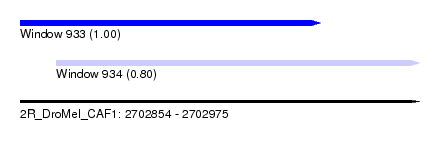

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 2,702,854 – 2,702,975 |
| Length | 121 |
| Max. P | 0.999385 |

| Location | 2,702,854 – 2,702,945 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 94.62 |
| Mean single sequence MFE | -27.00 |
| Consensus MFE | -24.52 |
| Energy contribution | -24.96 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.56 |
| SVM RNA-class probability | 0.999385 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
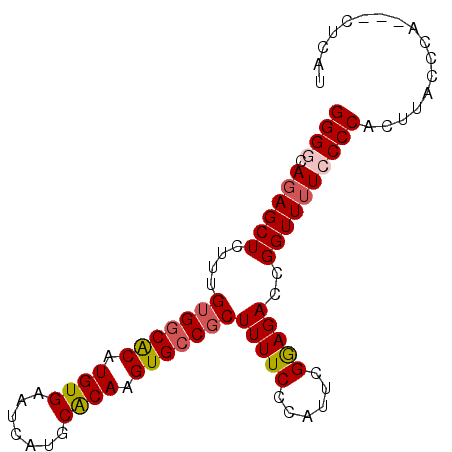
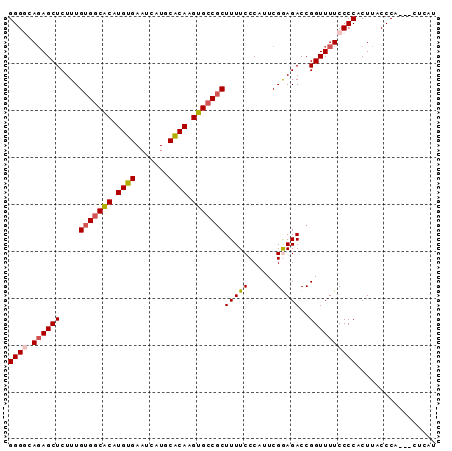
>2R_DroMel_CAF1 2702854 91 + 20766785 GGGGCAUAGCUCUUUGUGGCGCAUGUGAAUCAUGCACAAGUGCCGCUUUUCCCAUUCGGAGACCGGUUUUCCCCACUUACCCACUACUCAU ((((...((((....(((((((.((((.......)))).))))))).(((((.....)))))..))))..))))................. ( -29.30) >DroSec_CAF1 17749 88 + 1 GGGCCAGAGCUCUUUGUGCCACAUGUGAAUCAUGCACAAGUGCCGCUUUUCCCAUUCGGAGACCGGUUUUCCCCACUUACCCA---CUCAU (((..((.....(((((((...(((.....)))))))))).((((..(((((.....))))).))))........))..))).---..... ( -20.30) >DroSim_CAF1 15154 88 + 1 GGGCCAGAGCUCUUUGUGGCACAUGUGAAUCAUGCACAAGUGCCGCUUUUCCCAUUCGGAGACCGGUUUUCCCCACUUACCCA---CUCAU (((..((((((....(((((((.((((.......)))).))))))).(((((.....)))))..)))))).))).........---..... ( -27.30) >DroEre_CAF1 15181 88 + 1 GGGGCAGAGCUCUUUGUGGCACAUGUGAAUCAUGCACAAGUGCCCCUUUUCCCAUUCGAAGACCGGUUUUCCCCACUUACCCA---CUCAU ((((.(((((((((((.(((((.((((.......)))).)))))............))))))...))))))))).........---..... ( -26.22) >DroYak_CAF1 14876 88 + 1 GGGGAAGAGCUCUUUGUGGCACAUGUGAAUCAUGCGCAAGUGCCGCUUUUCCCAUUCGAAGACCGGUUUUCCCCACUUACCCA---CUCAU ((((((((.(.....(((((((.((((.......)))).)))))))(((((......)))))..).)))))))).........---..... ( -31.90) >consensus GGGGCAGAGCUCUUUGUGGCACAUGUGAAUCAUGCACAAGUGCCGCUUUUCCCAUUCGGAGACCGGUUUUCCCCACUUACCCA___CUCAU ((((.((((((....(((((((.((((.......)))).)))))))(((((......)))))..))))))))))................. (-24.52 = -24.96 + 0.44)

| Location | 2,702,865 – 2,702,975 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 96.12 |
| Mean single sequence MFE | -24.44 |
| Consensus MFE | -22.63 |
| Energy contribution | -22.91 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.804680 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
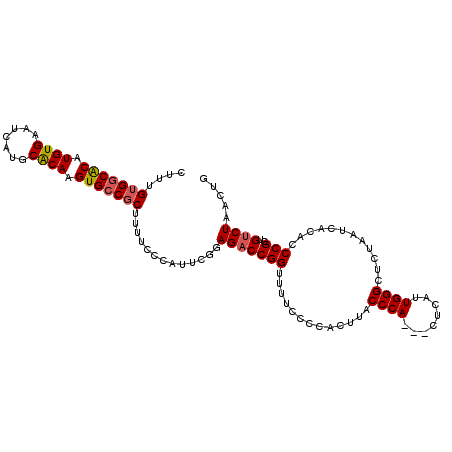
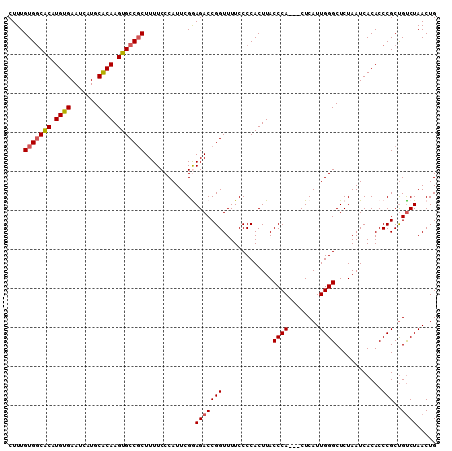
>2R_DroMel_CAF1 2702865 110 + 20766785 CUUUGUGGCGCAUGUGAAUCAUGCACAAGUGCCGCUUUUCCCAUUCGGAGACCGGUUUUCCCCACUUACCCACUACUCAUUGGGCUCUAAUCACACCCGCUGUCUAACUG ....(((((((.((((.......)))).))))))).....((....))(((((((.............((((........))))............)))..))))..... ( -26.51) >DroSec_CAF1 17760 107 + 1 CUUUGUGCCACAUGUGAAUCAUGCACAAGUGCCGCUUUUCCCAUUCGGAGACCGGUUUUCCCCACUUACCCA---CUCAUUGGGCUCUAAUCACACCCGCUGCCUAACUG .(((((((...(((.....)))))))))).((((..(((((.....))))).))))................---....((((((................))))))... ( -20.79) >DroSim_CAF1 15165 107 + 1 CUUUGUGGCACAUGUGAAUCAUGCACAAGUGCCGCUUUUCCCAUUCGGAGACCGGUUUUCCCCACUUACCCA---CUCAUUGGGCUCUAAUCACACCCGCUGUCUAACUG ....(((((((.((((.......)))).))))))).....((....))(((((((.............((((---.....))))............)))..))))..... ( -26.91) >DroEre_CAF1 15192 107 + 1 CUUUGUGGCACAUGUGAAUCAUGCACAAGUGCCCCUUUUCCCAUUCGAAGACCGGUUUUCCCCACUUACCCA---CUCAUUGGGCUCUAAUCACACCCGCCGUCUAACUG ......(((((.((((.......)))).)))))...............((((.(((............((((---.....))))..............)))))))..... ( -22.17) >DroYak_CAF1 14887 107 + 1 CUUUGUGGCACAUGUGAAUCAUGCGCAAGUGCCGCUUUUCCCAUUCGAAGACCGGUUUUCCCCACUUACCCA---CUCAUUGGGCUCUAAUCACACCCGCUGUCUAACUG ....(((((((.((((.......)))).))))))).............(((((((.............((((---.....))))............)))..))))..... ( -25.81) >consensus CUUUGUGGCACAUGUGAAUCAUGCACAAGUGCCGCUUUUCCCAUUCGGAGACCGGUUUUCCCCACUUACCCA___CUCAUUGGGCUCUAAUCACACCCGCUGUCUAACUG ....(((((((.((((.......)))).))))))).............(((((((.............((((........))))............)))..))))..... (-22.63 = -22.91 + 0.28)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:40:01 2006