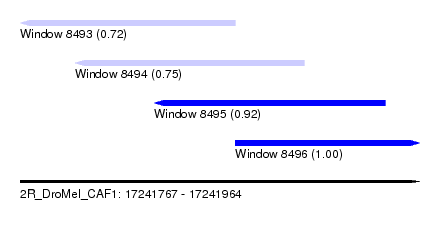

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,241,767 – 17,241,964 |
| Length | 197 |
| Max. P | 0.995566 |

| Location | 17,241,767 – 17,241,873 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.30 |
| Mean single sequence MFE | -18.34 |
| Consensus MFE | -14.20 |
| Energy contribution | -14.45 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724996 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17241767 106 - 20766785 AACAACGAGCAAUAACAACAAAAGGCGCCACAAGUUGGCAGCCUUUUC-GCUCCGCCGUUUUCAUACUUCAUUCGCUUUGUGCACUUUCUUCCCCCU----------ACCCCUUUCC--- ...((((.((.........(((((((((((.....)))).))))))).-.....))))))..............((.....))..............----------..........--- ( -18.46) >DroSim_CAF1 44444 106 - 1 AACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCAGCCUUUUC-GCUCCGCCGUUUUCAUACUUCAUUCGCUUUGUGCACUUUCUUCCCCCU----------ACCCCUUCCC--- .(((((((((.........(((((((((((.....)))).))))))).-........((......))...)))))..))))................----------..........--- ( -17.60) >DroEre_CAF1 43622 111 - 1 AACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCAGCCUUUUU-GCUCCGCCGUUUUCAUACUUCAUUCGCUUUGUGCAUUUUCUUCCC-CCA----CACUCACCCUUCACC--- ......((((.........(((((((((((.....)))).))))))).-.........................((.....))...........-...----.))))..........--- ( -19.80) >DroAna_CAF1 39308 113 - 1 AACAAC-------UACAACAAAAGGCGCCACAAGUUGGCAGCCCUUUCGGCUCUACAAUUUUCAUACUUCAUUCGCUUUGUGCAUAUUCUUCUGCCCAUUUCCCCUUGCCCCUUGCCCCC ......-------.....(((..((((....((((.(((((((.....))).......................((.....)).........)))).)))).....))))..)))..... ( -17.50) >consensus AACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCAGCCUUUUC_GCUCCGCCGUUUUCAUACUUCAUUCGCUUUGUGCACUUUCUUCCCCCC__________ACCCCUUACC___ ...................(((((((((((.....)))).)))))))...........................((.....))..................................... (-14.20 = -14.45 + 0.25)

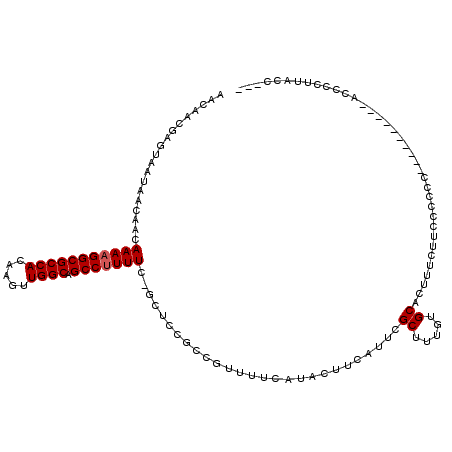
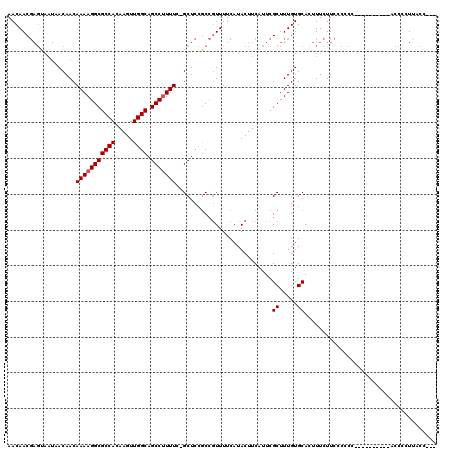
| Location | 17,241,794 – 17,241,907 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.77 |
| Mean single sequence MFE | -24.95 |
| Consensus MFE | -17.99 |
| Energy contribution | -18.48 |
| Covariance contribution | 0.49 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.747523 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17241794 113 - 20766785 AAAAGUGCAAUAACAAACAGGAA------ACAGCAACAGCAACAACGAGCAAUAACAACAAAAGGCGCCACAAGUUGGCAGCCUUUUC-GCUCCGCCGUUUUCAUACUUCAUUCGCUUUG .((((((.(((........((((------((.((....((........)).........(((((((((((.....)))).))))))).-.....)).)))))).......))))))))). ( -26.56) >DroSec_CAF1 41417 113 - 1 AAAAGUGCAAUAACAAACAGGAA------ACAGCAACAGCAACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCAGCCUUUUC-GCUCCGCCGUUUUCAUACUUCAUUCGCUUUG .((((((.(((........((((------((.((...(((........((....))...(((((((((((.....)))).))))))).-)))..)).)))))).......))))))))). ( -26.86) >DroSim_CAF1 44471 113 - 1 AAAAGUGCAAUAACAAACAGGAA------ACAGCAACAGCAACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCAGCCUUUUC-GCUCCGCCGUUUUCAUACUUCAUUCGCUUUG .((((((.(((........((((------((.((...(((........((....))...(((((((((((.....)))).))))))).-)))..)).)))))).......))))))))). ( -26.86) >DroEre_CAF1 43654 119 - 1 AAAAGUGCAAUAACAAACAGGAAGCAGCAACAGCAAAAGCAACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCAGCCUUUUU-GCUCCGCCGUUUUCAUACUUCAUUCGCUUUG .((((((.(((........((((((.((....((....))......((((((........((((((((((.....)))).))))))))-)))).)).)))))).......))))))))). ( -30.06) >DroAna_CAF1 39348 97 - 1 AAAAGUGCAAUAACAAACAGGAA----------------CAACAAC-------UACAACAAAAGGCGCCACAAGUUGGCAGCCCUUUCGGCUCUACAAUUUUCAUACUUCAUUCGCUUUG ....((......)).........----------------.......-------........(((((((((.....))))((((.....))))......................))))). ( -14.40) >consensus AAAAGUGCAAUAACAAACAGGAA______ACAGCAACAGCAACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCAGCCUUUUC_GCUCCGCCGUUUUCAUACUUCAUUCGCUUUG .((((((.(((........(((..........((....))...................(((((((((((.....)))).)))))))....)))...((......))...))))))))). (-17.99 = -18.48 + 0.49)

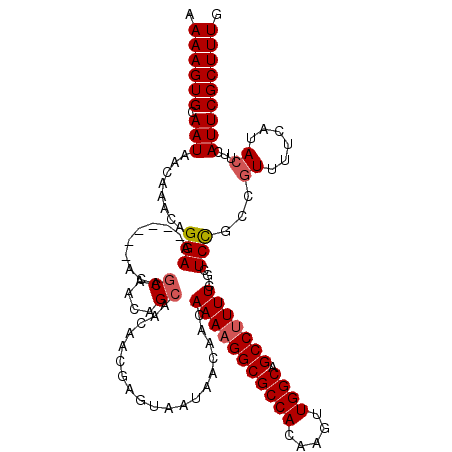
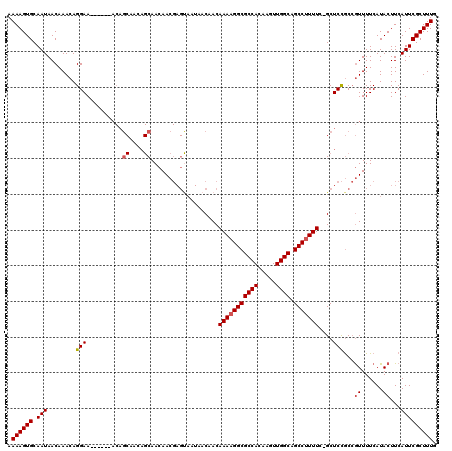
| Location | 17,241,833 – 17,241,947 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.30 |
| Mean single sequence MFE | -23.16 |
| Consensus MFE | -22.00 |
| Energy contribution | -22.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.95 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.915736 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17241833 114 - 20766785 CCCUUUCGCACACCUCUGCCUUUGUCUCUUUUCAGCACGCAAAAGUGCAAUAACAAACAGGAA------ACAGCAACAGCAACAACGAGCAAUAACAACAAAAGGCGCCACAAGUUGGCA .................(((((((((((......((((......))))...........(...------.).((....))......)))........)).))))))((((.....)))). ( -23.30) >DroSec_CAF1 41456 114 - 1 CCUUUUCCCACACCUCUGCCUUUGUCUCUUUCCAGCACGCAAAAGUGCAAUAACAAACAGGAA------ACAGCAACAGCAACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCA .................((((((..(((......((((......))))...........(...------.).((....))......)))...........))))))((((.....)))). ( -23.32) >DroSim_CAF1 44510 114 - 1 CCCUUUCCCACACCUCUGCCUUUGUCUCUUUUCAGCACGCAAAAGUGCAAUAACAAACAGGAA------ACAGCAACAGCAACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCA .................((((((..(((......((((......))))...........(...------.).((....))......)))...........))))))((((.....)))). ( -23.32) >DroEre_CAF1 43693 120 - 1 CUUUUUCCCGCACCUCUGCCUUUGUCUCUUUUCAGCACGCAAAAGUGCAAUAACAAACAGGAAGCAGCAACAGCAAAAGCAACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCA .................((((((((..(((((..((((......))))..................((....)))))))..)).....((....))....))))))((((.....)))). ( -22.70) >consensus CCCUUUCCCACACCUCUGCCUUUGUCUCUUUUCAGCACGCAAAAGUGCAAUAACAAACAGGAA______ACAGCAACAGCAACAACGAGUAAUAACAACAAAAGGCGCCACAAGUUGGCA .................(((((((((((......((((......))))........................((....))......)))........)).))))))((((.....)))). (-22.00 = -22.00 + 0.00)

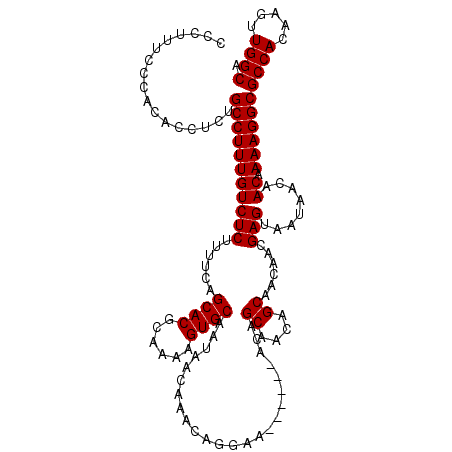
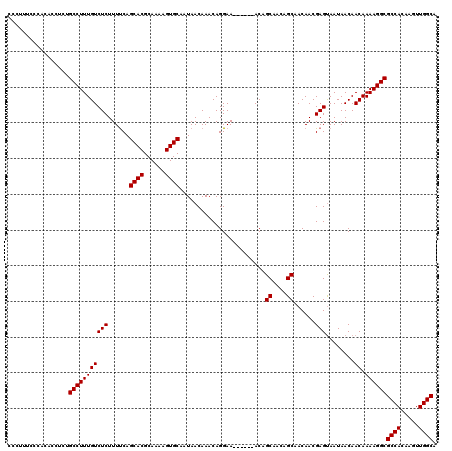
| Location | 17,241,873 – 17,241,964 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 83.02 |
| Mean single sequence MFE | -36.98 |
| Consensus MFE | -24.97 |
| Energy contribution | -24.66 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -3.45 |
| Structure conservation index | 0.68 |
| SVM decision value | 2.59 |
| SVM RNA-class probability | 0.995566 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17241873 91 + 20766785 GCUGUUGCUGU------UUCCUGUUUGUUAUUGCACUUUUGCGUGCUGAAAAGAGACAAAGGCAGAGGUGUGCGAAAGGGAGGGGU------------------GGGCAGCCAAA (((((..((.(------(((((.(((((....((((((((((.(.((....))......).))))))))))))))))))))).)).------------------.).)))).... ( -35.00) >DroSec_CAF1 41496 104 + 1 GCUGUUGCUGU------UUCCUGUUUGUUAUUGCACUUUUGCGUGCUGGAAAGAGACAAAGGCAGAGGUGUGGGAAAAGGAGGGGUGU-----GCGGCGGAGCAGGGCAGCCAAA (((.(((((((------..(((.(((....(..(((((((((.(..((........)).).)))))))))..).....))).)))...-----)))))))))).((....))... ( -37.00) >DroSim_CAF1 44550 96 + 1 GCUGUUGCUGU------UUCCUGUUUGUUAUUGCACUUUUGCGUGCUGAAAAGAGACAAAGGCAGAGGUGUGGGAAAGGGAGGGGCGU-----G--------CAGGGCAGCCAAA ((((((.((((------..(((.(((.((.(..(((((((((.(.((....))......).)))))))))..)..)).))).)))...-----)--------))))))))).... ( -35.40) >DroEre_CAF1 43733 108 + 1 GCUUUUGCUGUUGCUGCUUCCUGUUUGUUAUUGCACUUUUGCGUGCUGAAAAGAGACAAAGGCAGAGGUGCGGGAAAAAGAGAGAGGUGGUGGGGG-------GGGGCAGCCAAA ((((((.((.(..(..((((((.(((....((((((((((((.(.((....))......).))))))))))))...))).)).))))..)..).))-------.))).))).... ( -40.50) >consensus GCUGUUGCUGU______UUCCUGUUUGUUAUUGCACUUUUGCGUGCUGAAAAGAGACAAAGGCAGAGGUGUGGGAAAAGGAGGGGUGU_____G_________AGGGCAGCCAAA ......(((((......(((((.(((....((((((((((((.(..((........)).).))))))))))))...))).))))).....................))))).... (-24.97 = -24.66 + -0.31)

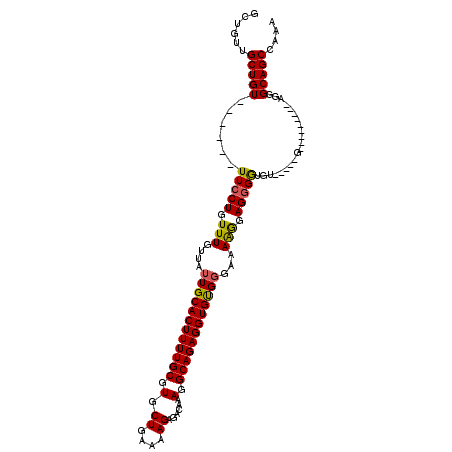
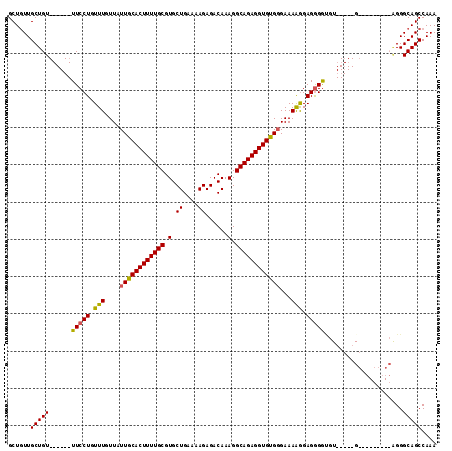
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:41:58 2006