
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,668,074 – 17,668,308 |
| Length | 234 |
| Max. P | 0.552107 |

| Location | 17,668,074 – 17,668,190 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.19 |
| Mean single sequence MFE | -38.85 |
| Consensus MFE | -34.46 |
| Energy contribution | -34.27 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.552107 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17668074 116 - 20766785 AUGCCGUACAUAUAGAAGGCGCUCGUAUAUAUAUCUUCGAUGUGGCCUAUCUACCAUGCGCCUACAUAUCGCAGU----GUGUGUGCGAGUGAGUGUGUCUACUACUGGCAAGUCCGCAU .(((((((....((((...((((((((((((((.((.(((((((((.(((.....))).)))...)))))).)))----)))))))))))))......)))).))).))))......... ( -37.90) >DroSec_CAF1 21491 116 - 1 AUGCCGUACAUAUAGAAGGCGCUCGUAUAUAUAUCUUCGAUGUGGCCUAUCUACCGUGCGCCUACAUAUCGCAGU----GUGUGUGCGAGUGAGUGUGUCUACUACUGGCAAGUCCGCAU .(((((((....((((...((((((((((((((.((.(((((((((.(((.....))).)))...)))))).)))----)))))))))))))......)))).))).))))......... ( -38.20) >DroEre_CAF1 21904 118 - 1 AUGCCGUACAUAUAGCAGGCGCUCGUAUAUAUAUCUUCGAUGUGGCCUAUCUACCGUGCGCCUACAUAUCGCAGU--GUGUGUGUGCGAGUGAGUGUGUCUACUACUGGCAAGUCCGCAU .(((((((......(((..(((((((((((((((((.(((((((((.(((.....))).)))...)))))).)).--)))))))))))))))..)))......))).))))......... ( -41.80) >DroYak_CAF1 33633 120 - 1 AUGCCGUACAUAUAGAAGGCGCUCGUAUAUAUAUCUUCGAUGUGGCCUAUCUACCGUGCGCCUACAUAUCGCAGUAUGUGUGUGUGCGAGUGAGUGUGUCUACUACUGGCAAGUCCGCAU .(((((((....((((...(((((((((((((((((.(((((((((.(((.....))).)))...)))))).))...)))))))))))))))......)))).))).))))......... ( -37.50) >consensus AUGCCGUACAUAUAGAAGGCGCUCGUAUAUAUAUCUUCGAUGUGGCCUAUCUACCGUGCGCCUACAUAUCGCAGU____GUGUGUGCGAGUGAGUGUGUCUACUACUGGCAAGUCCGCAU .(((((((.(((((.....(((((((((((((..((.(((((((((.(((.....))).)))...)))))).)).....)))))))))))))..))))).)))....))))......... (-34.46 = -34.27 + -0.19)


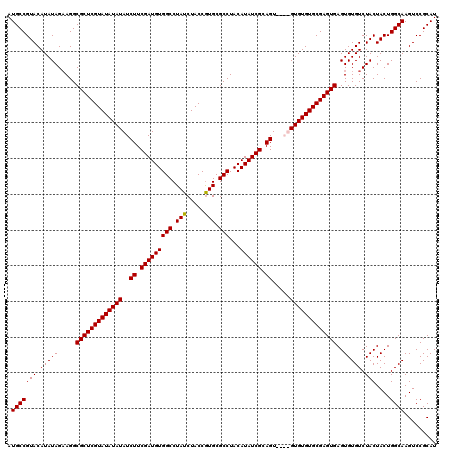
| Location | 17,668,190 – 17,668,308 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.51 |
| Mean single sequence MFE | -29.05 |
| Consensus MFE | -27.70 |
| Energy contribution | -27.95 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.95 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.517058 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
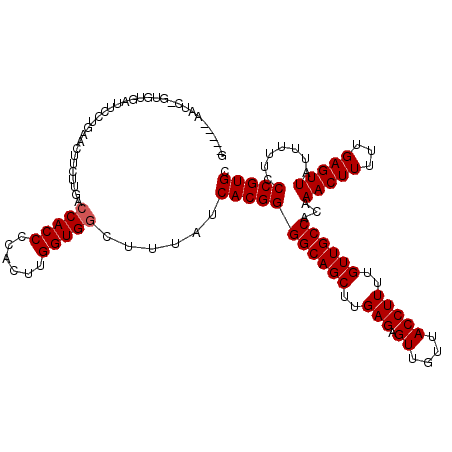
>2R_DroMel_CAF1 17668190 118 - 20766785 ACGU--GAUCCGUGUGAUUCCUGAACUUCUUGAGCACCCCACUUGGUGGCUUUAUCACGGGGCAGCUUGAGAGUUGUUACCUUUUGUUGCCACAAACUUUUUGAGUUAUUUUUCCCGUGC ..(.--.(((.....)))..)..........(((((((......))).))))...(((((((((((..(((.((....)))))..))))))...(((((...))))).......))))). ( -28.80) >DroSec_CAF1 21607 97 - 1 -----------------------AACUUCUUGACCACCCCACUUGGUGGCUUUAUCACGGGGCAGCUUGAGAGUUGUUACCUUUUGUUGCCACAAACUUUUUGAGUUAUUUUUCCCGUGC -----------------------..........(((((......)))))......(((((((((((..(((.((....)))))..))))))...(((((...))))).......))))). ( -29.10) >DroEre_CAF1 22022 115 - 1 G-----AACCAAUGUGCUAUCCGAACUUCUUGACCACCCCACUUGGUGGCUUUAUCACGGGGCAGCUUGAGAGUUGUUACCUUUUGUUGCCACAAACUUUUUGAGUUAUUUUUCCCGUGC .-----...........................(((((......)))))......(((((((((((..(((.((....)))))..))))))...(((((...))))).......))))). ( -29.10) >DroYak_CAF1 33753 120 - 1 GUGUGUAAUCUGUGUAAUUCCUGAACUUCUUGACCACCCCACGCGGUGGCUUUAUCACGGGGCAGCUUGAGAGUUGUUACCUUUUGUUGCCACAAACUUUUUGAGUUAUUUUUCCCGUGC .................................(((((......)))))......(((((((((((..(((.((....)))))..))))))...(((((...))))).......))))). ( -29.20) >consensus G_____AAUC_GUGUGAUUCCUGAACUUCUUGACCACCCCACUUGGUGGCUUUAUCACGGGGCAGCUUGAGAGUUGUUACCUUUUGUUGCCACAAACUUUUUGAGUUAUUUUUCCCGUGC .................................(((((......)))))......(((((((((((..(((.((....)))))..))))))...(((((...))))).......))))). (-27.70 = -27.95 + 0.25)

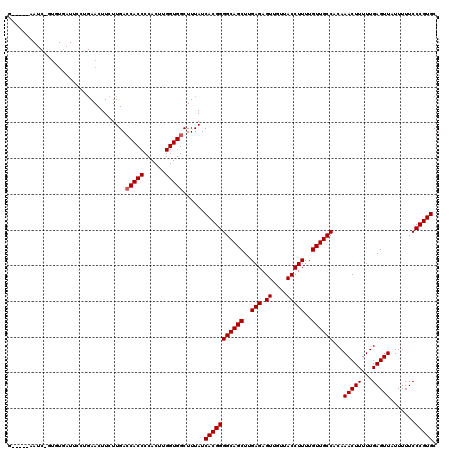
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:48:09 2006