
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,349,226 – 17,349,330 |
| Length | 104 |
| Max. P | 0.871119 |

| Location | 17,349,226 – 17,349,330 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 73.69 |
| Mean single sequence MFE | -28.00 |
| Consensus MFE | -15.50 |
| Energy contribution | -15.33 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.871119 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
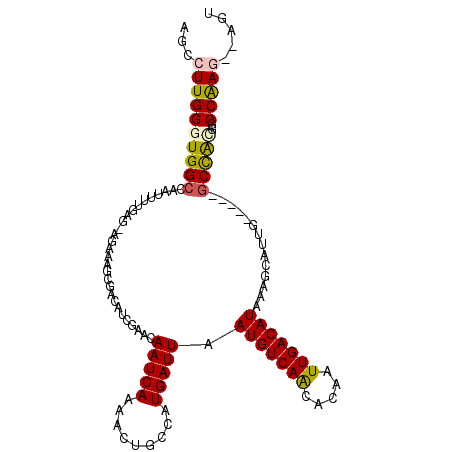
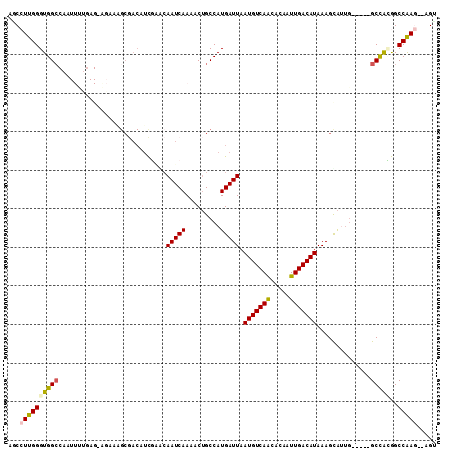
>2R_DroMel_CAF1 17349226 104 + 20766785 ACCCUUGGGUGGCCAAUUUUAAGCAGAACGCGACAUCGAGCAAUCAAAACUGCCAUGAUUAAUGUCAGCACAAUUGACAUAAAGCAUUG-----GCCAUGGCCGAG--AGU ...((((((((((((((.....((((.....(....)((....))....))))........(((((((.....))))))).....))))-----)))))..)))))--... ( -30.10) >DroVir_CAF1 57238 99 + 1 GGCCUUGGGUGGCCAAUUUCGAGGAG----------UGCACAAUCAAAACUGCCAUGAUUAAUGUCAACACAAUUGACAUAAAGUAGGAACGCGGCUGCAGCCAAA--UGU (((.....(..(((....(((.((((----------(...........))).)).)))...(((((((.....))))))).............)))..).)))...--... ( -23.80) >DroPse_CAF1 72954 93 + 1 AGCCUUGGGUGGC-----------AUCCAGCGACAUCCAACAAUCAAAACUGCCAUGAUUAAUGUCAACACAAUUGACAUAAAGCAUUG-----GCCAGGGCCAAG--AGC .(((((((((.((-----------.....)).))...............((((((((.((.(((((((.....))))))).)).)).))-----).)))..)))))--.)) ( -26.40) >DroMoj_CAF1 62865 108 + 1 AACGUUGGGUGGCCAAUUUUGAGUGGUUGGCAACGUUGCGAAAUCAAAACGGCCAUGAUUAAUGUCAACACAAUUGACAUAAAGUGCUGA-GCGGCUGCAGCCAAA--UGU ...(((.(((((((..((((((....((.(((....))).)).)))))).)))).......(((((((.....))))))).....))).)-))(((....)))...--... ( -31.90) >DroAna_CAF1 57905 106 + 1 ACCCUUGGGUGGGCAAUUUUGAGCAGAACGCGACAUCGGGCAAUCAAAACUGCCAUGAUUAAUGUCAGCACAAUUGACAUAAAGCAUUG-----GCCAUGGCCGAGAGAGU (((....))).((((.((((((...(..((......))..)..)))))).)))).((.((.(((((((.....))))))).)).))(((-----((....)))))...... ( -29.40) >DroPer_CAF1 72106 93 + 1 AGCCUUGGGUGGC-----------AUCCAGCGACAUCCAACAAUCAAAACUGCCAUGAUUAAUGUCAACACAAUUGACAUAAAGCAUUG-----GCCAGGGCCAAG--AGC .(((((((((.((-----------.....)).))...............((((((((.((.(((((((.....))))))).)).)).))-----).)))..)))))--.)) ( -26.40) >consensus AGCCUUGGGUGGCCAAUUUUGAG_AGAAAGCGACAUCGAACAAUCAAAACUGCCAUGAUUAAUGUCAACACAAUUGACAUAAAGCAUUG_____GCCACGGCCAAG__AGU ...((((((((((............................(((((.........))))).(((((((.....)))))))..............)))))..)))))..... (-15.50 = -15.33 + -0.16)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:43:16 2006