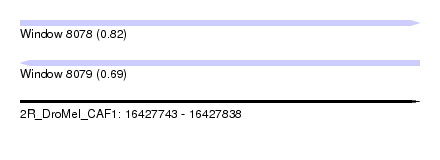

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 16,427,743 – 16,427,838 |
| Length | 95 |
| Max. P | 0.822361 |

| Location | 16,427,743 – 16,427,838 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 77.17 |
| Mean single sequence MFE | -25.42 |
| Consensus MFE | -21.43 |
| Energy contribution | -21.43 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.12 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.822361 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
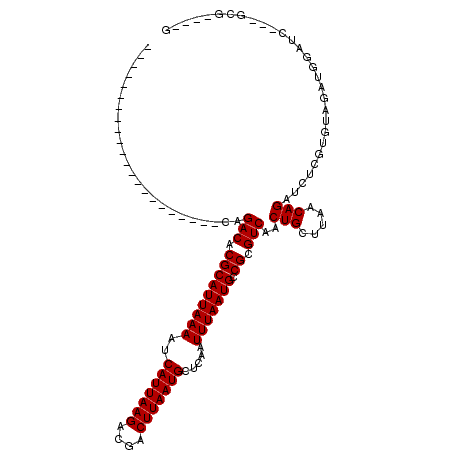
>2R_DroMel_CAF1 16427743 95 + 20766785 CCU-UCACGACGAUCCAUCUACACGAGAUCUGUUAAGCAGUUGACGCGGCAUUAAAUUGAGCAUUAAGGCGUCUUAAUGAUUUUAAUGCGUGUCUG---------------------- ...-....((((((((........).)))).)))........((((((.(((((((.....(((((((....)))))))..)))))))))))))..---------------------- ( -24.20) >DroPse_CAF1 132564 115 + 1 CAAGCCGC---GAUCCAUCUACACGAGAUCUGUUAAGCAGUUGACGCGGCAUUAAAUUGAGCAUUAAGUCAUCUUAAUGAUUUUAAUGCGUGUCCGUGUCCGAGUCUCUGUCCGUAUC ........---.........(((.((((((((.....)))..((((((((((...(((((.(((((((....)))))))...)))))..))).)))))))...))))))))....... ( -29.30) >DroWil_CAF1 244301 85 + 1 ------UC---GAUCCAUCUAUGC--GACCUGUUAAGCAGUUGACGCGGCAUUAAAUUGAGCAUUAAGUCGCCUUAAUGAUUUUAAUGCGUGUCCG---------------------- ------..---..........(((--..........)))...((((((.(((((((.....(((((((....)))))))..)))))))))))))..---------------------- ( -21.50) >DroYak_CAF1 103883 92 + 1 CCU-UCAC---GAUCCAUCUACACGAGAUCUGUUCGCCAGUUGACGCGGCAUUAAAUUGAGCAUUAAGGCGUCUUAAUGAUUUUAAUGCGUGUCUG---------------------- ...-...(---((...((((.....))))....)))......((((((.(((((((.....(((((((....)))))))..)))))))))))))..---------------------- ( -23.20) >DroAna_CAF1 98509 84 + 1 --------------CCGUCUACACGAGAUCUGUUAAGCAGUUGACGCGGCAUUAAAGUGAGCAUUAAGUCGUCUUAAUGAUUUUAAUGCGUGUCUGUG-------------------- --------------.......((((....(((.....)))..((((((.(((((((((...(((((((....))))))))))))))))))))))))))-------------------- ( -25.00) >DroPer_CAF1 133628 115 + 1 CAAGCCGC---GAUCCAUCUACACGAGAUCUGUUAAGCAGUUGACGCGGCAUUAAAUUGAGCAUUAAGUCAUCUUAAUGAUUUUAAUGCGUGUCCGUGUCCGAGUCUCUGUCCGUAUC ........---.........(((.((((((((.....)))..((((((((((...(((((.(((((((....)))))))...)))))..))).)))))))...))))))))....... ( -29.30) >consensus C____CAC___GAUCCAUCUACACGAGAUCUGUUAAGCAGUUGACGCGGCAUUAAAUUGAGCAUUAAGUCGUCUUAAUGAUUUUAAUGCGUGUCCG______________________ .............................(((.....)))..((((((.(((((((.....(((((((....)))))))..)))))))))))))........................ (-21.43 = -21.43 + 0.00)


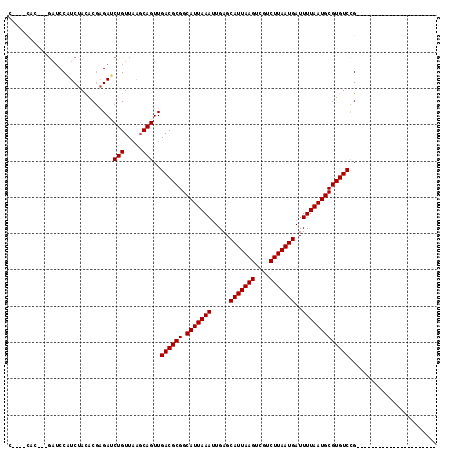
| Location | 16,427,743 – 16,427,838 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 77.17 |
| Mean single sequence MFE | -23.67 |
| Consensus MFE | -17.65 |
| Energy contribution | -17.65 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.02 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.690384 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 16427743 95 - 20766785 ----------------------CAGACACGCAUUAAAAUCAUUAAGACGCCUUAAUGCUCAAUUUAAUGCCGCGUCAACUGCUUAACAGAUCUCGUGUAGAUGGAUCGUCGUGA-AGG ----------------------..(((.(((((((((..(((((((....))))))).....))))))).)).))).....(((.((.((((.(((....)))))))...)).)-)). ( -22.70) >DroPse_CAF1 132564 115 - 1 GAUACGGACAGAGACUCGGACACGGACACGCAUUAAAAUCAUUAAGAUGACUUAAUGCUCAAUUUAAUGCCGCGUCAACUGCUUAACAGAUCUCGUGUAGAUGGAUC---GCGGCUUG ....((....((..(((..((((((((.(((((((((..(((((((....))))))).....))))))).)).)))..(((.....)))....))))).)).)..))---.))..... ( -27.40) >DroWil_CAF1 244301 85 - 1 ----------------------CGGACACGCAUUAAAAUCAUUAAGGCGACUUAAUGCUCAAUUUAAUGCCGCGUCAACUGCUUAACAGGUC--GCAUAGAUGGAUC---GA------ ----------------------..(((.(((((((((..(((((((....))))))).....))))))).)).)))............((((--.((....))))))---..------ ( -21.70) >DroYak_CAF1 103883 92 - 1 ----------------------CAGACACGCAUUAAAAUCAUUAAGACGCCUUAAUGCUCAAUUUAAUGCCGCGUCAACUGGCGAACAGAUCUCGUGUAGAUGGAUC---GUGA-AGG ----------------------....(((((((((((..(((((((....))))))).....))))))))..((((....))))....((((.(((....)))))))---))).-... ( -23.60) >DroAna_CAF1 98509 84 - 1 --------------------CACAGACACGCAUUAAAAUCAUUAAGACGACUUAAUGCUCACUUUAAUGCCGCGUCAACUGCUUAACAGAUCUCGUGUAGACGG-------------- --------------------(((.(((.(((((((((..(((((((....))))))).....))))))).)).)))..(((.....))).....))).......-------------- ( -19.20) >DroPer_CAF1 133628 115 - 1 GAUACGGACAGAGACUCGGACACGGACACGCAUUAAAAUCAUUAAGAUGACUUAAUGCUCAAUUUAAUGCCGCGUCAACUGCUUAACAGAUCUCGUGUAGAUGGAUC---GCGGCUUG ....((....((..(((..((((((((.(((((((((..(((((((....))))))).....))))))).)).)))..(((.....)))....))))).)).)..))---.))..... ( -27.40) >consensus ______________________CAGACACGCAUUAAAAUCAUUAAGACGACUUAAUGCUCAAUUUAAUGCCGCGUCAACUGCUUAACAGAUCUCGUGUAGAUGGAUC___GCG____G ........................(((.(((((((((..(((((((....))))))).....))))))).)).)))..(((.....)))............................. (-17.65 = -17.65 + 0.00)


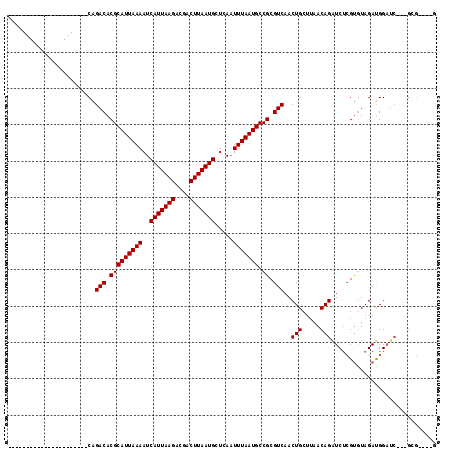
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:35:20 2006