
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 16,473,743 – 16,473,841 |
| Length | 98 |
| Max. P | 0.635895 |

| Location | 16,473,743 – 16,473,841 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 87.41 |
| Mean single sequence MFE | -31.12 |
| Consensus MFE | -20.32 |
| Energy contribution | -19.80 |
| Covariance contribution | -0.52 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.548879 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
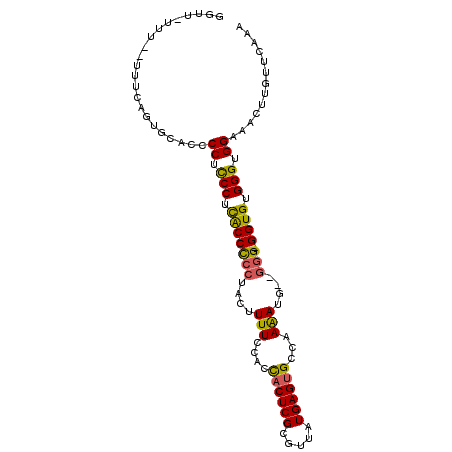
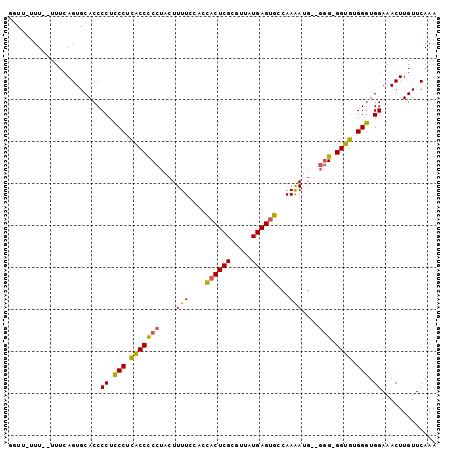
>2R_DroMel_CAF1 16473743 98 + 20766785 GAUUUUUU--UUUCAGUGCACCCCUUCCUCACCUCCUACUUUUCCGCCACUCGCGUUAUGAGUGCCAAAAUG--GGGCGGUGUGGGUGGAAACUUGUUCAAA ........--.....................((.(((((....((((((((((.....))))).((.....)--)))))).))))).))............. ( -25.50) >DroSec_CAF1 148958 96 + 1 GGUU-UUU--UUUCAGUGCACCCCUUCCGCACCCCCUACUUUUCCACCACUCGCGUUAUGAGUGCCAAAAUG--GUG-GGUGUGGGUGGAAACUUGUUCAAA ....-...--............((.(((((((((((...((((....((((((.....))))))..)))).)--).)-)))))))).))............. ( -27.00) >DroSim_CAF1 145714 97 + 1 GGUAAUUU--UUUCAGUGCACCCCUCCCACGCCCCCUACUUUUCCACCACUCGCGUUAUGAGCGCCAAAAUG--GGG-GGUGUGGGUGGAAACUUGUUCAAA (((.(((.--....)))..)))((.(((((((((((((.((((....(.((((.....)))).)..))))))--)))-)))))))).))............. ( -33.90) >DroEre_CAF1 150372 97 + 1 UGCU-UUU--UUUCAGUGCACCCCUCCCUUACCCCCUUCUUUUCCGCUACUCGCGUUAUGAGUGCCAAAA-GAGGGG-GGUGUGGGUGGAAACUUGUUCAAA (((.-...--.......)))..((.(((.((((((((((((((..((.(((((.....))))))).))))-))))))-)))).))).))............. ( -38.70) >DroYak_CAF1 150982 99 + 1 UGCC-UUUUCUUUGAGUGCACCCCUCCCUUACCCCCCACUUUUCC-CCACUCGCGUUAUGAGUGCCAAGAUGAGAGG-GGUGUGGGUGGAAACUUGUUCAAA ((((-((......))).)))..((.(((.((((((...(((.((.-.((((((.....))))))....)).))).))-)))).))).))............. ( -30.50) >consensus GGUU_UUU__UUUCAGUGCACCCCUCCCUCACCCCCUACUUUUCCACCACUCGCGUUAUGAGUGCCAAAAUG__GGG_GGUGUGGGUGGAAACUUGUUCAAA ......................((.(((.(((((((....(((....((((((.....))))))...)))....))).)))).))).))............. (-20.32 = -19.80 + -0.52)

| Location | 16,473,743 – 16,473,841 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 87.41 |
| Mean single sequence MFE | -31.58 |
| Consensus MFE | -21.46 |
| Energy contribution | -22.98 |
| Covariance contribution | 1.52 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.635895 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
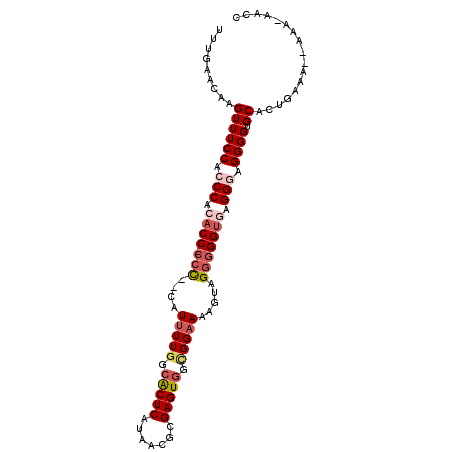
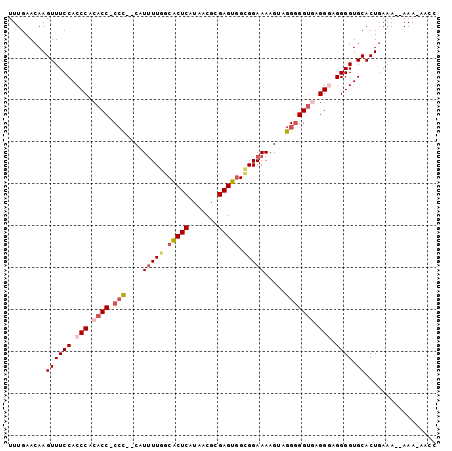
>2R_DroMel_CAF1 16473743 98 - 20766785 UUUGAACAAGUUUCCACCCACACCGCCC--CAUUUUGGCACUCAUAACGCGAGUGGCGGAAAAGUAGGAGGUGAGGAAGGGGUGCACUGAAA--AAAAAAUC ........(((...(((((.(..(((((--(.(((((.(((((.......))))).))))).....)).))))..)...))))).)))....--........ ( -27.60) >DroSec_CAF1 148958 96 - 1 UUUGAACAAGUUUCCACCCACACC-CAC--CAUUUUGGCACUCAUAACGCGAGUGGUGGAAAAGUAGGGGGUGCGGAAGGGGUGCACUGAAA--AAA-AACC ........(((...(((((.((((-(.(--((((((..((((....((....))))))..))))).)))))))(....)))))).)))....--...-.... ( -29.60) >DroSim_CAF1 145714 97 - 1 UUUGAACAAGUUUCCACCCACACC-CCC--CAUUUUGGCGCUCAUAACGCGAGUGGUGGAAAAGUAGGGGGCGUGGGAGGGGUGCACUGAAA--AAAUUACC .........((.(((.(((((.((-(((--.(((((..((((....((....))))))..))))).))))).)))))..))).)).......--........ ( -35.60) >DroEre_CAF1 150372 97 - 1 UUUGAACAAGUUUCCACCCACACC-CCCCUC-UUUUGGCACUCAUAACGCGAGUAGCGGAAAAGAAGGGGGUAAGGGAGGGGUGCACUGAAA--AAA-AGCA .........((.(((.(((..(((-(((.((-((((.((((((.......)))).)).).))))).))))))..)))..))).)).......--...-.... ( -34.10) >DroYak_CAF1 150982 99 - 1 UUUGAACAAGUUUCCACCCACACC-CCUCUCAUCUUGGCACUCAUAACGCGAGUGG-GGAAAAGUGGGGGGUAAGGGAGGGGUGCACUCAAAGAAAA-GGCA (((((......((((.(((..(((-((((.(.((((..(((((.......))))))-)))...).)))))))..))).)))).....))))).....-.... ( -31.00) >consensus UUUGAACAAGUUUCCACCCACACC_CCC__CAUUUUGGCACUCAUAACGCGAGUGGCGGAAAAGUAGGGGGUGAGGGAGGGGUGCACUGAAA__AAA_AACC .........((((((.(((.((((.(((....(((((.(((((.......))))).))))).....))))))).))).)))).))................. (-21.46 = -22.98 + 1.52)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:35:50 2006