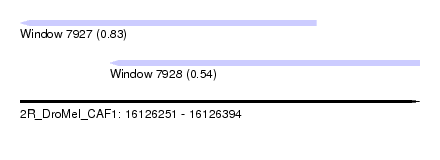

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 16,126,251 – 16,126,394 |
| Length | 143 |
| Max. P | 0.833929 |

| Location | 16,126,251 – 16,126,357 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 82.73 |
| Mean single sequence MFE | -36.70 |
| Consensus MFE | -26.52 |
| Energy contribution | -26.76 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.833929 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
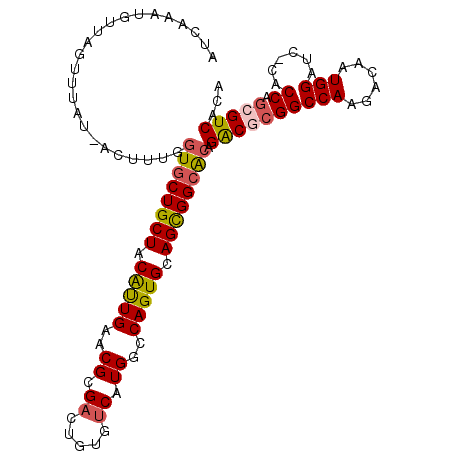
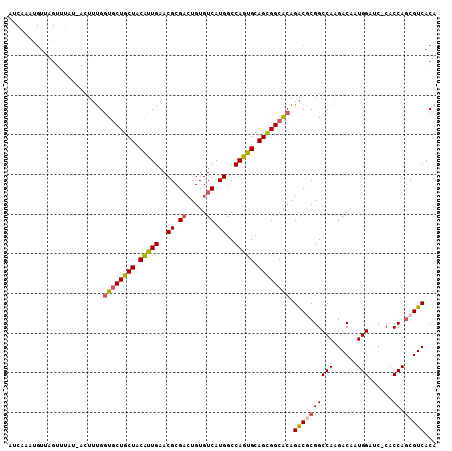
>2R_DroMel_CAF1 16126251 106 - 20766785 AUCCAAAGUUGGUUUAU-ACUUUGCUGCUGCUGCAUUGAACGCGAGUGUGUCAUGACCAGUGCAGCGGCGCAGACGCGGCCAAGACAAUGGAUC-CACCAGCGUCACA ........((((((...-..(((((.((((((((((((.((....))..((....)))))))))))))))))))...))))))(((..(((...-..)))..)))... ( -38.70) >DroSec_CAF1 15490 107 - 1 AUCAAAUGUUAGUUUUUCACUUUGGUGCUGCUACAUUGAACGCGACUGUGUCAUGGCCAGUGCAGCGGCACAGACGCGGCCAAGACAAUGGAAC-CACCAGCGUCACA ........................((((((((.(((((..((.(((...))).))..))))).)))))))).((((((((((......)))...-..)).)))))... ( -34.00) >DroSim_CAF1 15583 107 - 1 AUCAAAUGUUAGUUUUUCACUUUGGUACUGCUGCAUUGAACGCGUCUGUGUCAUGGCCAGUGCAGUGGCACAGACGCGGCCAAGACAAUGGAAC-CACCAGCGUCACA .....((((((((.....))).((((.((..((..(((..((((((((((((((.((....)).))))))))))))))..)))..))..)).))-))..))))).... ( -39.60) >DroEre_CAF1 16038 106 - 1 GUCAAAUGUUAGGUUAU-AUUUCGGUGCUGCUACGCUGAACGCGACUGUGUCAUGGCCAGUGCAGCGGCACAGACACGGCCAAGACAAUGGAUC-CACCAGCGUCACA (((((((((......))-))))..((((((((.(((((..((.(((...))).))..))))).)))))))).)))........(((..(((...-..)))..)))... ( -35.90) >DroYak_CAF1 15906 89 - 1 -------------------UGGUGGUGCUGCUACGCUGAACGCGACUGUGUCAUGGCCAGUGCAGCGGCACAGGCACGGCCAAGACAAUGGAUCCCACCAACGUCACA -------------------(((((((((((((.(((((..((.(((...))).))..))))).)))))))).(((...)))..............)))))........ ( -35.30) >consensus AUCAAAUGUUAGUUUAU_ACUUUGGUGCUGCUACAUUGAACGCGACUGUGUCAUGGCCAGUGCAGCGGCACAGACGCGGCCAAGACAAUGGAUC_CACCAGCGUCACA ........................((((((((.(((((..((.((.....)).))..))))).)))))))).((((((((((......)))......)).)))))... (-26.52 = -26.76 + 0.24)

| Location | 16,126,283 – 16,126,394 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 79.06 |
| Mean single sequence MFE | -28.76 |
| Consensus MFE | -21.62 |
| Energy contribution | -22.12 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.537503 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
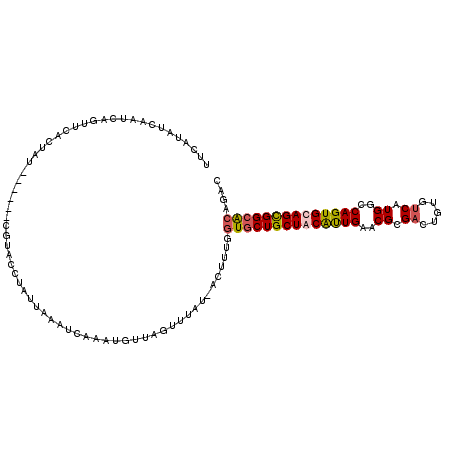
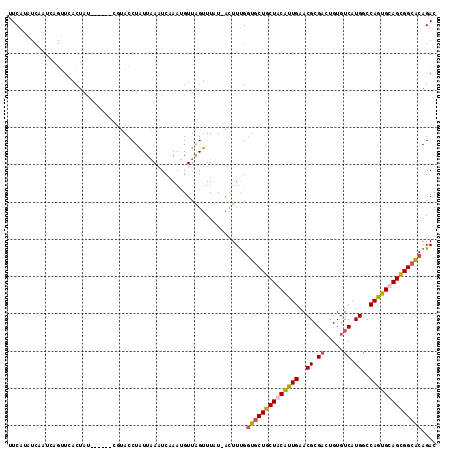
>2R_DroMel_CAF1 16126283 111 - 20766785 UUCAUAUCAAUCGGUUCACUAUGUA---CCUACCUAUUCAAUCCAAAGUUGGUUUAU-ACUUUGCUGCUGCUGCAUUGAACGCGAGUGUGUCAUGACCAGUGCAGCGGCGCAGAC ......................(((---...(((.(((........))).)))...)-))(((((.((((((((((((.((....))..((....))))))))))))))))))). ( -30.40) >DroSec_CAF1 15522 115 - 1 UUCAUAUCAAUUAGUUCACUACAUACUACAUACCUAUUAAAUCAAAUGUUAGUUUUUCACUUUGGUGCUGCUACAUUGAACGCGACUGUGUCAUGGCCAGUGCAGCGGCACAGAC .......(((.......(((((((.....................))).))))........)))((((((((.(((((..((.(((...))).))..))))).)))))))).... ( -24.16) >DroSim_CAF1 15615 108 - 1 UUCAUAUCAAUCAGUUCACUAU-------GUACCUAUUAAAUCAAAUGUUAGUUUUUCACUUUGGUACUGCUGCAUUGAACGCGUCUGUGUCAUGGCCAGUGCAGUGGCACAGAC ......(((((((((.......-------(((((.........((((....))))........))))).)))).)))))....(((((((((((.((....)).))))))))))) ( -30.03) >DroEre_CAF1 16070 104 - 1 UUCGAA--AAU-----UACGAU---CCUGGUACCUCUUAAGUCAAAUGUUAGGUUAU-AUUUCGGUGCUGCUACGCUGAACGCGACUGUGUCAUGGCCAGUGCAGCGGCACAGAC ..((((--(..-----((..(.---(((..((..............))..))))..)-))))))((((((((.(((((..((.(((...))).))..))))).)))))))).... ( -30.44) >consensus UUCAUAUCAAUCAGUUCACUAU______CGUACCUAUUAAAUCAAAUGUUAGUUUAU_ACUUUGGUGCUGCUACAUUGAACGCGACUGUGUCAUGGCCAGUGCAGCGGCACAGAC ................................................................((((((((((((((..((.((.....)).))..)))))))))))))).... (-21.62 = -22.12 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:32:57 2006