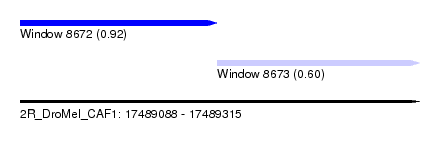
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,489,088 – 17,489,315 |
| Length | 227 |
| Max. P | 0.924157 |

| Location | 17,489,088 – 17,489,200 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.66 |
| Mean single sequence MFE | -34.98 |
| Consensus MFE | -29.11 |
| Energy contribution | -29.92 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.924157 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
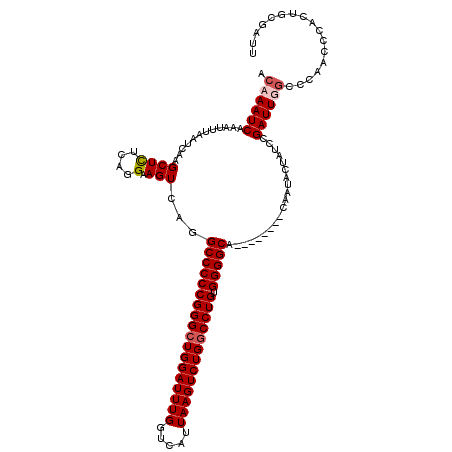
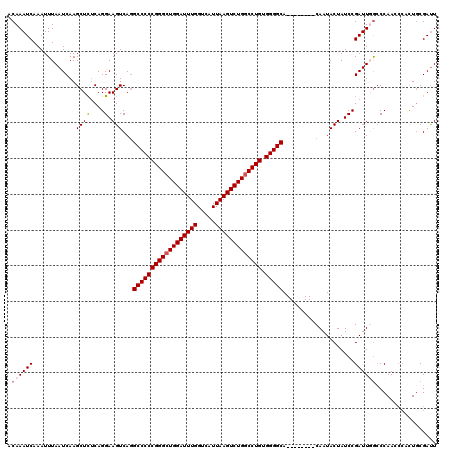
>2R_DroMel_CAF1 17489088 112 + 20766785 ACAAAUCAAAUUUAAUCAAGCUCUCAGGAAGUCAGGCCCCCGGGCUGGAUUUGGUCAUUAAGUCUGGCCUGUGGGGCA--------CAAUACUAUCCGAUUUAACCAACCCACUGCGACU ..........................((((((...((((((((((..((((((.....))))))..))))).))))).--------....))).)))....................... ( -34.00) >DroSec_CAF1 16904 112 + 1 ACAAAUCAAAUUUAAUCAAGCUUUCAGGAAGUCAGGCCCCCGGGAUGGAUUUGGUCAUUAAGUCUGGCCUGUGGGGCA--------CAAUACUAUCCGAUUUGCCCAACCCACUGCGAUU .((((((............(....).((((((...(((((((((..(((((((.....)))))))..)))).))))).--------....))).)))))))))................. ( -30.00) >DroSim_CAF1 29763 108 + 1 ACAAAUCAAAUUUAAUCAAGCUCUCAGGAAGUCAGGCCCCCGGGCUGGAUUUGGUCAUUAAGUCUGGCCUGUGGGGCA--------CAAUACUAUCCGAUUGGCCCAA----UUGCGAUU ...................(((....((((((...((((((((((..((((((.....))))))..))))).))))).--------....))).)))....)))....----........ ( -36.70) >DroYak_CAF1 25922 120 + 1 ACAAAUCAAAUUUAAUCAAGCUCGCAGGAAGUCAUGCCCCCGGGCUGGAUUUGGUCAUUAAGUCUGGCCUGCGGGGCAGAAUGCUACAAUACUAUCCGAUUGGCCCAAACCACUCCGAUU .......................(((.....((.(((((((((((..((((((.....))))))..))))).)))))))).))).........(((.((.(((......))).)).))). ( -39.20) >consensus ACAAAUCAAAUUUAAUCAAGCUCUCAGGAAGUCAGGCCCCCGGGCUGGAUUUGGUCAUUAAGUCUGGCCUGUGGGGCA________CAAUACUAUCCGAUUGGCCCAACCCACUGCGAUU .((((((............((((....).)))...((((((((((((((((((.....))))))))))))).)))))....................))))))................. (-29.11 = -29.92 + 0.81)

| Location | 17,489,200 – 17,489,315 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.85 |
| Mean single sequence MFE | -32.68 |
| Consensus MFE | -25.58 |
| Energy contribution | -26.08 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.596526 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
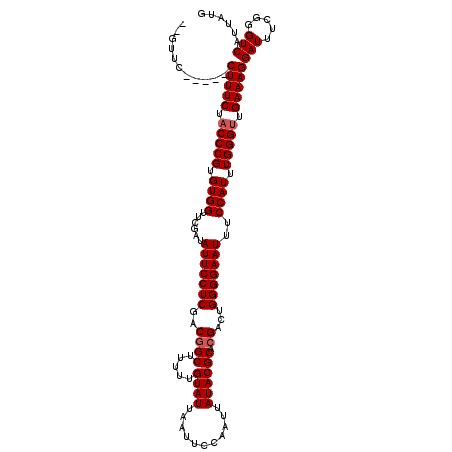
>2R_DroMel_CAF1 17489200 115 + 20766785 GUGUUC-----CUUUCUACCCGUGUGGUUCGAUAUUCCUCGACGGCUUUUUGUAUUAAUUCCAAUUAUACGCACGACUGGGGAAUUUCCAUUUGGGUUGAAAGGAUUUUGGGUCAUUAUG ....((-----(((((.(((((.((((......((((((((.((((.....((((...........)))))).))..))))))))..)))).))))).)))))))............... ( -32.70) >DroSec_CAF1 17016 113 + 1 --GUUC-----CUUUCUACCCGUGUGGUUCGAUAUUCCUCGACGGCUUUUUGUAUUAAUUCCAAUUAUACGCACGACUGGGGAAUUUCCAUUUGGGUUGAAAGGAUUUCGGGUCAUUAUG --..((-----(((((.(((((.((((......((((((((.((((.....((((...........)))))).))..))))))))..)))).))))).)))))))............... ( -32.70) >DroSim_CAF1 29871 113 + 1 --GUUC-----CUUUCUACCCGUGUGGUUCGAUAUUCCUCGACGGCUUUUUGUAUUAAUUCCAAUUAUACGCACGACUGGGGAAUUUCCAUUUGGGUUGAAAGGAUUUCGGGUCAUUAUG --..((-----(((((.(((((.((((......((((((((.((((.....((((...........)))))).))..))))))))..)))).))))).)))))))............... ( -32.70) >DroYak_CAF1 26042 118 + 1 --GUUCCUUUCCUUUCGCCCCGUGUGGUUCGAUAUUCCUCUACUGCUUUUUGUAUUAAUUCCAAUUAUACGCACGACUGGGGAAUUUCCAUUUGGGUUGAAAGGAUUUCGGGUCAUUAUG --(..((..((((((((.((((.((((......(((((((..((((.....((((...........))))))).)...)))))))..)))).)))).))))))))....))..)...... ( -32.60) >consensus __GUUC_____CUUUCUACCCGUGUGGUUCGAUAUUCCUCGACGGCUUUUUGUAUUAAUUCCAAUUAUACGCACGACUGGGGAAUUUCCAUUUGGGUUGAAAGGAUUUCGGGUCAUUAUG ...........(((((.(((((.((((......(((((((..((((.....((((...........)))))).))...)))))))..)))).))))).)))))(((.....)))...... (-25.58 = -26.08 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:44:48 2006