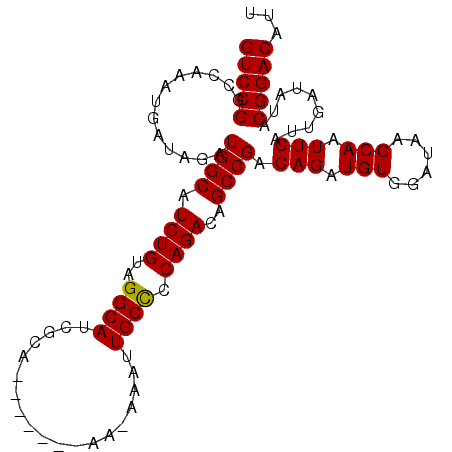
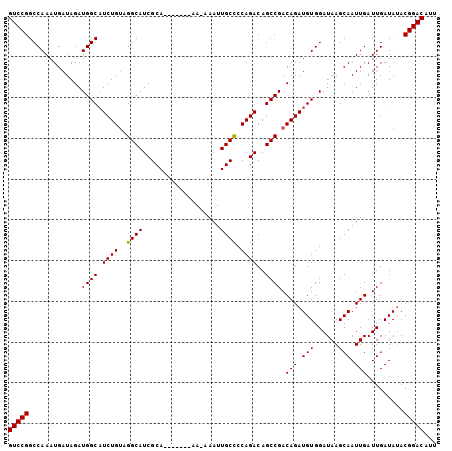
>2R_DroMel_CAF1 17944194 98 + 20766785 GUCCGGCCAAAUGAUAGAUGGCAUCUGUAGGCAUCGCA-------AAAAAAUUGCCCCAGACAGCCGACAGAUGUGGAUAAGCAAUUGAUUGAUAUACGGACAGU (((((.............(.((((((((.(((.(((((-------(.....))))....))..))).)))))))).).(((....))).........)))))... ( -26.80) >DroSec_CAF1 1569 97 + 1 GUCCGGCCAAAUGAUAGAUGGCAUCUGUAGGCAUCGCA-------AA-AAAUUGCUCCAGACAGCCGACAGAUGUGGAUAAGCAAUUGAUUGAUACACGGACAUU (((((.............(.((((((((.(((.(((((-------(.-...))))....))..))).)))))))).).(((....))).........)))))... ( -26.80) >DroEre_CAF1 6325 97 + 1 GUCCGGCCAAAUGAUAGAUGGCUUCUGUAGGCAUCGCA-------AA-AAAUUGCCCCAGACAGCCGACAGAUGUGGAUAAGCAAUUGAUUGAUAUACGGACAUU (((((.............(((((((((..((((.....-------..-....)))).)))).))))).(((.(((......))).))).........)))))... ( -27.10) >DroYak_CAF1 6113 104 + 1 GUCCGGCCAAAUGAUAGAUGGCAUCUGAAGGCAUCGCAAAAGGCAAA-AAAUUGCCCCAGACAGCCGACAGAUGUGGAUAAGCAAUUGAUUGAUAUACGGACAUU (((((.............(.(((((((..(((.((......(((((.-...)))))...))..)))..))))))).).(((....))).........)))))... ( -30.00) >consensus GUCCGGCCAAAUGAUAGAUGGCAUCUGUAGGCAUCGCA_______AA_AAAUUGCCCCAGACAGCCGACAGAUGUGGAUAAGCAAUUGAUUGAUAUACGGACAUU (((((.............((((.((((..((((...................)))).))))..)))).(((.(((......))).))).........)))))... (-23.67 = -23.48 + -0.19)

| Location | 17,944,194 – 17,944,292 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 93.56 |
| Mean single sequence MFE | -27.80 |
| Consensus MFE | -21.89 |
| Energy contribution | -22.39 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.766236 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
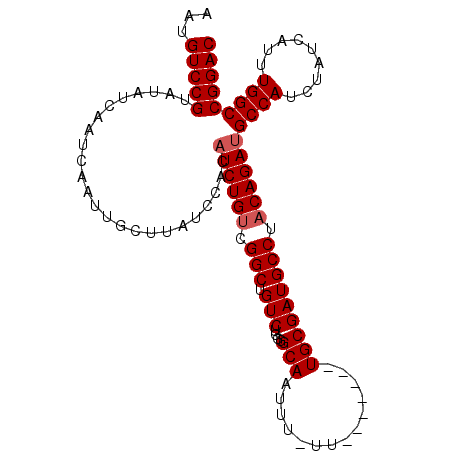
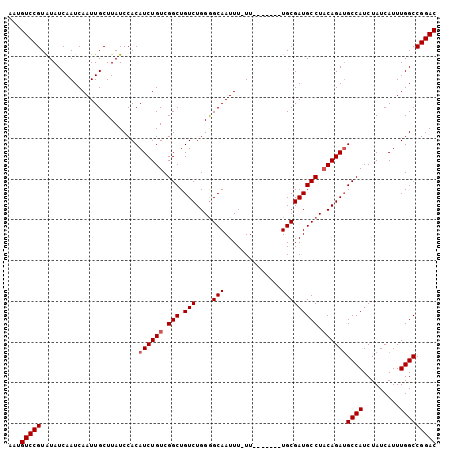
>2R_DroMel_CAF1 17944194 98 - 20766785 ACUGUCCGUAUAUCAAUCAAUUGCUUAUCCACAUCUGUCGGCUGUCUGGGGCAAUUUUUU-------UGCGAUGCCUACAGAUGCCAUCUAUCAUUUGGCCGGAC ...(((((........................((((((.(((.(((....((((.....)-------))))))))).))))))((((.........))))))))) ( -26.60) >DroSec_CAF1 1569 97 - 1 AAUGUCCGUGUAUCAAUCAAUUGCUUAUCCACAUCUGUCGGCUGUCUGGAGCAAUUU-UU-------UGCGAUGCCUACAGAUGCCAUCUAUCAUUUGGCCGGAC ...(((((........................((((((.(((.(((....((((...-.)-------))))))))).))))))((((.........))))))))) ( -26.60) >DroEre_CAF1 6325 97 - 1 AAUGUCCGUAUAUCAAUCAAUUGCUUAUCCACAUCUGUCGGCUGUCUGGGGCAAUUU-UU-------UGCGAUGCCUACAGAAGCCAUCUAUCAUUUGGCCGGAC ...(((((.........................(((((.(((.(((....((((...-.)-------))))))))).))))).((((.........))))))))) ( -25.80) >DroYak_CAF1 6113 104 - 1 AAUGUCCGUAUAUCAAUCAAUUGCUUAUCCACAUCUGUCGGCUGUCUGGGGCAAUUU-UUUGCCUUUUGCGAUGCCUUCAGAUGCCAUCUAUCAUUUGGCCGGAC ...(((((........................(((((..((((((..(((((((...-.)))))))..)))..)))..)))))((((.........))))))))) ( -32.20) >consensus AAUGUCCGUAUAUCAAUCAAUUGCUUAUCCACAUCUGUCGGCUGUCUGGGGCAAUUU_UU_______UGCGAUGCCUACAGAUGCCAUCUAUCAUUUGGCCGGAC ...(((((........................((((((.(((.(((....(((..............))))))))).))))))((((.........))))))))) (-21.89 = -22.39 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:50:45 2006