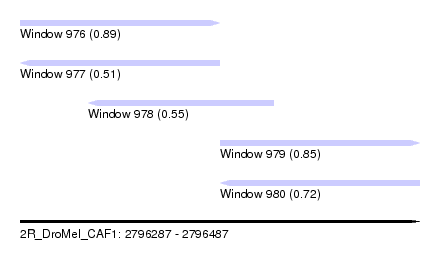

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 2,796,287 – 2,796,487 |
| Length | 200 |
| Max. P | 0.887025 |

| Location | 2,796,287 – 2,796,387 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 82.76 |
| Mean single sequence MFE | -34.34 |
| Consensus MFE | -20.13 |
| Energy contribution | -20.65 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.887025 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
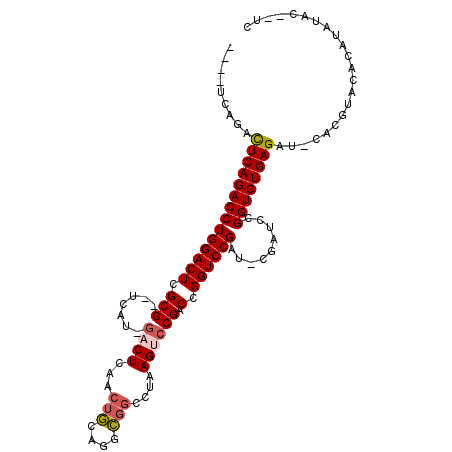
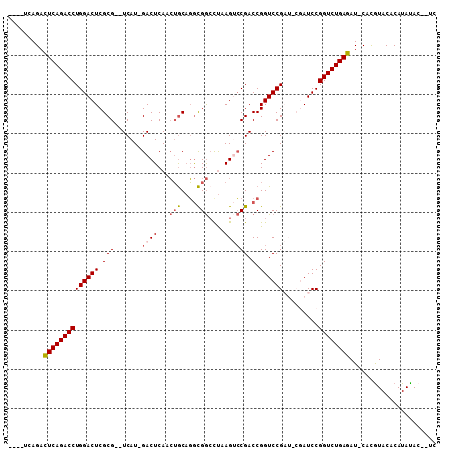
>2R_DroMel_CAF1 2796287 100 + 20766785 ---CUCAGACUCAGACCUGGACUCGCG--UCAU-GACUCAACUGCAGGCGGCCUAAGUCCGACCGGUCCGAUCCGAUCCGGUCUGAGAU-CACGUACACAUAUAC--UC ---....((((((((((.((((.((.(--((..-((((...(((....)))....)))).)))))))))(((...))).)))))))).)-)..............--.. ( -31.60) >DroSec_CAF1 3241 98 + 1 -----CAGACUCAGACCUGGACUCGCG--UCAU-GACUCAACUGCAGGCGGCCUAAGUCCGACCGGUCCGAUGCGAUCCGGUCUGAGAU-CACGUACACAUAUAC--UC -----..((((((((((.(((.(((((--((..-((((........((((((....).))).)))))).)))))))))))))))))).)-)..............--.. ( -37.20) >DroSim_CAF1 3315 98 + 1 -----CAGACUCAGACCUGGACUCGCG--UCAU-GACUCAACUGCAGGCGGGCUAAGUCCGACCGGUCCGAUGCGAUCCGGUCUGAGAU-CACGUACACAUAUAC--UC -----..((((((((((.(((.(((((--((..-((((........(((((((...))))).)))))).)))))))))))))))))).)-)..............--.. ( -40.50) >DroEre_CAF1 3226 100 + 1 --UAUCAGACUCAGACCUGGACUCGCGACUCAUGGACUCAUGGACUCAUGACUCAAGAGCGACCGGUCCG-----AUCCGGUCUGAGAU-CACGUACACAUACAAAUG- --.....((((((((((((((((.(((.(((...((.(((((....))))).))..))))).).))))))-----....)))))))).)-).................- ( -34.40) >DroYak_CAF1 3292 101 + 1 UGUCUCAGAUUCAGACCUGGACUCGCG--UCAU-GACUCAACUGCAGGCGGCUUAAGACCGACCGGUCCG-----AUCCGGUCUGAGAUACAUAUACACAUAUACAUUU (((((((((.......(((((.(((((--((.(-(.(......)))))))......((((....))))))-----)))))))))))))))................... ( -28.01) >consensus ____UCAGACUCAGACCUGGACUCGCG__UCAU_GACUCAACUGCAGGCGGCCUAAGUCCGACCGGUCCGAU_CGAUCCGGUCUGAGAU_CACGUACACAUAUAC__UC .........((((((((((((((.(((.......((((...(((....)))....)))))).).)))))).........))))))))...................... (-20.13 = -20.65 + 0.52)

| Location | 2,796,287 – 2,796,387 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 82.76 |
| Mean single sequence MFE | -38.14 |
| Consensus MFE | -20.86 |
| Energy contribution | -21.46 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.55 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.510044 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
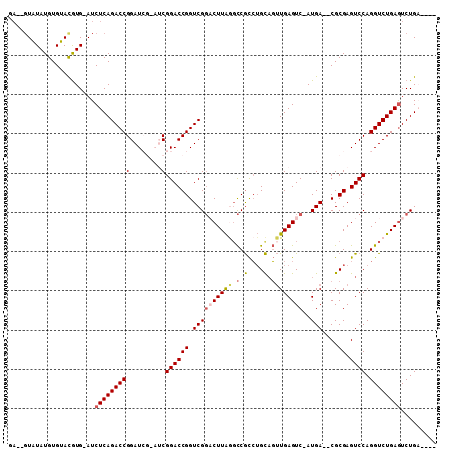
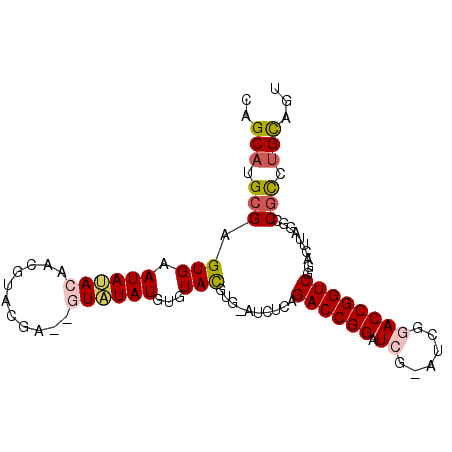
>2R_DroMel_CAF1 2796287 100 - 20766785 GA--GUAUAUGUGUACGUG-AUCUCAGACCGGAUCGGAUCGGACCGGUCGGACUUAGGCCGCCUGCAGUUGAGUC-AUGA--CGCGAGUCCAGGUCUGAGUCUGAG--- ..--((((....))))...-..(((((((....(((((((((((((((((((((((.((.....))...))))))-.)))--).)).)))).))))))))))))))--- ( -39.70) >DroSec_CAF1 3241 98 - 1 GA--GUAUAUGUGUACGUG-AUCUCAGACCGGAUCGCAUCGGACCGGUCGGACUUAGGCCGCCUGCAGUUGAGUC-AUGA--CGCGAGUCCAGGUCUGAGUCUG----- ((--(..((((....))))-..))).((((((.(((...))).))))))((((((((((((....).........-..((--(....)))..))))))))))).----- ( -38.40) >DroSim_CAF1 3315 98 - 1 GA--GUAUAUGUGUACGUG-AUCUCAGACCGGAUCGCAUCGGACCGGUCGGACUUAGCCCGCCUGCAGUUGAGUC-AUGA--CGCGAGUCCAGGUCUGAGUCUG----- ..--((((....))))..(-(.((((((((.(((...)))((((((((((((((((((..(....).))))))))-.)))--).)).)))).))))))))))..----- ( -39.40) >DroEre_CAF1 3226 100 - 1 -CAUUUGUAUGUGUACGUG-AUCUCAGACCGGAU-----CGGACCGGUCGCUCUUGAGUCAUGAGUCCAUGAGUCCAUGAGUCGCGAGUCCAGGUCUGAGUCUGAUA-- -......((((....))))-...(((((((((((-----(((((..((.((((..((.(((((....))))).))...)))).))..)))).)))))).))))))..-- ( -38.50) >DroYak_CAF1 3292 101 - 1 AAAUGUAUAUGUGUAUAUGUAUCUCAGACCGGAU-----CGGACCGGUCGGUCUUAAGCCGCCUGCAGUUGAGUC-AUGA--CGCGAGUCCAGGUCUGAAUCUGAGACA ..(((((((....))))))).(((((((.(((((-----((((((((((((.(((((((.....))..))))).)-.)))--).)).)))).))))))..))))))).. ( -34.70) >consensus GA__GUAUAUGUGUACGUG_AUCUCAGACCGGAUCG_AUCGGACCGGUCGGACUUAGGCCGCCUGCAGUUGAGUC_AUGA__CGCGAGUCCAGGUCUGAGUCUGA____ ......................(((((((((........)((((((.((((((((((.(.(....).))))))))..)))....)).)))).))))))))......... (-20.86 = -21.46 + 0.60)
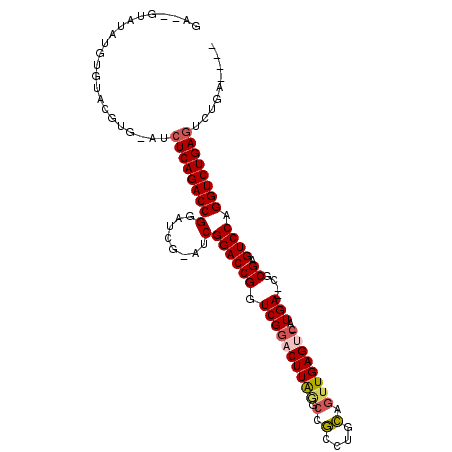


| Location | 2,796,321 – 2,796,414 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 80.79 |
| Mean single sequence MFE | -29.92 |
| Consensus MFE | -19.52 |
| Energy contribution | -20.20 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.546465 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 2796321 93 - 20766785 CAGCAUGCGAGUGAAUACACAACGUACGA--GUAUAUGUGUACGUG-AUCUCAGACCGGAUCGGAUCGGACCGGUCGGACUUAGGCCGCCUGCAGU ..(((.(((.(((....))).(((((((.--.......))))))).-...((.((((((.(((...))).)))))).)).......))).)))... ( -29.90) >DroSec_CAF1 3273 93 - 1 CAGCAUGCGAGUGAAUAUACAACGUACGA--GUAUAUGUGUACGUG-AUCUCAGACCGGAUCGCAUCGGACCGGUCGGACUUAGGCCGCCUGCAGU ..(((.(((.((.........(((((((.--.......))))))).-...((.((((((.(((...))).)))))).)).....))))).)))... ( -29.80) >DroSim_CAF1 3347 93 - 1 CAGCAUGCGAGUGAAUAUACAACGUACGA--GUAUAUGUGUACGUG-AUCUCAGACCGGAUCGCAUCGGACCGGUCGGACUUAGCCCGCCUGCAGU ..(((.(((.((.........(((((((.--.......))))))).-...((.((((((.(((...))).)))))).))....)).))).)))... ( -29.90) >DroEre_CAF1 3264 82 - 1 CAGCAUGCGAGUGAAUACAC--------CAUUUGUAUGUGUACGUG-AUCUCAGACCGGAU-----CGGACCGGUCGCUCUUGAGUCAUGAGUCCA ..(((((((((((.......--------)))))))))))...((((-(.((((((((((..-----....)))))).....)))))))))...... ( -30.80) >DroYak_CAF1 3329 90 - 1 CAGCAUGCGAGUGAAUAUAA-AUGUAUAAAUGUAUAUGUGUAUAUGUAUCUCAGACCGGAU-----CGGACCGGUCGGUCUUAAGCCGCCUGCAGU ..(((.(((((...(((((.-(((((((....))))))).)))))....))).((((((..-----....))))))(((.....))))).)))... ( -29.20) >consensus CAGCAUGCGAGUGAAUAUACAACGUACGA__GUAUAUGUGUACGUG_AUCUCAGACCGGAUCG_AUCGGACCGGUCGGACUUAGGCCGCCUGCAGU ..(((.(((.(((.((((((...........))))))...)))..........((((((.((......))))))))..........))).)))... (-19.52 = -20.20 + 0.68)

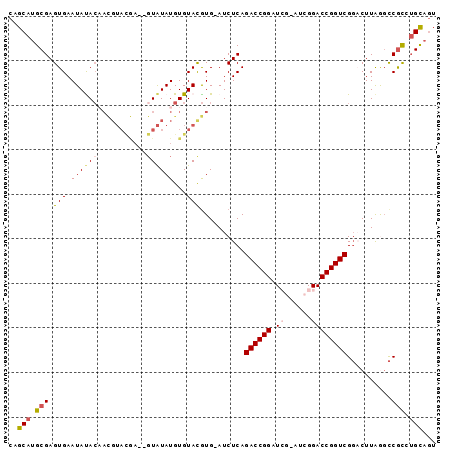
| Location | 2,796,387 – 2,796,487 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 88.66 |
| Mean single sequence MFE | -20.60 |
| Consensus MFE | -16.42 |
| Energy contribution | -17.10 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.850324 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 2796387 100 + 20766785 GUACGUUGUGUAUUCACUCGCAUGCUGCAGAAGCAACCUUUACCUUUCCUUUCCAUUCGCUACCUC------CGUUGCCGCUGCUCUGGAUGCUUUCACUUUCCAU .......(((....)))..((((.(.((((..(((((.............................------.)))))..))))...).))))............. ( -17.91) >DroSec_CAF1 3339 100 + 1 GUACGUUGUAUAUUCACUCGCAUGCUGCAGAGGCAACCUUUACCUUUCCUUUCCAUUCGCUACCUC------CGUUGCCGCUGCUCUGGAUGCUUUCACUUUCCAU ...................((((.(.((((.((((((.............................------.)))))).))))...).))))............. ( -20.61) >DroSim_CAF1 3413 100 + 1 GUACGUUGUAUAUUCACUCGCAUGCUGCAGAGGCAACCUUUACCUUUCCUUUCCAUUCGCUACCUC------CGUCGCCGCUGCUCUGGAUGCUUUCACUUUCCAU ...................((((.(.((((.(((.((.............................------.)).))).))))...).))))............. ( -16.51) >DroEre_CAF1 3326 93 + 1 -------GUGUAUUCACUCGCAUGCUGCAGCGGCAACCUUUACCUUUCCUUUCCAUUUGCUAUCUC------CGCUGCCGCUGCUCUGGAUGCUUUCACUUUCCAG -------(((....)))..((((.(.(((((((((...............................------...)))))))))...).))))............. ( -24.08) >DroYak_CAF1 3393 105 + 1 AUACAU-UUAUAUUCACUCGCAUGCUGCAGAGGCAACCUUUGCCUUUCCUUUCUAUUUGCUAUCUCCAGUGCCGCUGCUGCUGCUCUGGAUGCUUUCACUUUCCAU ......-............(((....(((((((...))))))).....................(((((.((.((....)).)).))))))))............. ( -23.90) >consensus GUACGUUGUAUAUUCACUCGCAUGCUGCAGAGGCAACCUUUACCUUUCCUUUCCAUUCGCUACCUC______CGUUGCCGCUGCUCUGGAUGCUUUCACUUUCCAU ...................((((.(.((((.((((((....................................)))))).))))...).))))............. (-16.42 = -17.10 + 0.68)

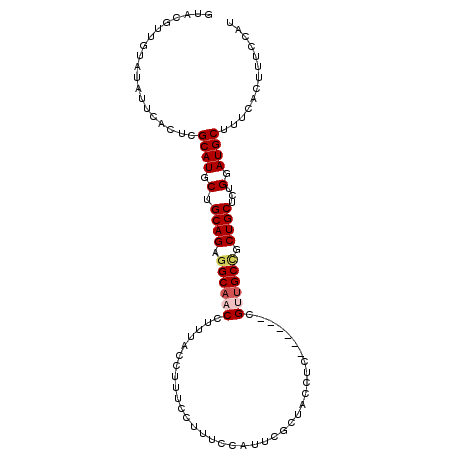
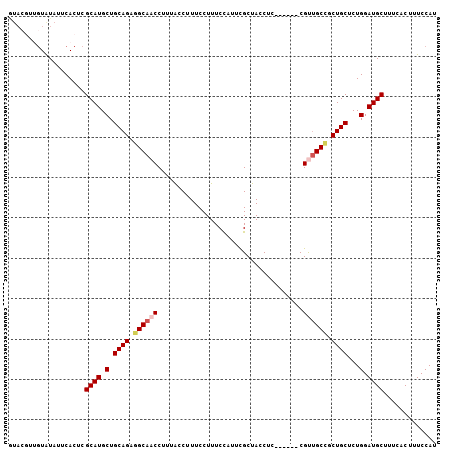
| Location | 2,796,387 – 2,796,487 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 88.66 |
| Mean single sequence MFE | -26.09 |
| Consensus MFE | -18.82 |
| Energy contribution | -18.50 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.719820 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 2796387 100 - 20766785 AUGGAAAGUGAAAGCAUCCAGAGCAGCGGCAACG------GAGGUAGCGAAUGGAAAGGAAAGGUAAAGGUUGCUUCUGCAGCAUGCGAGUGAAUACACAACGUAC .((((..((....)).))))..(((...((..((------((((((((.....................))))))))))..)).)))..(((....)))....... ( -24.60) >DroSec_CAF1 3339 100 - 1 AUGGAAAGUGAAAGCAUCCAGAGCAGCGGCAACG------GAGGUAGCGAAUGGAAAGGAAAGGUAAAGGUUGCCUCUGCAGCAUGCGAGUGAAUAUACAACGUAC .((((..((....)).))))..(((...((..((------((((((((.....................))))))))))..)).)))................... ( -25.60) >DroSim_CAF1 3413 100 - 1 AUGGAAAGUGAAAGCAUCCAGAGCAGCGGCGACG------GAGGUAGCGAAUGGAAAGGAAAGGUAAAGGUUGCCUCUGCAGCAUGCGAGUGAAUAUACAACGUAC .((((..((....)).))))..(((...((..((------((((((((.....................))))))))))..)).)))................... ( -27.10) >DroEre_CAF1 3326 93 - 1 CUGGAAAGUGAAAGCAUCCAGAGCAGCGGCAGCG------GAGAUAGCAAAUGGAAAGGAAAGGUAAAGGUUGCCGCUGCAGCAUGCGAGUGAAUACAC------- (((((..((....)).))))).(((((((((((.------...((......(....)......))....)))))))))))...................------- ( -29.60) >DroYak_CAF1 3393 105 - 1 AUGGAAAGUGAAAGCAUCCAGAGCAGCAGCAGCGGCACUGGAGAUAGCAAAUAGAAAGGAAAGGCAAAGGUUGCCUCUGCAGCAUGCGAGUGAAUAUAA-AUGUAU .((((..((....)).))))..(((((.((((.((((((.......((...............))...)).)))).)))).)).)))............-...... ( -23.56) >consensus AUGGAAAGUGAAAGCAUCCAGAGCAGCGGCAACG______GAGGUAGCGAAUGGAAAGGAAAGGUAAAGGUUGCCUCUGCAGCAUGCGAGUGAAUAUACAACGUAC .((((..((....)).))))..((((.((((((....................................)))))).)))).......................... (-18.82 = -18.50 + -0.32)

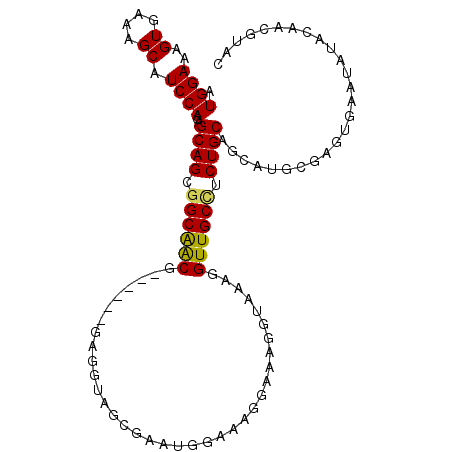
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:40:45 2006