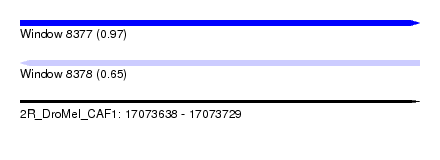

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,073,638 – 17,073,729 |
| Length | 91 |
| Max. P | 0.971381 |

| Location | 17,073,638 – 17,073,729 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 89.31 |
| Mean single sequence MFE | -23.28 |
| Consensus MFE | -20.70 |
| Energy contribution | -21.08 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.67 |
| SVM RNA-class probability | 0.971381 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
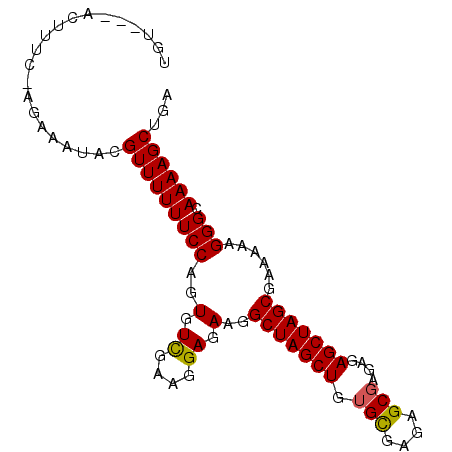
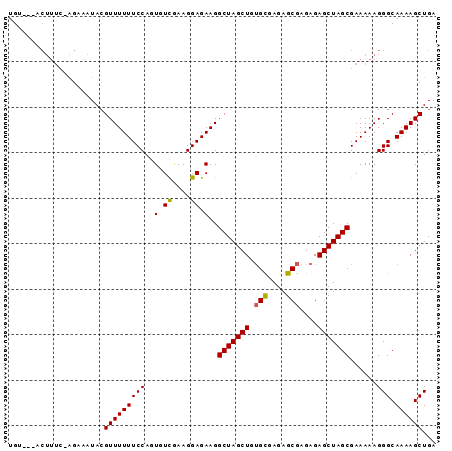
>2R_DroMel_CAF1 17073638 91 + 20766785 UGU---ACUUUCCAGUAAUACGUUUUUUCCAGUGUCGAAGGAGAAGGCUAGCUGGGUGAGAGCGAGAGAGCUAGCGAAAAAGGGCAAAAGCUGA ((.---.(((((((((..((.(((((((((.........))))))))))))))))((...(((......))).))...))))..))........ ( -22.50) >DroSec_CAF1 11169 91 + 1 UGU---ACUUUCAAGAAAUACGUUUUUUCCAGUGUCAAAGGAGAAGGCUAGCUGUGCGAGAGCGAGAGAGCUAGCGAAAAAGGGCAAAAGCUGA ...---..((((...........(((((((.........)))))))(((((((.(((....)))....)))))))))))...(((....))).. ( -24.60) >DroSim_CAF1 11267 91 + 1 UGU---GCUUUCAAGAAAUACGUUUUUUCCAGUGUCGAAGGAGAAGGCUAGCUGUGCGAGAGCGAGAGAGCUAGCGAAAAAGGGCAAAAGCUGA (((---.((((............(((((((.........)))))))(((((((.(((....)))....)))))))...)))).)))........ ( -25.70) >DroEre_CAF1 11507 86 + 1 UGGGU--------GGCAAUACGUUUUUUCCAGUGUCGAAGGAGAAGGCUAGCUGUGCAAGAGCGAGAGAGCUAGCGAAAAAGGGCAAAAGCUGA ...((--------(....)))(((((((((..(.((....)).)..(((((((.(((....)))....)))))))......))).))))))... ( -22.10) >DroYak_CAF1 12818 92 + 1 UGGAUUACUUU--AGGAAUAAGUUUUUUCCAGUGUUGAAGGAGAAGGCUAGCUGUGCAAGAGCGAGAGAGCUAGCGAAAAAGGGCAAAAGCUGA .......((((--..........(((((((.........)))))))(((((((.(((....)))....)))))))...))))(((....))).. ( -21.50) >consensus UGU___ACUUUC_AGAAAUACGUUUUUUCCAGUGUCGAAGGAGAAGGCUAGCUGUGCGAGAGCGAGAGAGCUAGCGAAAAAGGGCAAAAGCUGA .....................(((((((((..(.((....)).)..(((((((.(((....)))....)))))))......))).))))))... (-20.70 = -21.08 + 0.38)

| Location | 17,073,638 – 17,073,729 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 89.31 |
| Mean single sequence MFE | -16.55 |
| Consensus MFE | -11.88 |
| Energy contribution | -12.08 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.648058 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
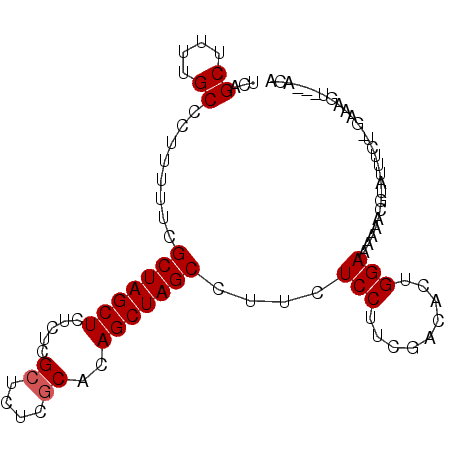
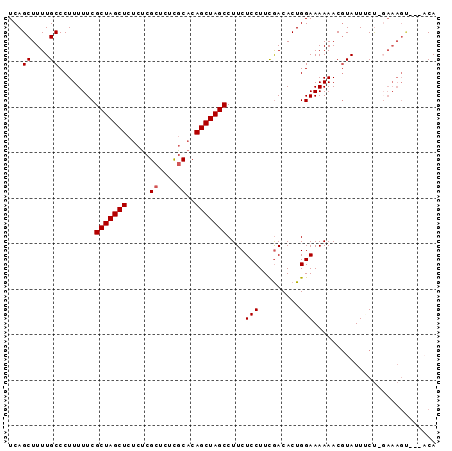
>2R_DroMel_CAF1 17073638 91 - 20766785 UCAGCUUUUGCCCUUUUUCGCUAGCUCUCUCGCUCUCACCCAGCUAGCCUUCUCCUUCGACACUGGAAAAAACGUAUUACUGGAAAGU---ACA ((((.(..(((..(((((((((((((...............)))))))..((......)).....))))))..)))..))))).....---... ( -15.36) >DroSec_CAF1 11169 91 - 1 UCAGCUUUUGCCCUUUUUCGCUAGCUCUCUCGCUCUCGCACAGCUAGCCUUCUCCUUUGACACUGGAAAAAACGUAUUUCUUGAAAGU---ACA ...((....))..(((((((((((((.....((....))..)))))))..((......)).....))))))..((((((.....))))---)). ( -16.80) >DroSim_CAF1 11267 91 - 1 UCAGCUUUUGCCCUUUUUCGCUAGCUCUCUCGCUCUCGCACAGCUAGCCUUCUCCUUCGACACUGGAAAAAACGUAUUUCUUGAAAGC---ACA ...((((((..........(((((((.....((....))..)))))))....(((.........)))...............))))))---... ( -18.30) >DroEre_CAF1 11507 86 - 1 UCAGCUUUUGCCCUUUUUCGCUAGCUCUCUCGCUCUUGCACAGCUAGCCUUCUCCUUCGACACUGGAAAAAACGUAUUGCC--------ACCCA ...((...(((..(((((((((((((.....((....))..)))))))..((......)).....))))))..)))..)).--------..... ( -16.80) >DroYak_CAF1 12818 92 - 1 UCAGCUUUUGCCCUUUUUCGCUAGCUCUCUCGCUCUUGCACAGCUAGCCUUCUCCUUCAACACUGGAAAAAACUUAUUCCU--AAAGUAAUCCA ...(((((...........(((((((.....((....))..)))))))................((((........)))).--)))))...... ( -15.50) >consensus UCAGCUUUUGCCCUUUUUCGCUAGCUCUCUCGCUCUCGCACAGCUAGCCUUCUCCUUCGACACUGGAAAAAACGUAUUUCU_GAAAGU___ACA ...((....))........(((((((.....((....))..)))))))....(((.........)))........................... (-11.88 = -12.08 + 0.20)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:40:03 2006