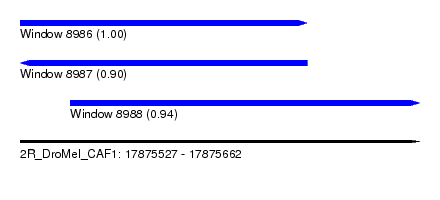

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,875,527 – 17,875,662 |
| Length | 135 |
| Max. P | 0.998535 |

| Location | 17,875,527 – 17,875,624 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 96.49 |
| Mean single sequence MFE | -31.08 |
| Consensus MFE | -27.34 |
| Energy contribution | -27.54 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.88 |
| SVM decision value | 3.13 |
| SVM RNA-class probability | 0.998535 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17875527 97 + 20766785 UCGCCCAUUUGCUGACCGCAUUCACGGAGCUGGCCUCCAUUUGGCCUGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAAGCAAAGCC ..((...(((((((((((......)))(((.((((.......)))).)))((((((((........))))))))...........)).)))))))). ( -30.70) >DroSec_CAF1 40395 97 + 1 UCGCCCAUUUGCUGACCGCAUUUACGGAGCUGGCCUCCAUUUGGCCAGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAAGCAAAGCC ..((...(((((((((((......)))((((((((.......))))))))((((((((........))))))))...........)).)))))))). ( -35.30) >DroSim_CAF1 40853 97 + 1 UCGCCCAUUUGCUGACCGCAUUCACGGAGCUGGCCUCCAUUUGGCCUGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAAGCAAAGCC ..((...(((((((((((......)))(((.((((.......)))).)))((((((((........))))))))...........)).)))))))). ( -30.70) >DroEre_CAF1 42785 97 + 1 UCGCCCAUUUGCUGACCGCAUUCACGGAGCUGGCCUCCAUUUGGCCUGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAGGCCAAGCC ..(((.....(((..(((......)))))).))).....(((((((((...(((((((........)))))))(((......))).))))))))).. ( -31.50) >DroYak_CAF1 39548 97 + 1 UCGCCCAUUUGCUGACCGCAUUCACGAAGCUGGCCUCCAUUUGGCCUGCUUUGUCCGCAAUUAAAGCCAGGCAGUUCAAAGAAACUCAGGCAAAGCC ..(((.....((((.((......(((((((.((((.......)))).)))))))..((.......))..)))))).....(.....).)))...... ( -27.20) >consensus UCGCCCAUUUGCUGACCGCAUUCACGGAGCUGGCCUCCAUUUGGCCUGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAAGCAAAGCC ..((...((((((....((....(((((((.((((.......)))).)))))))((((........))))))((((......))))..)))))))). (-27.34 = -27.54 + 0.20)


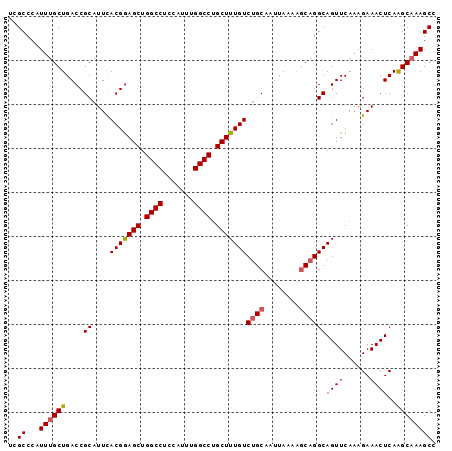
| Location | 17,875,527 – 17,875,624 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 96.49 |
| Mean single sequence MFE | -32.74 |
| Consensus MFE | -28.94 |
| Energy contribution | -29.02 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.901861 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17875527 97 - 20766785 GGCUUUGCUUGAGUUUCUUUGAACUGCCUGCUUUUAAUUGCAGACAAAGCAGGCCAAAUGGAGGCCAGCUCCGUGAAUGCGGUCAGCAAAUGGGCGA ((((((..((((((((....)))))(((((((((...........))))))))))))...)))))).((.((((...(((.....))).)))))).. ( -32.50) >DroSec_CAF1 40395 97 - 1 GGCUUUGCUUGAGUUUCUUUGAACUGCCUGCUUUUAAUUGCAGACAAAGCUGGCCAAAUGGAGGCCAGCUCCGUAAAUGCGGUCAGCAAAUGGGCGA .((((((((.((............((.((((........)))).)).((((((((.......))))))))((((....)))))))))))...))).. ( -32.80) >DroSim_CAF1 40853 97 - 1 GGCUUUGCUUGAGUUUCUUUGAACUGCCUGCUUUUAAUUGCAGACAAAGCAGGCCAAAUGGAGGCCAGCUCCGUGAAUGCGGUCAGCAAAUGGGCGA ((((((..((((((((....)))))(((((((((...........))))))))))))...)))))).((.((((...(((.....))).)))))).. ( -32.50) >DroEre_CAF1 42785 97 - 1 GGCUUGGCCUGAGUUUCUUUGAACUGCCUGCUUUUAAUUGCAGACAAAGCAGGCCAAAUGGAGGCCAGCUCCGUGAAUGCGGUCAGCAAAUGGGCGA ((((((((((((((((....)))))..((((........))))......)))))))......)))).((.((((...(((.....))).)))))).. ( -35.00) >DroYak_CAF1 39548 97 - 1 GGCUUUGCCUGAGUUUCUUUGAACUGCCUGGCUUUAAUUGCGGACAAAGCAGGCCAAAUGGAGGCCAGCUUCGUGAAUGCGGUCAGCAAAUGGGCGA ....((((((.(((((....)))))((.(((((.((.(..(((....(((.((((.......)))).))))))..).)).)))))))....)))))) ( -30.90) >consensus GGCUUUGCUUGAGUUUCUUUGAACUGCCUGCUUUUAAUUGCAGACAAAGCAGGCCAAAUGGAGGCCAGCUCCGUGAAUGCGGUCAGCAAAUGGGCGA .((((((....(((((....)))))..((((........)))).)))))).((((.......)))).((.((((...(((.....))).)))))).. (-28.94 = -29.02 + 0.08)


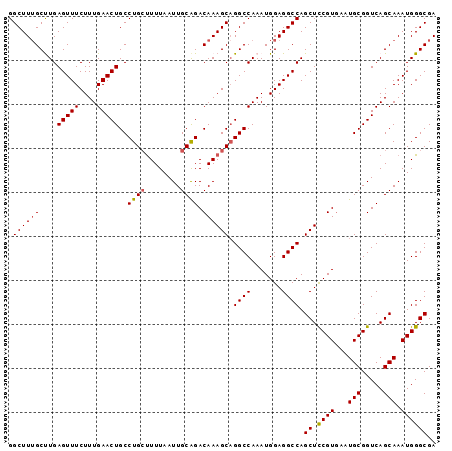
| Location | 17,875,544 – 17,875,662 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.79 |
| Mean single sequence MFE | -33.74 |
| Consensus MFE | -24.40 |
| Energy contribution | -24.84 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.68 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.31 |
| SVM RNA-class probability | 0.940516 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17875544 118 + 20766785 GCAUUCACGGAGCUGGCCUCCAUUUGGCCUGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAAGCAAAGCCAAGUACACUGGGGAAA--AAACCCUCCCAAUGCUGCUAAG (((((...((((.....))))(((((((.((((((((((((........)))))))(((......)))..)))))..)))))))......((((..--......)))))))))....... ( -36.40) >DroSec_CAF1 40412 107 + 1 GCAUUUACGGAGCUGGCCUCCAUUUGGCCAGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAAGCAAAGCCAAGUACACUGGGGAAA--AAACCAUCCCA----------- ((......((((((((((.......))))))))))((((((........)))))).................))................((((..--......)))).----------- ( -35.10) >DroSim_CAF1 40870 109 + 1 GCAUUCACGGAGCUGGCCUCCAUUUGGCCUGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAAGCAAAGCCAAGUACACUGGGGAAAAAAAACCCUCCCA----------- ........((((.....))))(((((((.((((((((((((........)))))))(((......)))..)))))..)))))))......((((..........)))).----------- ( -31.90) >DroEre_CAF1 42802 107 + 1 GCAUUCACGGAGCUGGCCUCCAUUUGGCCUGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAGGCCAAGCCGAGUACACUGGGGAAA--AUACUGGCUCA----------- .........((((..(......(((((((((...(((((((........)))))))(((......))).)))))))))((.((....)).))....--...)..)))).----------- ( -37.70) >DroYak_CAF1 39565 102 + 1 GCAUUCACGAAGCUGGCCUCCAUUUGGCCUGCUUUGUCCGCAAUUAAAGCCAGGCAGUUCAAAGAAACUCAGGCAAAGCCAAGUACACUGUGGAAA--AU-----CUCA----------- .......((((((.((((.......)))).))))))(((((.......(((..(.((((......))))).))).......((....)))))))..--..-----....----------- ( -27.60) >consensus GCAUUCACGGAGCUGGCCUCCAUUUGGCCUGCUUUGUCUGCAAUUAAAAGCAGGCAGUUCAAAGAAACUCAAGCAAAGCCAAGUACACUGGGGAAA__AAACCCUCCCA___________ ((......(((((.((((.......)))).)))))((((((........)))))).................))....((.((....)).))............................ (-24.40 = -24.84 + 0.44)


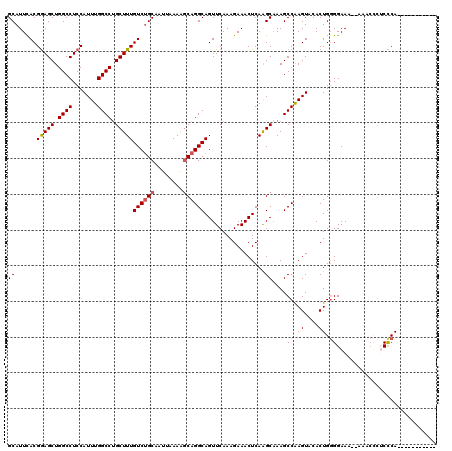
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:49:53 2006