
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 2,800,660 – 2,800,768 |
| Length | 108 |
| Max. P | 0.751995 |

| Location | 2,800,660 – 2,800,768 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.08 |
| Mean single sequence MFE | -35.87 |
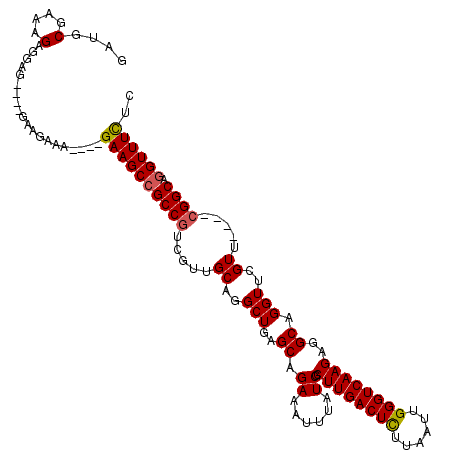
| Consensus MFE | -30.56 |
| Energy contribution | -30.62 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.751995 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
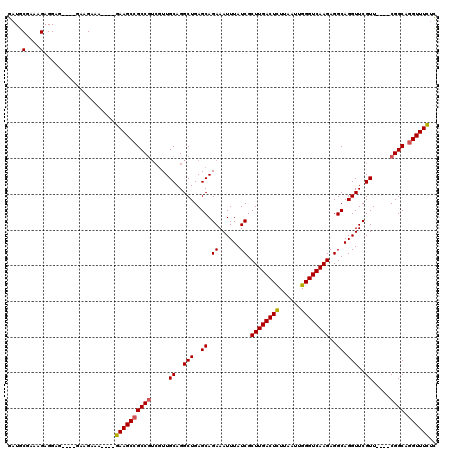
>2R_DroMel_CAF1 2800660 108 + 20766785 GAUGCGAAAGAGGAG----GAAGAAA----GAAGCCGCCGUCGUUGCAGGCUGAGCAGAAAUUUAUCGCUUGACUCUUAAUUGGGUCAAGAGGCAGGUUCGUU----CGGCAGGUUUCUC ....(....).....----......(----((((((((((((......))).((((.(((.(((..(.((((((((......)))))))).)..)))))))))----)))).))))))). ( -36.00) >DroSec_CAF1 7506 108 + 1 GAUGCAAAAGAGGUG----GAAGAAA----GAAGCCGCCGUCGUUGCAGGCUGAGCAGAAAUUUAUCGCUUGACUCUUAAUUGGGUCAAGAGGCAGGUUCGUU----CGGCAGGUUUCCC ..(((((....((((----(......----....)))))....))))).(((((((.(((.(((..(.((((((((......)))))))).)..)))))))))----))))......... ( -38.90) >DroSim_CAF1 7554 108 + 1 GAUGCGAAAGAGGUG----GAAGAAA----GAAGCCGCCAUCGUUGCAGGCUGAGCAGAAAUUUAUCGCUUGACUCUUAAUUGGGUCAAGAGGCAGGUUCGUU----CGGCAGGUUUCUC ..(((((..((((((----(......----....))))).)).))))).(((((((.(((.(((..(.((((((((......)))))))).)..)))))))))----))))......... ( -37.60) >DroEre_CAF1 7772 108 + 1 GAUGCGAAAGAGGAG----GAAGAAA----GAAGCCGCCGUCGUUGCAGGCUGAGCAGAAAUUUAUCGCUUGACUCUUAAUUGGGUCAAGAGGCAGGUUCGUU----CGGCAGGUUUCUC ....(....).....----......(----((((((((((((......))).((((.(((.(((..(.((((((((......)))))))).)..)))))))))----)))).))))))). ( -36.00) >DroYak_CAF1 7501 112 + 1 GAUGCGAAAGAGGAG----GAAGAAAGAGAGAAGCCGCCGUCGUUGCAGGCUGAGCAGAAAUUUAUCGCUUGACUCUUAAUUGGGUCAAGAGGCAGGUUCGUU----CGGCAUGUUUCUC ....(....).....----.........(((((((.((((((......))).((((.(((.(((..(.((((((((......)))))))).)..)))))))))----))))..))))))) ( -34.60) >DroAna_CAF1 7308 115 + 1 UAUUCGAAAGAGGAGAAGAGAAGAAA----GAAGCCGCCGA-AUUGCAGGCUGAGCAGAAAUUUAUCGCUUGACUUUUUAUUGGGUCAAGAGGCAGGUUCGUUCCUCCGGCAGGUUUUUA ..(((....))).............(----((((((((((.-...((..(((..((.((......)).((((((((......))))))))..)).)))..)).....)))).))))))). ( -32.10) >consensus GAUGCGAAAGAGGAG____GAAGAAA____GAAGCCGCCGUCGUUGCAGGCUGAGCAGAAAUUUAUCGCUUGACUCUUAAUUGGGUCAAGAGGCAGGUUCGUU____CGGCAGGUUUCUC ....(....)....................((((((((((.....((..(((..((.((......)).((((((((......))))))))..)).)))..)).....)))).)))))).. (-30.56 = -30.62 + 0.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:40:49 2006