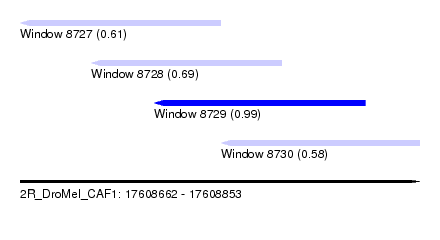
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,608,662 – 17,608,853 |
| Length | 191 |
| Max. P | 0.991237 |

| Location | 17,608,662 – 17,608,758 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.23 |
| Mean single sequence MFE | -28.18 |
| Consensus MFE | -15.19 |
| Energy contribution | -15.07 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.614836 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17608662 96 - 20766785 CACCACUGCCCCCCA--------UCAACUGGGCAAUCACUGACAUCGCAG---------UGGCAUGGUUU-UCGGGGCUAUUCCAAAUAGAUAUCAUUCAAAGAUAUA------UUUCUC .......(((((...--------..((((..((...(((((......)))---------))))..)))).-..)))))............(((((.......))))).------...... ( -21.10) >DroSec_CAF1 23335 99 - 1 CACCACUGCCCCCCA--------UCAACUGGGCAAUCACUGACAUCGCAG---------AGGCAUGGUUU-UGGGGGCUAUUCCAAAUAGAUAUCAUUGAGAGAUAUA---AUAAUUAUC .((((.((((.((((--------.....))))......(((......)))---------.))))))))((-(((((....)))))))...(((((.......))))).---......... ( -22.70) >DroSim_CAF1 24110 99 - 1 CACCACUGCCCCCCA--------UCAACUGGGCAAUCACUGACAUCGCAG---------AGGCAUGGUUU-UGGGGGCUAUUCCAAAUAGAUAUCAUUGAGAGAUAUA---AUAAUUCUC .((((.((((.((((--------.....))))......(((......)))---------.))))))))((-(((((....)))))))...(((((.......))))).---......... ( -22.70) >DroYak_CAF1 25062 120 - 1 CACCACUGCCCCCCACCAACCGACCAACUGGGCAAUCACUGACAUCGCAGUGAUUGCAGGGGCAUGGAUUUUUGGGGUAAUCCCAGAAAGAUAUUCGUGGAAGAAAUUAUCAAAAAUAUU ..(((.((((((.......(((......)))((((((((((......)))))))))).))))))))).((((((((.....))))))))((((((..(((((....)).)))..)))))) ( -46.20) >consensus CACCACUGCCCCCCA________UCAACUGGGCAAUCACUGACAUCGCAG_________AGGCAUGGUUU_UGGGGGCUAUUCCAAAUAGAUAUCAUUGAAAGAUAUA___AUAAUUAUC ..(((.((((.((((.............))))......(((......)))..........))))))).......(((....)))......(((((.......)))))............. (-15.19 = -15.07 + -0.12)

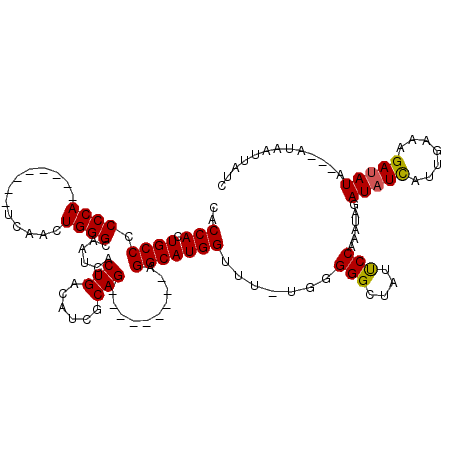

| Location | 17,608,696 – 17,608,787 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 79.14 |
| Mean single sequence MFE | -30.51 |
| Consensus MFE | -15.91 |
| Energy contribution | -15.95 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.694375 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17608696 91 - 20766785 GUUACAAUAAUAACAGCCGUGAUUCCCGACACCACUGCCCCCCA--------UCAACUGGGCAAUCACUGACAUCGCAG---------UGGCAUGGUUU-UCGGGGCUA ((((......))))((((.(((...(((...(((((((..((((--------.....))))((.....)).....))))---------)))..)))...-))).)))). ( -28.70) >DroSec_CAF1 23372 91 - 1 GUUACAAUAAUAACAGCCGUGAUUCCCGACACCACUGCCCCCCA--------UCAACUGGGCAAUCACUGACAUCGCAG---------AGGCAUGGUUU-UGGGGGCUA ((((......))))((((......(((((.((((.((((.((((--------.....))))......(((......)))---------.)))))))).)-)))))))). ( -29.20) >DroSim_CAF1 24147 91 - 1 GUUACAAUAAUAACAGCCGUGAUUCCCGACACCACUGCCCCCCA--------UCAACUGGGCAAUCACUGACAUCGCAG---------AGGCAUGGUUU-UGGGGGCUA ((((......))))((((......(((((.((((.((((.((((--------.....))))......(((......)))---------.)))))))).)-)))))))). ( -29.20) >DroEre_CAF1 25121 94 - 1 GUUACACUAAUAACAGCCGUGAUUCCCGACACCA--------------CCAACCCACUGGGCAAUCACCGACUUCGCAGUGGCGGCAGGGGCAUGAACC-UGGGGGCAU ((((......)))).(((((((((((((......--------------.........)))).)))))).....((((....))))((((........))-))..))).. ( -25.16) >DroYak_CAF1 25102 109 - 1 GUUACACUAAUAACAGCCGUGAUUCCCGACACCACUGCCCCCCACCAACCGACCAACUGGGCAAUCACUGACAUCGCAGUGAUUGCAGGGGCAUGGAUUUUUGGGGUAA ((((......)))).........((((((..(((.((((((.......(((......)))((((((((((......)))))))))).)))))))))....))))))... ( -40.30) >consensus GUUACAAUAAUAACAGCCGUGAUUCCCGACACCACUGCCCCCCA________UCAACUGGGCAAUCACUGACAUCGCAG_________AGGCAUGGUUU_UGGGGGCUA ((((......)))).(((((((((((((.............................)))).))))))...(((.((.............))))).........))).. (-15.91 = -15.95 + 0.04)

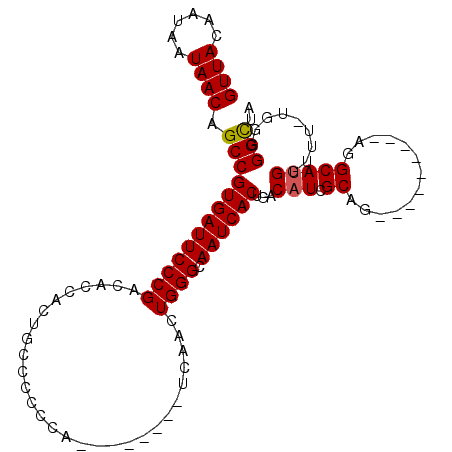

| Location | 17,608,726 – 17,608,827 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.46 |
| Mean single sequence MFE | -29.26 |
| Consensus MFE | -18.67 |
| Energy contribution | -19.70 |
| Covariance contribution | 1.03 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.64 |
| SVM decision value | 2.26 |
| SVM RNA-class probability | 0.991237 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17608726 101 - 20766785 CCGGGAUCGCGGUCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACAAUAAUAAC-----------AGCCGUGAUUCCCGACACCACUGCCCCCCA--------UCAACUGGGCAAUCACU .((((((((((((......(((....)))((((((((.....))))))))....-----------.))))))).))))).......(((((....--------......)))))...... ( -32.70) >DroVir_CAF1 32366 95 - 1 GCGCCG----UGUCCAUUUGGUAAUAUCUUUGCCGUAAUUGUAACAGUAAUGAUAAUGCCCAGCUAGCCGUGAUGCCUAAUGCCCAUGCUCAUUA--------UUAG------------- (.((.(----((..((((.((((.((((.((((.((.......)).)))).))))..((.......)).....)))).))))..))))).)....--------....------------- ( -16.40) >DroSec_CAF1 23402 101 - 1 CCGGGAUCGCGGCCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACAAUAAUAAC-----------AGCCGUGAUUCCCGACACCACUGCCCCCCA--------UCAACUGGGCAAUCACU .((((((((((((......(((....)))((((((((.....))))))))....-----------.))))))).))))).......(((((....--------......)))))...... ( -34.70) >DroSim_CAF1 24177 101 - 1 CCGGGAUCGCGGCCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACAAUAAUAAC-----------AGCCGUGAUUCCCGACACCACUGCCCCCCA--------UCAACUGGGCAAUCACU .((((((((((((......(((....)))((((((((.....))))))))....-----------.))))))).))))).......(((((....--------......)))))...... ( -34.70) >DroEre_CAF1 25160 95 - 1 CCGGGAUCGCGGUCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACACUAAUAAC-----------AGCCGUGAUUCCCGACACCA--------------CCAACCCACUGGGCAAUCACC .((((((((((((.......((((((....))))))...(((((......))))-----------)))))))).)))))......--------------(((......)))......... ( -26.60) >DroYak_CAF1 25142 109 - 1 CCGGGAUCGCGGUCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACACUAAUAAC-----------AGCCGUGAUUCCCGACACCACUGCCCCCCACCAACCGACCAACUGGGCAAUCACU .((((((((((((.......((((((....))))))...(((((......))))-----------)))))))).))))).......(((((..................)))))...... ( -30.47) >consensus CCGGGAUCGCGGUCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACAAUAAUAAC___________AGCCGUGAUUCCCGACACCACUGCCCCCCA________UCAACUGGGCAAUCACU ..(((((((((((......(((....)))((((((((.....))))))))................)))))))))))........................................... (-18.67 = -19.70 + 1.03)

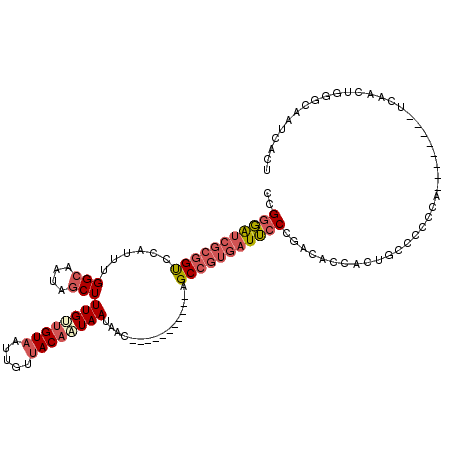
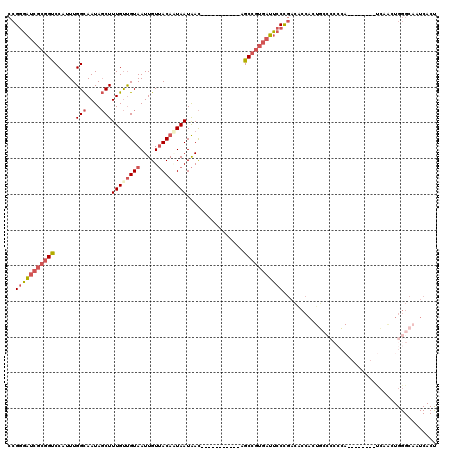
| Location | 17,608,758 – 17,608,853 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 83.93 |
| Mean single sequence MFE | -31.08 |
| Consensus MFE | -21.16 |
| Energy contribution | -23.55 |
| Covariance contribution | 2.39 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.579684 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
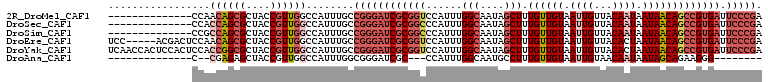
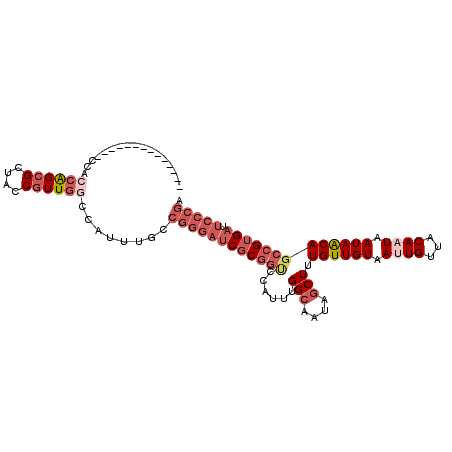
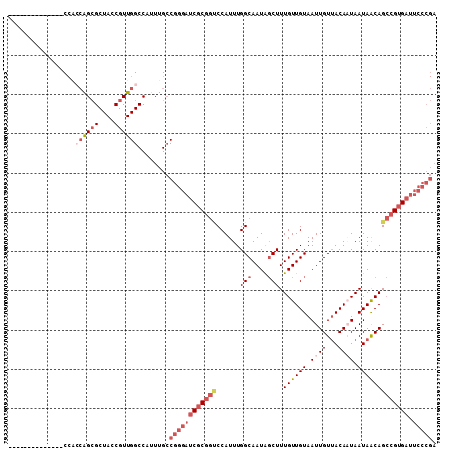
>2R_DroMel_CAF1 17608758 95 - 20766785 --------------CCAACAGCGCUACCGUUGGCCAUUUGCCGGGAUCGCGGUCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACAAUAAUAACAGCCGUGAUUCCCGA --------------....(((((....))))).........((((((((((((......(((....)))((((((((.....)))))))).....))))))).))))). ( -30.40) >DroSec_CAF1 23434 95 - 1 --------------CCACCAGCGCUACCGUUGGCCAUUUGCCGGGAUCGCGGCCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACAAUAAUAACAGCCGUGAUUCCCGA --------------...((((((....))))))........((((((((((((......(((....)))((((((((.....)))))))).....))))))).))))). ( -34.80) >DroSim_CAF1 24209 95 - 1 --------------CCGCCAGCGCUACCGUUGGCCAUUUGCCGGGAUCGCGGCCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACAAUAAUAACAGCCGUGAUUCCCGA --------------..(((((((....))))))).......((((((((((((......(((....)))((((((((.....)))))))).....))))))).))))). ( -38.40) >DroEre_CAF1 25186 104 - 1 UCC-----ACGACUCCAACAGCGCUACCGUUGGCCAUUUGCCGGGAUCGCGGUCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACACUAAUAACAGCCGUGAUUCCCGA ...-----..........(((((....))))).........((((((((((((.......((((((....))))))...(((((......)))))))))))).))))). ( -28.80) >DroYak_CAF1 25182 109 - 1 UCAACCACUCCACUCCACCGGCGCUACCGUUGGCCAUUUGCCGGGAUCGCGGUCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACACUAAUAACAGCCGUGAUUCCCGA .................((((((....))))))........((((((((((((.......((((((....))))))...(((((......)))))))))))).))))). ( -30.40) >DroAna_CAF1 23864 82 - 1 --------------C--CGAGAGCUACCGUUGGCCAUUUGGCGGGAUCGC---CCAUUUGGCAAUGCCUUUGUUGUAAUUGUAACAAUAAUAGCAGAAGGG-------- --------------.--.........((....((((..(((((....)).---)))..))))..(((..(((((((.......)))))))..)))...)).-------- ( -23.70) >consensus ______________CCACCAGCGCUACCGUUGGCCAUUUGCCGGGAUCGCGGUCCAUUUGGCAAUAGCUUUGUUGUAAUUGUUACAAUAAUAACAGCCGUGAUUCCCGA .................((((((....))))))........((((((((((((......(((....))).((((((.((((...)))).))))))))))))).))))). (-21.16 = -23.55 + 2.39)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:45:42 2006