
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,263,097 – 17,263,192 |
| Length | 95 |
| Max. P | 0.908300 |

| Location | 17,263,097 – 17,263,192 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 77.63 |
| Mean single sequence MFE | -24.97 |
| Consensus MFE | -13.29 |
| Energy contribution | -13.93 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.53 |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.908300 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
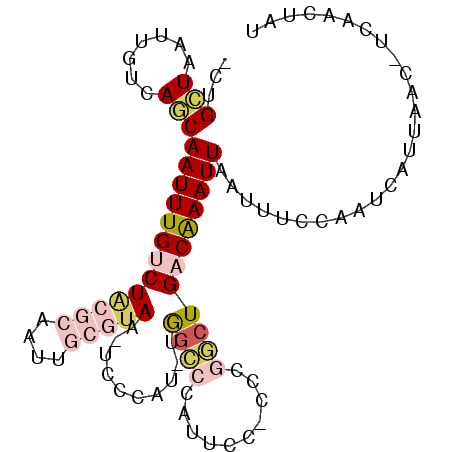
>2R_DroMel_CAF1 17263097 95 + 20766785 AUAGCUGA-GUUAAUGAUUGGAAAUUAAUUUGUCAGCCGGG-AGAAUGGGCCA-AUGGGA-UUACGCAAUUGCGUAGACAAAUUGCUGACAAUUAGCAG- ...(((((-((((((........))))))((((((((..((-........))(-((..(.-((((((....)))))).)..))))))))))))))))..- ( -26.90) >DroVir_CAF1 72864 88 + 1 AUUGUUGA-CUUAAUGAUUGGAUAUUAAUUUGGCAGGCUGU-GUUA-GCCUCA-AUAGUACCUACUU-------CAGACAAAUUGUUGUCGAUUAGCCG- ...(((((-.....(((..((.(((((..(((..((((((.-..))-))))))-))))))))....)-------))(((((....)))))..)))))..- ( -16.80) >DroSim_CAF1 56535 95 + 1 AUAGCUGA-GUUAAUGAUUGGAAAUUAAUUCGUCAGCCGGG-GGAACGGGCCA-AUGGGA-UUACGCAAUUGCGUAGACAAAUUGCUGACAAUUAGCAG- ...(((((-((((((........))))))))((((((((..-....)))..((-((..(.-((((((....)))))).)..)))))))))....)))..- ( -30.10) >DroEre_CAF1 55878 95 + 1 AUAGUUGA-GUUAAUGAUUGGAAAUUAAUUUGUCAGCCGGG-GGAAUGGGCCA-AUGGGA-UUACGCAAUUGCGUAGACAAAUUGCUGACAAUUAGCAG- ........-((((((..(..(....(((((((((..(((..-((......)).-.)))..-.(((((....)))))))))))))))..)..))))))..- ( -26.60) >DroWil_CAF1 77528 91 + 1 AUAGUUAAACUUAAUGAUUGGAAAUUAAUUUGACAGGCUGA-AGCAUUUGCUGUGUGUAG-AUGCUUA-------AGACAAAUUGUUGUCGAUUAACCAG ...((((((.(((((........))))))))))).((...(-(((((((((.....))))-)))))).-------.(((((....)))))......)).. ( -23.20) >DroYak_CAF1 55086 96 + 1 AUAGUUGA-GUUAAUGAUUGGAAAUUAAUUUGUCAGCCGGGGGGAAUGGGCCA-AUGGGA-UUACGCAAUUGCGUAGACAAAUUGCUGACAAUUAGCAG- ........-((((((..(..(....(((((((((..(((..((.......)).-.)))..-.(((((....)))))))))))))))..)..))))))..- ( -26.20) >consensus AUAGUUGA_GUUAAUGAUUGGAAAUUAAUUUGUCAGCCGGG_GGAAUGGGCCA_AUGGGA_UUACGCAAUUGCGUAGACAAAUUGCUGACAAUUAGCAG_ ...(((((..(((((........))))).((((((((.....((......)).........((((((....)))))).......)))))))))))))... (-13.29 = -13.93 + 0.64)



| Location | 17,263,097 – 17,263,192 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 77.63 |
| Mean single sequence MFE | -17.47 |
| Consensus MFE | -7.86 |
| Energy contribution | -9.37 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.45 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.682835 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17263097 95 - 20766785 -CUGCUAAUUGUCAGCAAUUUGUCUACGCAAUUGCGUAA-UCCCAU-UGGCCCAUUCU-CCCGGCUGACAAAUUAAUUUCCAAUCAUUAAC-UCAGCUAU -..(((..((((((((........(((((....))))).-......-(((........-.))))))))))).(((((........))))).-..)))... ( -19.90) >DroVir_CAF1 72864 88 - 1 -CGGCUAAUCGACAACAAUUUGUCUG-------AAGUAGGUACUAU-UGAGGC-UAAC-ACAGCCUGCCAAAUUAAUAUCCAAUCAUUAAG-UCAACAAU -.((((..(((((((....))))).)-------)....(((.....-..((((-(...-..))))))))....................))-))...... ( -15.51) >DroSim_CAF1 56535 95 - 1 -CUGCUAAUUGUCAGCAAUUUGUCUACGCAAUUGCGUAA-UCCCAU-UGGCCCGUUCC-CCCGGCUGACGAAUUAAUUUCCAAUCAUUAAC-UCAGCUAU -..(((....((((((........(((((....))))).-......-.....((....-..))))))))((.(((((........))))).-)))))... ( -20.50) >DroEre_CAF1 55878 95 - 1 -CUGCUAAUUGUCAGCAAUUUGUCUACGCAAUUGCGUAA-UCCCAU-UGGCCCAUUCC-CCCGGCUGACAAAUUAAUUUCCAAUCAUUAAC-UCAACUAU -..(((.......)))(((((((((((((....))))).-......-.((((......-...)))))))))))).................-........ ( -18.80) >DroWil_CAF1 77528 91 - 1 CUGGUUAAUCGACAACAAUUUGUCU-------UAAGCAU-CUACACACAGCAAAUGCU-UCAGCCUGUCAAAUUAAUUUCCAAUCAUUAAGUUUAACUAU .(((((((.((((((....))))).-------.((((((-.............)))))-)..............................).))))))). ( -11.42) >DroYak_CAF1 55086 96 - 1 -CUGCUAAUUGUCAGCAAUUUGUCUACGCAAUUGCGUAA-UCCCAU-UGGCCCAUUCCCCCCGGCUGACAAAUUAAUUUCCAAUCAUUAAC-UCAACUAU -..(((.......)))(((((((((((((....))))).-......-.((((..........)))))))))))).................-........ ( -18.70) >consensus _CUGCUAAUUGUCAGCAAUUUGUCUACGCAAUUGCGUAA_UCCCAU_UGGCCCAUUCC_CCCGGCUGACAAAUUAAUUUCCAAUCAUUAAC_UCAACUAU ...(((.......)))(((((((((((((....)))))..........((((..........)))))))))))).......................... ( -7.86 = -9.37 + 1.50)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:42:16 2006