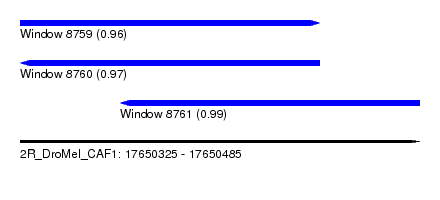

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,650,325 – 17,650,485 |
| Length | 160 |
| Max. P | 0.990407 |

| Location | 17,650,325 – 17,650,445 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -33.48 |
| Consensus MFE | -31.22 |
| Energy contribution | -31.82 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.49 |
| SVM RNA-class probability | 0.958827 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
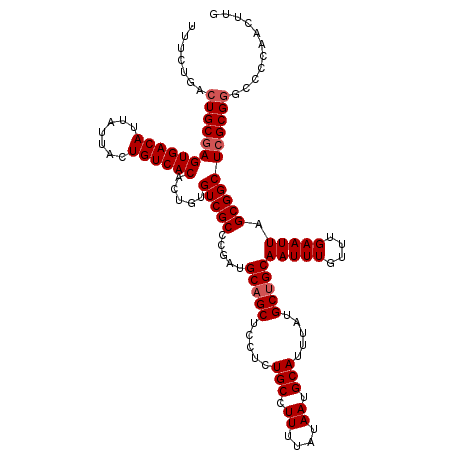
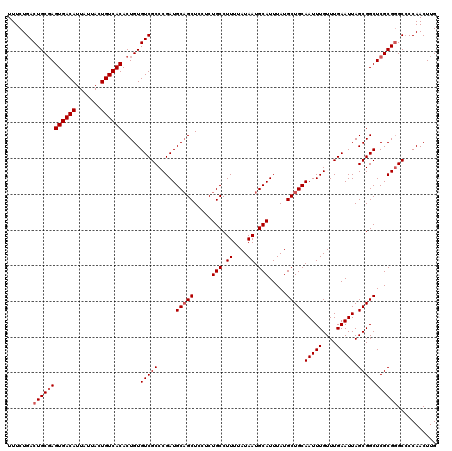
>2R_DroMel_CAF1 17650325 120 + 20766785 UUUCUGACUGCGAGUGACAUUAUUACUGUCACACUGUGUCGCCCGAUGCAGCUUCUCUGCCUUUUAUAAUGCAUUUAUGCUGCAAUUUGUUUGAAUUAGCGGCUCGCGGGCACCAACUUG .......((((((((((((.......)))))).....(((((.....(((((.....(((.((....)).))).....)))))(((((....))))).)))))))))))........... ( -34.50) >DroSec_CAF1 3925 120 + 1 UUUCUGACUGCGAGUGACAUUAUUACUGUCACACUGUGUCGCCCGAUGCAGCUCCUCUGCCUUUUAUAAUGCAUUUAUGCUGCAAUUUGUUUGAAUUAGCGGCUCGCGGGCCCCAACUUG .......((((((((((((.......)))))).....(((((.....(((((.....(((.((....)).))).....)))))(((((....))))).)))))))))))........... ( -34.50) >DroSim_CAF1 9844 120 + 1 UUUCUGACUGCGAGUGACAUUAUUACUGUCACACUGUGUCGCCCGAUGCAGCUCCUCUGCCUUUUAUAAUGCAUUUAUGCUGCAAUUUGUUUGAAUUAGCGGCUCGCGGGCCCCAACUUG .......((((((((((((.......)))))).....(((((.....(((((.....(((.((....)).))).....)))))(((((....))))).)))))))))))........... ( -34.50) >DroEre_CAF1 3988 119 + 1 UUUCUGACUGCCAGUGACAUUAUUACUGUCACACUGUGUCGCCCGAUGCUGCUCCUCUGCCUUUUAUAAUGCAUUUAUGCUGCAAUUUGUUUGAAUUAGCGGCUCGCG-GCCCCAACUUG .....(((.....((((((.......)))))).....)))((((((.((((((.....((.((....)).))(((((.((........)).))))).))))))))).)-))......... ( -29.40) >DroYak_CAF1 15260 120 + 1 UUUCUGACUGCGAGUGACAUUAUUACUGUCACACUGUGUCGCCCGAUGCAGCUCCUCUGCCUUUUAUAAUGCAUUUAUGCUGCAAUUUGUUUGAAUUAGCGGCUCGCGGGCCCCAACUUG .......((((((((((((.......)))))).....(((((.....(((((.....(((.((....)).))).....)))))(((((....))))).)))))))))))........... ( -34.50) >consensus UUUCUGACUGCGAGUGACAUUAUUACUGUCACACUGUGUCGCCCGAUGCAGCUCCUCUGCCUUUUAUAAUGCAUUUAUGCUGCAAUUUGUUUGAAUUAGCGGCUCGCGGGCCCCAACUUG .......((((((((((((.......)))))).....(((((.....(((((.....(((.((....)).))).....)))))(((((....))))).)))))))))))........... (-31.22 = -31.82 + 0.60)

| Location | 17,650,325 – 17,650,445 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -33.40 |
| Consensus MFE | -31.38 |
| Energy contribution | -31.98 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966571 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
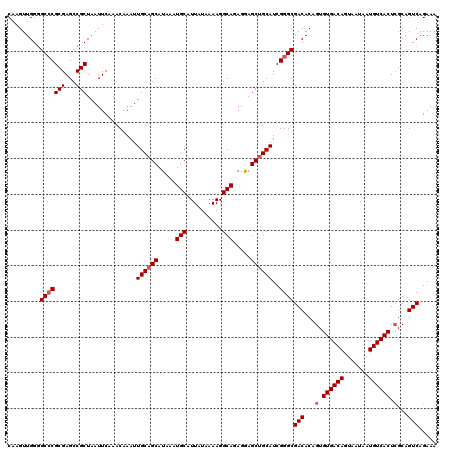
>2R_DroMel_CAF1 17650325 120 - 20766785 CAAGUUGGUGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGAAGCUGCAUCGGGCGACACAGUGUGACAGUAAUAAUGUCACUCGCAGUCAGAAA .........(((((((...)))..............((((((.....(((..........))).....))))))..))))(((...(.((((((.......)))))).)...)))..... ( -34.60) >DroSec_CAF1 3925 120 - 1 CAAGUUGGGGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCUGCAUCGGGCGACACAGUGUGACAGUAAUAAUGUCACUCGCAGUCAGAAA .........(((((((...)))..............((((((.....(((..........))).....))))))..))))(((...(.((((((.......)))))).)...)))..... ( -34.60) >DroSim_CAF1 9844 120 - 1 CAAGUUGGGGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCUGCAUCGGGCGACACAGUGUGACAGUAAUAAUGUCACUCGCAGUCAGAAA .........(((((((...)))..............((((((.....(((..........))).....))))))..))))(((...(.((((((.......)))))).)...)))..... ( -34.60) >DroEre_CAF1 3988 119 - 1 CAAGUUGGGGC-CGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCAGCAUCGGGCGACACAGUGUGACAGUAAUAAUGUCACUGGCAGUCAGAAA .........((-(.(((((.(((..((.........((((.......))))............))..))).)).))))))(((.....((((((.......)))))).....)))..... ( -28.60) >DroYak_CAF1 15260 120 - 1 CAAGUUGGGGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCUGCAUCGGGCGACACAGUGUGACAGUAAUAAUGUCACUCGCAGUCAGAAA .........(((((((...)))..............((((((.....(((..........))).....))))))..))))(((...(.((((((.......)))))).)...)))..... ( -34.60) >consensus CAAGUUGGGGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCUGCAUCGGGCGACACAGUGUGACAGUAAUAAUGUCACUCGCAGUCAGAAA .........(((((((...)))..............((((((.....(((..........))).....))))))..))))(((...(.((((((.......)))))).)...)))..... (-31.38 = -31.98 + 0.60)


| Location | 17,650,365 – 17,650,485 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -40.98 |
| Consensus MFE | -34.16 |
| Energy contribution | -35.40 |
| Covariance contribution | 1.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.83 |
| SVM decision value | 2.21 |
| SVM RNA-class probability | 0.990407 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
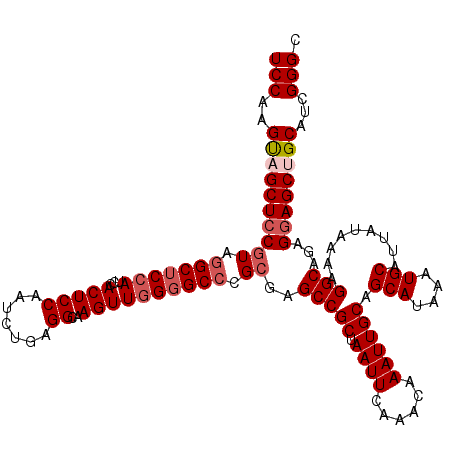
>2R_DroMel_CAF1 17650365 120 - 20766785 UCCAAGUUGCUCCGUAGGCUCCAUCCACUCCAAUCUGAGGCAAGUUGGUGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGAAGCUGCAUCGGGC (((.(((.((((.(..(((.(((....(((......)))......))).)))..))))).))).............((((((.....(((..........))).....))))))..))). ( -36.20) >DroSec_CAF1 3965 120 - 1 UCCAAGUAGCUCCGUAGGCUCCAUCCACUCCAAUCUGAGGCAAGUUGGGGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCUGCAUCGGGC (((..(((((((((..(((((((....(((......)))......)))))))..)..(((((.((((......)))))).(((....))).........)))...))))))))...))). ( -43.10) >DroSim_CAF1 9884 120 - 1 UCCAAGUAGCUCCGUAGGCUCCAUCCACUCCAAUCUGAGGCAAGUUGGGGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCUGCAUCGGGC (((..(((((((((..(((((((....(((......)))......)))))))..)..(((((.((((......)))))).(((....))).........)))...))))))))...))). ( -43.10) >DroEre_CAF1 4028 119 - 1 UCCAAGUAGCUCCGUUGGCUCGAUCCACUCCAAUCCGUGGCAAGUUGGGGC-CGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCAGCAUCGGGC .................((((((((..((((....((((((........))-)))).(((((.((((......)))))).(((....))).........)))...))))..).))))))) ( -35.20) >DroYak_CAF1 15300 120 - 1 UCCAAGCAGCUCCAUCGGCUCCAGCCACUCCGAUCCGCGGCAAGUUGGGGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCUGCAUCGGGC (((..((((((((.(((((((((((....(((.....)))...))))))))).....(((((.((((......)))))).(((....))).........))).))))))))))...))). ( -47.30) >consensus UCCAAGUAGCUCCGUAGGCUCCAUCCACUCCAAUCUGAGGCAAGUUGGGGCCCGCGAGCCGCUAAUUCAAACAAAUUGCAGCAUAAAUGCAUUAUAAAAGGCAGAGGAGCUGCAUCGGGC (((..((((((((((.(((((((...(((((.......))..)))))))))).))..(((((.((((......)))))).(((....))).........)))...))))))))...))). (-34.16 = -35.40 + 1.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:46:12 2006