| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 17,001,099 – 17,001,190 |
| Length | 91 |
| Max. P | 0.987801 |

| Location | 17,001,099 – 17,001,190 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 69.97 |
| Mean single sequence MFE | -23.30 |
| Consensus MFE | -7.24 |
| Energy contribution | -8.16 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.31 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.930131 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17001099 91 + 20766785 AGGAGCUGUGUUCGCAGCUUGAGGCUGAAAAGCAAGAAGUGGCUGAAAAGGAUUUCUCGCACAAGGCGAACAGA---AGAUGGCGAAUGAGGAG---- ....(((((.(((.(((((....(((....))).......)))))...........((((.....))))...))---).)))))..........---- ( -24.70) >DroPse_CAF1 113422 77 + 1 ACAAGCUG--UUUGCAGCCAGAGGC------------AGUGGCUGAAAACAGAUUUUUGCAUAGC---AACAAAAGAAAACUCUUUGCGAGGGG---- .....(((--(((.((((((.....------------..)))))).)))))).((((((((....---.....((((....)))))))))))).---- ( -20.60) >DroSec_CAF1 97471 94 + 1 AGGAGCUGUGUUCGCAGCUUGAGGCUGAAAAGCAAGAAGUGCCUGAAAAGGAUUUUUCGCACAAGGCGAACAGAAGAAGAUGGCGAAUGAGGAG---- ....(((((((((((..((((..(((....)))..((((..(((....)))...))))...)))))))))))........))))..........---- ( -25.90) >DroSim_CAF1 104340 94 + 1 AGGAGCUGUGUUCGCAGCUUGAGGCUGAAAAGCAAGAAGUGGCUGAAAAGGAUUUUUCGCACAAGGCGAACAGAAGAAGAUGCCGAAUGAGGAG---- ..(((((((....)))))))...(((....)))......((((............(((((.....)))))...........)))).........---- ( -23.80) >DroPer_CAF1 112232 81 + 1 ACAAGCUG--UUUGCAGCCAGAGGC------------AGUGGCUGAAAACAGAUUUUUGCAUAGC---AACAAAAGAAAACUCUUUGCGAGGGGCACC .....(((--(((.((((((.....------------..)))))).)))))).....(((...((---((....(((....))))))).....))).. ( -21.50) >consensus AGGAGCUGUGUUCGCAGCUUGAGGCUGAAAAGCAAGAAGUGGCUGAAAAGGAUUUUUCGCACAAGGCGAACAGAAGAAGAUGCCGAAUGAGGAG____ ..(((((((....)))))))..................(((.(.(((((....))))))))).................................... ( -7.24 = -8.16 + 0.92)

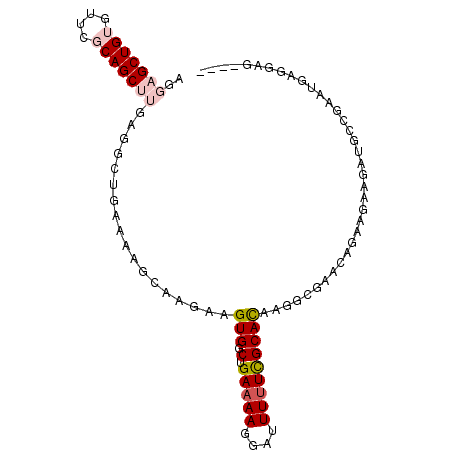
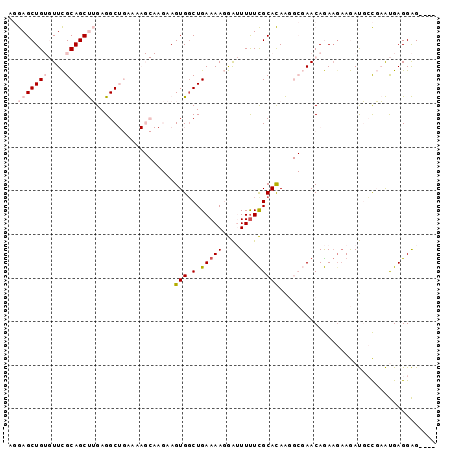
| Location | 17,001,099 – 17,001,190 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 69.97 |
| Mean single sequence MFE | -19.24 |
| Consensus MFE | -5.44 |
| Energy contribution | -6.24 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.28 |
| SVM decision value | 2.09 |
| SVM RNA-class probability | 0.987801 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 17001099 91 - 20766785 ----CUCCUCAUUCGCCAUCU---UCUGUUCGCCUUGUGCGAGAAAUCCUUUUCAGCCACUUCUUGCUUUUCAGCCUCAAGCUGCGAACACAGCUCCU ----.................---..(((((((((((.(((((((....)))))(((........))).....))..))))..)))))))........ ( -18.00) >DroPse_CAF1 113422 77 - 1 ----CCCCUCGCAAAGAGUUUUCUUUUGUU---GCUAUGCAAAAAUCUGUUUUCAGCCACU------------GCCUCUGGCUGCAAA--CAGCUUGU ----...(((.....))).....((((((.---.....))))))..((((((.((((((..------------.....)))))).)))--)))..... ( -20.00) >DroSec_CAF1 97471 94 - 1 ----CUCCUCAUUCGCCAUCUUCUUCUGUUCGCCUUGUGCGAAAAAUCCUUUUCAGGCACUUCUUGCUUUUCAGCCUCAAGCUGCGAACACAGCUCCU ----......................(((((((((((.(((((((....)))))(((((.....)))))....))..))))..)))))))........ ( -20.00) >DroSim_CAF1 104340 94 - 1 ----CUCCUCAUUCGGCAUCUUCUUCUGUUCGCCUUGUGCGAAAAAUCCUUUUCAGCCACUUCUUGCUUUUCAGCCUCAAGCUGCGAACACAGCUCCU ----......................(((((((((((.(((((((....)))))(((........))).....))..))))..)))))))........ ( -17.90) >DroPer_CAF1 112232 81 - 1 GGUGCCCCUCGCAAAGAGUUUUCUUUUGUU---GCUAUGCAAAAAUCUGUUUUCAGCCACU------------GCCUCUGGCUGCAAA--CAGCUUGU ..(((.....((((.(((....)))...))---))...))).....((((((.((((((..------------.....)))))).)))--)))..... ( -20.30) >consensus ____CUCCUCAUUCGGCAUCUUCUUCUGUUCGCCUUGUGCGAAAAAUCCUUUUCAGCCACUUCUUGCUUUUCAGCCUCAAGCUGCGAACACAGCUCCU ............................(((((.....)))))....................................(((((......)))))... ( -5.44 = -6.24 + 0.80)


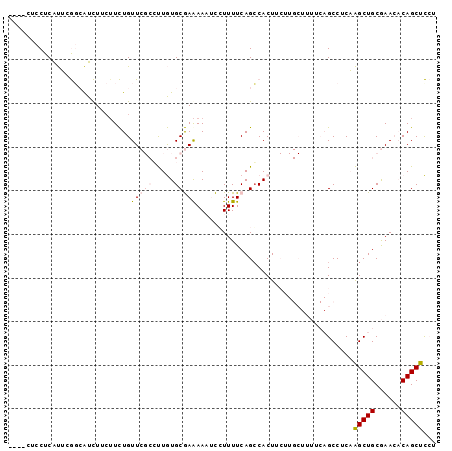
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:30 2006