
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 16,117,220 – 16,117,312 |
| Length | 92 |
| Max. P | 0.869430 |

| Location | 16,117,220 – 16,117,312 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 79.59 |
| Mean single sequence MFE | -15.23 |
| Consensus MFE | -5.90 |
| Energy contribution | -6.90 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.39 |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.869430 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 16117220 92 - 20766785 UUUCUUGCCCAACUAAAUUCGCAACUUUAUUGCUGUACAAACAGCUCAUAAAAAACUAUAACACCCUCCGUUGCGGCAUAGAAAGUUUA-UUU (((((((((((((((((........))))..(((((....)))))........................)))).)))).))))).....-... ( -18.00) >DroSec_CAF1 6539 85 - 1 --------CUAACUAAAUUCGCAACUUUAUUGCUGUACAAACAGCUCAUAAAAAACUAUAGCACCCUCCGCUGCGGCAUAUAAAGUUUAAUUC --------..............((((((((.(((((....)))))..........(..((((.......))))..)...))))))))...... ( -14.30) >DroSim_CAF1 6617 85 - 1 --------CCAACUAAAUUCGCAACUUUAUUGCUGUACAAACAGCUCAUAAAAAACUAUAGCACCCUCUGCUGCGGCAUAGAAAGUUUAAUUC --------....(((...((((...(((((.(((((....)))))..))))).......((((.....))))))))..)))............ ( -15.10) >DroEre_CAF1 6745 84 - 1 CUUCCCGCUCAACUAAAUUCUCAACUUUAUUGCUGUACAAACAGGUCAUAAA-AACUAUAGAACCCUCUGCGGC--------AAGUUAUAUUC (((.((((.................(((((.(((((....))).)).)))))-......(((....))))))).--------)))........ ( -10.90) >DroYak_CAF1 6743 92 - 1 CUAUCCGUUCAACUUAAUUCCCAACUUGAUUGCUGUACAAAUAGCUCAUAAA-AACUAUAGCACCAUCCGCAGCAGUAUAGAAAGUUGAAUUC ......(((((((((......(.....)((((((((.......(((.(((..-...)))))).......)))))))).....))))))))).. ( -17.84) >consensus _U____G_CCAACUAAAUUCGCAACUUUAUUGCUGUACAAACAGCUCAUAAAAAACUAUAGCACCCUCCGCUGCGGCAUAGAAAGUUUAAUUC ......................((((((...(((((....)))))..........((.((((.......)))).)).....))))))...... ( -5.90 = -6.90 + 1.00)

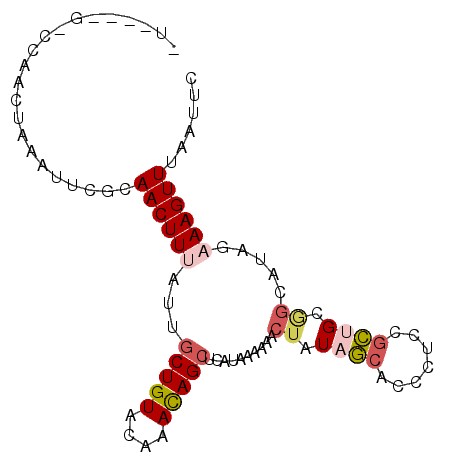
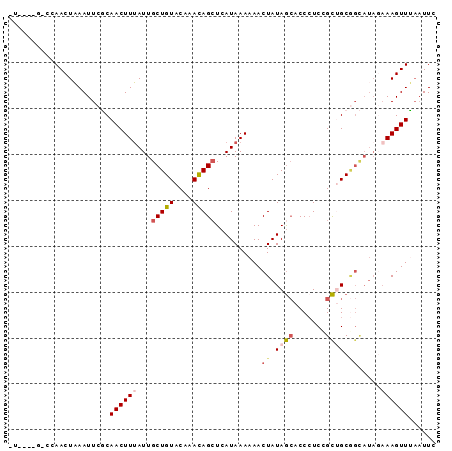
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:32:48 2006