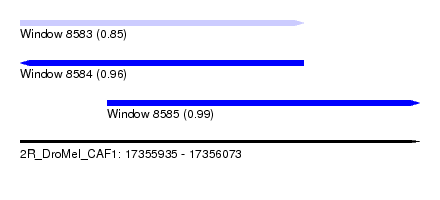
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,355,935 – 17,356,073 |
| Length | 138 |
| Max. P | 0.986512 |

| Location | 17,355,935 – 17,356,033 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 73.88 |
| Mean single sequence MFE | -17.84 |
| Consensus MFE | -13.30 |
| Energy contribution | -13.30 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.05 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.851221 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17355935 98 + 20766785 AUGAACAAAUAUUUUU---CA------GAAAU-AAAAAAAAA--AA---ACAACAAAAAGUUUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAA ...........(((((---((------(....-.........--((---((........))))......((((.(((((((.......))))).)).)).))))))))))... ( -16.00) >DroVir_CAF1 64479 89 + 1 --UACUAAAUAUAUUUCC---------U---U-AGAAAAAAA---------AAUAAAAUAUUUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAA --...........(((((---------.---.-.........---------......(((......)))((((.(((((((.......))))).)).)).))..))))).... ( -15.00) >DroGri_CAF1 57954 97 + 1 ---AGUUAAUUUAGUUUGAUGUGGUAAU---C-AAAAUUAAC---------AACAAAAAAUUAACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAA ---(((((((((..((((.(((......---.-.......))---------).))))))))))))).....((.(((((((.......))))).)).))((((.....)))). ( -21.04) >DroEre_CAF1 70450 87 + 1 AUGAACAAACAUUUUA---U-------------AAAA-UAAA---------AACAAAAAGCUUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAA ................---.-------------....-....---------........((((.((...((((.(((((((.......))))).)).)).))..)).)))).. ( -18.20) >DroYak_CAF1 52067 80 + 1 AUGAAUAAA------A---U-------------AAAA--AAA---------AUCAAAAAGUUUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAA .........------.---.-------------....--...---------........((((.((...((((.(((((((.......))))).)).)).))..)).)))).. ( -15.70) >DroPer_CAF1 78643 99 + 1 --GAACGAAAACAUAA---GC------CG--AAAAAA-UAGAAAAAUAUUUUACAAAAAAUGUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAA --((.(((...(((((---(.------..--..((((-((......))))))(((.....))).))))))..))).(((((.......))))).))...((((.....)))). ( -21.10) >consensus __GAACAAAUAUAUUA___U_____________AAAA_UAAA_________AACAAAAAAUUUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAA .......................................................................((.(((((((.......))))).)).))((((.....)))). (-13.30 = -13.30 + -0.00)



| Location | 17,355,935 – 17,356,033 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 73.88 |
| Mean single sequence MFE | -19.33 |
| Consensus MFE | -14.50 |
| Energy contribution | -14.50 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.46 |
| SVM RNA-class probability | 0.956116 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17355935 98 - 20766785 UUGCCUUCCAAGCAGUUGAUCCGGGUUCGAUUCCCGGCCGACGCACAAGUAAACUUUUUGUUGU---UU--UUUUUUUUU-AUUUC------UG---AAAAAUAUUUGUUCAU .(((.......)))((((..(((((.......))))).))))(.((((((((((.....)))..---..--...((((((-(....------))---)))))))))))).).. ( -19.90) >DroVir_CAF1 64479 89 - 1 UUGCCUUCCAAGCAGUUGAUCCGGGUUCGAUUCCCGGCCGACGCACAAGUAAAUAUUUUAUU---------UUUUUUUCU-A---A---------GGAAAUAUAUUUAGUA-- ...........((.((((..(((((.......))))).))))))...(.((((((...((((---------(((((....-)---)---------))))))))))))).).-- ( -19.40) >DroGri_CAF1 57954 97 - 1 UUGCCUUCCAAGCAGUUGAUCCGGGUUCGAUUCCCGGCCGACGCACAAGUUAAUUUUUUGUU---------GUUAAUUUU-G---AUUACCACAUCAAACUAAAUUAACU--- ...........((.((((..(((((.......))))).))))))...(((((((((((((.(---------((((((...-.---))))..))).))))..)))))))))--- ( -22.40) >DroEre_CAF1 70450 87 - 1 UUGCCUUCCAAGCAGUUGAUCCGGGUUCGAUUCCCGGCCGACGCACAAGUAAGCUUUUUGUU---------UUUA-UUUU-------------A---UAAAAUGUUUGUUCAU ........((((((((((..(((((.......))))).))))..(((((.......))))).---------....-....-------------.---.....))))))..... ( -18.50) >DroYak_CAF1 52067 80 - 1 UUGCCUUCCAAGCAGUUGAUCCGGGUUCGAUUCCCGGCCGACGCACAAGUAAACUUUUUGAU---------UUU--UUUU-------------A---U------UUUAUUCAU ...........((.((((..(((((.......))))).)))))).((((.......))))..---------...--....-------------.---.------......... ( -14.90) >DroPer_CAF1 78643 99 - 1 UUGCCUUCCAAGCAGUUGAUCCGGGUUCGAUUCCCGGCCGACGCACAAGUACAUUUUUUGUAAAAUAUUUUUCUA-UUUUUU--CG------GC---UUAUGUUUUCGUUC-- ..(((......((.((((..(((((.......))))).)))))).....((((.....)))).............-......--.)------))---..............-- ( -20.90) >consensus UUGCCUUCCAAGCAGUUGAUCCGGGUUCGAUUCCCGGCCGACGCACAAGUAAACUUUUUGUU_________UUUA_UUUU_____________A___AAAUAUAUUUAUUC__ ...........((.((((..(((((.......))))).))))))..................................................................... (-14.50 = -14.50 + 0.00)

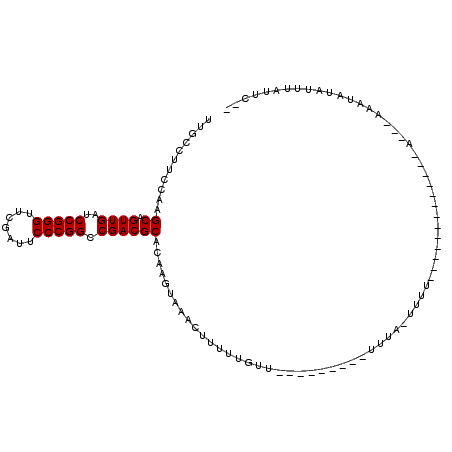
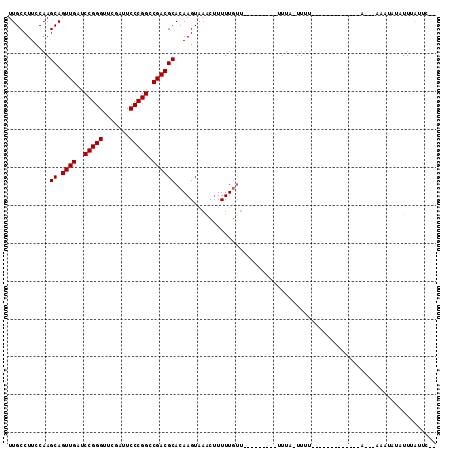
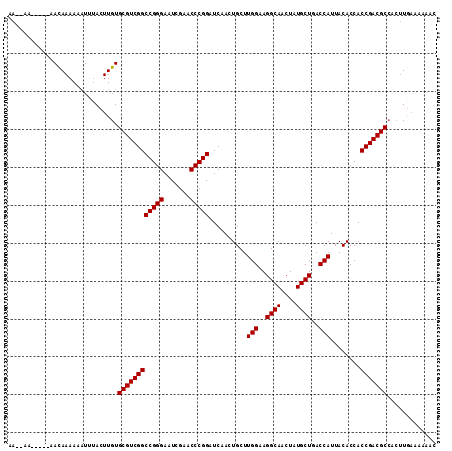
| Location | 17,355,965 – 17,356,073 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 85.97 |
| Mean single sequence MFE | -31.57 |
| Consensus MFE | -29.05 |
| Energy contribution | -29.05 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.92 |
| SVM decision value | 2.04 |
| SVM RNA-class probability | 0.986512 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17355965 108 + 20766785 AA--AA---ACAACAAAAAGUUUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAACUAUGCUGACCAUUACACCACCGACGCCCCUUAAAAAUAG ..--((---((........))))......((((((((((((.......)))))..........(((..((((....))))..)))........)))))))............. ( -29.30) >DroVir_CAF1 64504 104 + 1 AA---------AAUAAAAUAUUUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAACUAUGCUGACCAUUACACCACCGACGCCCGUUGUUUUCAA ..---------................(.((((((((((((.......)))))..........(((..((((....))))..)))........))))))).)........... ( -29.30) >DroPse_CAF1 79534 113 + 1 GAAAAAUAUUUUACAAAAAAUGUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAACUAUGCUGACCAUUACACCACCGACGCCACUUGAAAGAUC .....(((((((....))))))).(..((((((((((((((.......)))))..........(((..((((....))))..)))........)))))).)))..)....... ( -32.20) >DroGri_CAF1 57987 104 + 1 AC---------AACAAAAAAUUAACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAACUAUGCUGACCAUUACACCACCGACGCCACAUGAAUAUAG ..---------.............(.(((((((((((((((.......)))))..........(((..((((....))))..)))........)))))).)))).)....... ( -31.20) >DroSec_CAF1 71844 108 + 1 AA--AG---GAAACAAAAAAUUUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAACUAUGCUGACCAUUACACCACCGACGCGCCUUGAGGAGAU ..--.(---....)..........(((((((((((((((((.......)))))..........(((..((((....))))..)))........)))))))))...)))..... ( -35.20) >DroPer_CAF1 78669 113 + 1 GAAAAAUAUUUUACAAAAAAUGUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAACUAUGCUGACCAUUACACCACCGACGCCACUUGAAAGAUC .....(((((((....))))))).(..((((((((((((((.......)))))..........(((..((((....))))..)))........)))))).)))..)....... ( -32.20) >consensus AA__AA_____AACAAAAAAUUUACUUGUGCGUCGGCCGGGAAUCGAACCCGGAUCAACUGCUUGGAAGGCAACUAUGCUGACCAUUACACCACCGACGCCACUUGAAAAAAC .............................((((((((((((.......)))))..........(((..((((....))))..)))........)))))))............. (-29.05 = -29.05 + 0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:43:23 2006