
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,829,411 – 17,829,502 |
| Length | 91 |
| Max. P | 0.530380 |

| Location | 17,829,411 – 17,829,502 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.23 |
| Mean single sequence MFE | -22.73 |
| Consensus MFE | -8.64 |
| Energy contribution | -11.09 |
| Covariance contribution | 2.45 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.38 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.530380 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17829411 91 + 20766785 ----------------CUAGACACUACAAAGUGUCAUUGCGAUUAAACU--AUUUACGUAAACA---UUAACGACUUUAUUUUGAUUGUAAUAGAGCACAUUCGGUGC--------GAAG ----------------...((((((....))))))((((((((((((.(--(....(((.....---...)))....)).))))))))))))...((((.....))))--------.... ( -23.90) >DroSec_CAF1 97399 91 + 1 ----------------CUAGACACUACAAAGUGUCAUUGCGAUUAAACU--AUUUACGUAAACA---UUAACGACUUUAUUUUGAUUGUAAUAGAGCACAUUCGGUGC--------GCAG ----------------...((((((....))))))((((((((((((.(--(....(((.....---...)))....)).)))))))))))).(.((((.....))))--------.).. ( -24.00) >DroSim_CAF1 96723 91 + 1 ----------------CUAGACACUACAAAGUGUCAUUGCGAUUAAACU--AUUUACGUAAACA---UUAACGACUUUAUUUUGAUUGUAAUAGAGCACAUUCGGUGC--------GCAG ----------------...((((((....))))))((((((((((((.(--(....(((.....---...)))....)).)))))))))))).(.((((.....))))--------.).. ( -24.00) >DroEre_CAF1 97907 91 + 1 ----------------CUAGGCACUACAACGAGUCAUAGCGAUUAAACU--AUUUACGUACAGA---CUAACGACUUUAUUUUGAUUGUAAUAGAGCACUUUGUGUGC--------GCAG ----------------....((((.((((.(.((..(((((((((((.(--(....(((.....---...)))....)).)))))))))..))..)).).))))))))--------.... ( -17.50) >DroYak_CAF1 101753 91 + 1 ----------------CUAAACACUACAAAGUGUCAUUGCGAUUAAACU--GUUUACGUAAACA---UUAACGACUUUAUUUUGAUUGUAAUAGAGCACAUUCGGUGC--------GAAG ----------------....(((((....))))).((((((((((((.(--(....(((.....---...)))....)).))))))))))))...((((.....))))--------.... ( -19.20) >DroPer_CAF1 108568 120 + 1 UAAUCCACACAAGUUUCUAGACUGUAGAAAGUGUCAUAUUGAUUAAUCUGUAUUUAAUUAUAUAUAGCUAAUCAGUUUAGUUGGACUUCAGCAGACAGCGGGCGAGUCGGAAACUAUUUG ...........(((((((.((((((.....(((((..((((((((..((((((........)))))).))))))))...((((.....)))).)))).)..)).)))))))))))..... ( -27.80) >consensus ________________CUAGACACUACAAAGUGUCAUUGCGAUUAAACU__AUUUACGUAAACA___UUAACGACUUUAUUUUGAUUGUAAUAGAGCACAUUCGGUGC________GCAG ...................((((((....))))))((((((((((((.........(((...........))).......))))))))))))...((((.....))))............ ( -8.64 = -11.09 + 2.45)

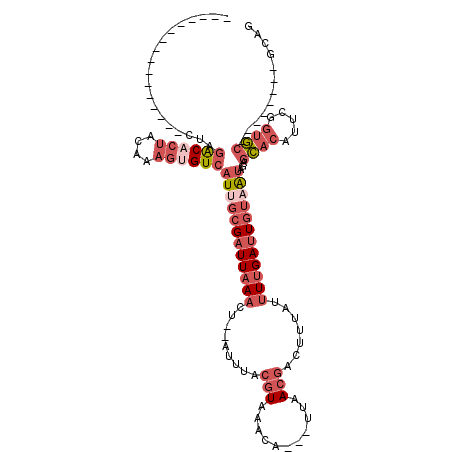

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:48:58 2006