
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 15,881,900 – 15,881,997 |
| Length | 97 |
| Max. P | 0.938603 |

| Location | 15,881,900 – 15,881,997 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 97.01 |
| Mean single sequence MFE | -8.58 |
| Consensus MFE | -8.48 |
| Energy contribution | -8.48 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.01 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.938603 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
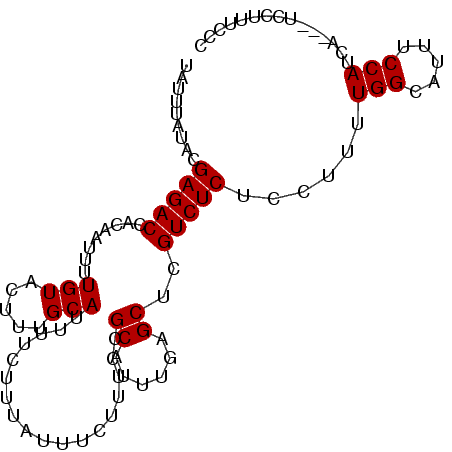
>2R_DroMel_CAF1 15881900 97 + 20766785 UAUUUAUACGAGACCACAAUUUUGUACUUUGCAUUUUCUUUAUUUCUUUCCGCAUUUGAGCUCGUCUCUCCUUUUGGCAUUUCCAUCA---UCCUUUCCC .........(((((........(((.....)))..................((......))..)))))......(((.....)))...---......... ( -8.50) >DroSec_CAF1 19013 97 + 1 UAUUUAUACGAGACCACAAUUUUGUACUUUGCAUUUUCUUUAUUUCUUUCCGCAUUUGAGCUCGUCUCUCCUUUUGGCAUUUCCAUCA---UCCUUUCCC .........(((((........(((.....)))..................((......))..)))))......(((.....)))...---......... ( -8.50) >DroSim_CAF1 16444 97 + 1 UAUUUAUACGAGACCACAAUUUUGUACUUUGCAUUUUCUUUAUUUCUUUCCGCAUUUGAGCUCGUCUCUCCUUUUGGCAUUUCCAUCA---UCCUUUCCC .........(((((........(((.....)))..................((......))..)))))......(((.....)))...---......... ( -8.50) >DroEre_CAF1 16792 97 + 1 UAUUUAUACGAGACCACAAUUUUGUACUUUGCAUUUUCUUUAUUUCUUUCCGCAUUUGAGCUCGUCUCUCCUUUUGGCAUUUCCAUCA---UCCUUUCAG .........(((((........(((.....)))..................((......))..)))))......(((.....)))...---......... ( -8.50) >DroYak_CAF1 19528 97 + 1 UAUUUAUACGAGACCACAAUUUUGUACUUUGCAUUUUCUUUAUUUCUUUCCGCAUUUGAGCUCGUCUCUCCUUUUGGCAUUUCCAUCA---UCCUUUCAA .........(((((........(((.....)))..................((......))..)))))......(((.....)))...---......... ( -8.50) >DroAna_CAF1 11838 100 + 1 UAUUUAUACGAGACCACAAUUUUGUACUUUGCAUUUUCUUUAUUUCUUUCAGCAUUUCAGCUCGUCUCUCCUUUUGGCAUUUCCAUCAUCAUCCUUUCCA .........(((((........(((.....))).................(((......))).)))))......(((.....)))............... ( -9.00) >consensus UAUUUAUACGAGACCACAAUUUUGUACUUUGCAUUUUCUUUAUUUCUUUCCGCAUUUGAGCUCGUCUCUCCUUUUGGCAUUUCCAUCA___UCCUUUCCC .........(((((........(((.....)))..................((......))..)))))......(((.....)))............... ( -8.48 = -8.48 + -0.00)


| Location | 15,881,900 – 15,881,997 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 97.01 |
| Mean single sequence MFE | -12.40 |
| Consensus MFE | -12.12 |
| Energy contribution | -12.12 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.95 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.882731 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
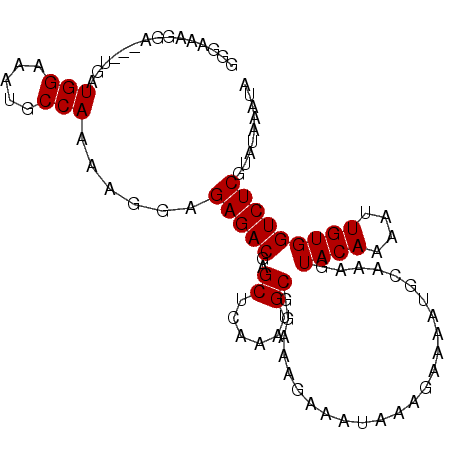
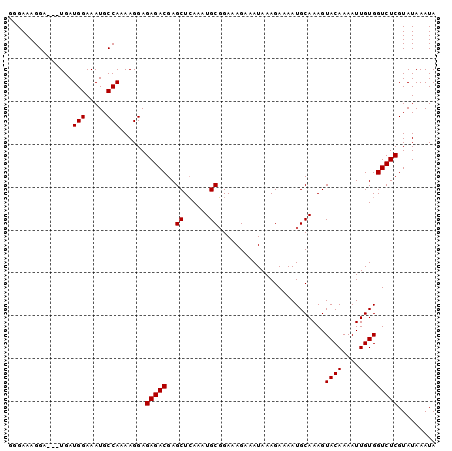
>2R_DroMel_CAF1 15881900 97 - 20766785 GGGAAAGGA---UGAUGGAAAUGCCAAAAGGAGAGACGAGCUCAAAUGCGGAAAGAAAUAAAGAAAAUGCAAAGUACAAAAUUGUGGUCUCGUAUAAAUA .........---...(((.....)))......(((((..((......)).........................((((....)))))))))......... ( -12.20) >DroSec_CAF1 19013 97 - 1 GGGAAAGGA---UGAUGGAAAUGCCAAAAGGAGAGACGAGCUCAAAUGCGGAAAGAAAUAAAGAAAAUGCAAAGUACAAAAUUGUGGUCUCGUAUAAAUA .........---...(((.....)))......(((((..((......)).........................((((....)))))))))......... ( -12.20) >DroSim_CAF1 16444 97 - 1 GGGAAAGGA---UGAUGGAAAUGCCAAAAGGAGAGACGAGCUCAAAUGCGGAAAGAAAUAAAGAAAAUGCAAAGUACAAAAUUGUGGUCUCGUAUAAAUA .........---...(((.....)))......(((((..((......)).........................((((....)))))))))......... ( -12.20) >DroEre_CAF1 16792 97 - 1 CUGAAAGGA---UGAUGGAAAUGCCAAAAGGAGAGACGAGCUCAAAUGCGGAAAGAAAUAAAGAAAAUGCAAAGUACAAAAUUGUGGUCUCGUAUAAAUA ((....)).---...(((.....)))......(((((..((......)).........................((((....)))))))))......... ( -12.60) >DroYak_CAF1 19528 97 - 1 UUGAAAGGA---UGAUGGAAAUGCCAAAAGGAGAGACGAGCUCAAAUGCGGAAAGAAAUAAAGAAAAUGCAAAGUACAAAAUUGUGGUCUCGUAUAAAUA .........---...(((.....)))......(((((..((......)).........................((((....)))))))))......... ( -12.20) >DroAna_CAF1 11838 100 - 1 UGGAAAGGAUGAUGAUGGAAAUGCCAAAAGGAGAGACGAGCUGAAAUGCUGAAAGAAAUAAAGAAAAUGCAAAGUACAAAAUUGUGGUCUCGUAUAAAUA ...............(((.....)))......(((((.(((......)))........................((((....)))))))))......... ( -13.00) >consensus GGGAAAGGA___UGAUGGAAAUGCCAAAAGGAGAGACGAGCUCAAAUGCGGAAAGAAAUAAAGAAAAUGCAAAGUACAAAAUUGUGGUCUCGUAUAAAUA ...............(((.....)))......(((((..((......)).........................((((....)))))))))......... (-12.12 = -12.12 + -0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:30:34 2006