
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 15,652,085 – 15,652,176 |
| Length | 91 |
| Max. P | 0.998426 |

| Location | 15,652,085 – 15,652,176 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 70.44 |
| Mean single sequence MFE | -30.07 |
| Consensus MFE | -10.90 |
| Energy contribution | -12.89 |
| Covariance contribution | 1.99 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.71 |
| Structure conservation index | 0.36 |
| SVM decision value | 3.10 |
| SVM RNA-class probability | 0.998426 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 15652085 91 + 20766785 -AAAAGGGUUGAGCUUGGCUG-C-CUACAAGGGCGAGUGCAAAGCGCGCUAUAAAAC----GUCUCUGAGCGCUUUUUGGCAGCCCAUUUAACCCGGC -....((((((((...(((((-(-(...((((((((((((.....)))))......(----(....))..))))))).)))))))..))))))))... ( -36.60) >DroSec_CAF1 529327 91 + 1 -AAAAGGUUCGAGCUUGGCUG-C-CAGCAAAGGCGAGUGCAGAGCGCGCUAUAAAAC----GUCUCUGAGUGCUUUUUGGCAGCCCAUUUAACCCGGC -....((((.((....(((((-(-((..(((((((....(((((.(((........)----)))))))..)))))))))))))))..)).)))).... ( -34.90) >DroSim_CAF1 426052 91 + 1 -AAAAGGGUUGAGCUUGGCUG-C-CAGCAAAGGCGAGUGCAGAGCGCGCUAUAAAAC----AUCUCUGAGUGCUUUUUGGCAGCCCAUUUAACCCGGC -....((((((((...(((((-(-((..(((((((....(((((.............----..)))))..)))))))))))))))..))))))))... ( -37.46) >DroWil_CAF1 824904 85 + 1 ---AAGAGUUUAACUU-ACUCACUAUACAAGUAUGUAUGUUAAAUGC------AUAUGUCUGGCUCU---GCCUUUUUGGCAGCCCAUUUAACCCGAC ---..((((.......-)))).........((((((((.....))))------))))...(((..((---(((.....))))).)))........... ( -21.00) >DroYak_CAF1 448414 70 + 1 AAAAAGGGUUGAGC------------------------ACAAAGCGCGCUAUAAAAC----GGCUCUGAGUGCUUUUUGGCAGCCCAUUUAACCCGGC .....((((((..(------------------------(.((((((((((.......----))).....))))))).)).))))))............ ( -20.40) >consensus _AAAAGGGUUGAGCUUGGCUG_C_CAACAAAGGCGAGUGCAAAGCGCGCUAUAAAAC____GUCUCUGAGUGCUUUUUGGCAGCCCAUUUAACCCGGC .....((((((.(((((.((..........)).))))).(((((.(((((..................))))).))))).))))))............ (-10.90 = -12.89 + 1.99)


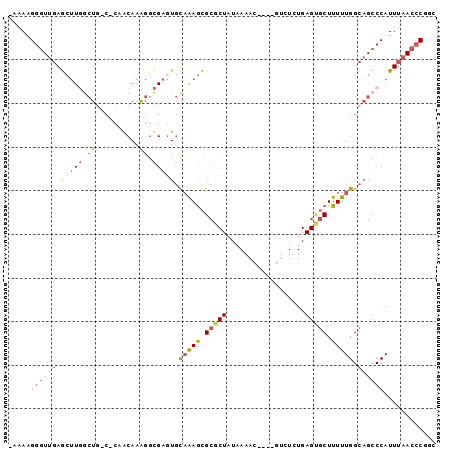
| Location | 15,652,085 – 15,652,176 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 70.44 |
| Mean single sequence MFE | -28.47 |
| Consensus MFE | -9.11 |
| Energy contribution | -9.43 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.32 |
| SVM decision value | 2.19 |
| SVM RNA-class probability | 0.989928 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
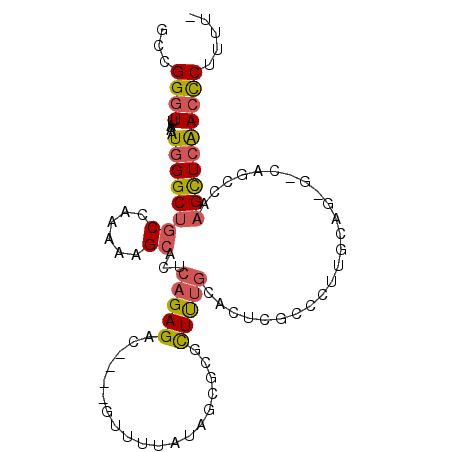
>2R_DroMel_CAF1 15652085 91 - 20766785 GCCGGGUUAAAUGGGCUGCCAAAAAGCGCUCAGAGAC----GUUUUAUAGCGCGCUUUGCACUCGCCCUUGUAG-G-CAGCCAAGCUCAACCCUUUU- ...(((((.....(((((((..(((((((.(......----........).)))))))((....)).......)-)-)))))......)))))....- ( -31.74) >DroSec_CAF1 529327 91 - 1 GCCGGGUUAAAUGGGCUGCCAAAAAGCACUCAGAGAC----GUUUUAUAGCGCGCUCUGCACUCGCCUUUGCUG-G-CAGCCAAGCUCGAACCUUUU- ..((((((.....((((((((.(((((...(((((.(----(((....))))..))))).....).))))..))-)-))))).))))))........- ( -32.50) >DroSim_CAF1 426052 91 - 1 GCCGGGUUAAAUGGGCUGCCAAAAAGCACUCAGAGAU----GUUUUAUAGCGCGCUCUGCACUCGCCUUUGCUG-G-CAGCCAAGCUCAACCCUUUU- ...(((((.....((((((((.(((((...(((((.(----(((....))))..))))).....).))))..))-)-)))))......)))))....- ( -32.30) >DroWil_CAF1 824904 85 - 1 GUCGGGUUAAAUGGGCUGCCAAAAAGGC---AGAGCCAGACAUAU------GCAUUUAACAUACAUACUUGUAUAGUGAGU-AAGUUAAACUCUU--- ...(((((...(((.(((((.....)))---))..))).......------.....((((.(((.((((.....)))).))-).)))))))))..--- ( -22.80) >DroYak_CAF1 448414 70 - 1 GCCGGGUUAAAUGGGCUGCCAAAAAGCACUCAGAGCC----GUUUUAUAGCGCGCUUUGU------------------------GCUCAACCCUUUUU ...((((....((((((((......)))..(((((((----(((....)))).)))))).------------------------)))))))))..... ( -23.00) >consensus GCCGGGUUAAAUGGGCUGCCAAAAAGCACUCAGAGAC____GUUUUAUAGCGCGCUUUGCACUCGCCCUUGCAG_G_CAGCCAAGCUCAACCCUUUU_ ...((((....((((((((......))...(((((...................)))))........................))))))))))..... ( -9.11 = -9.43 + 0.32)

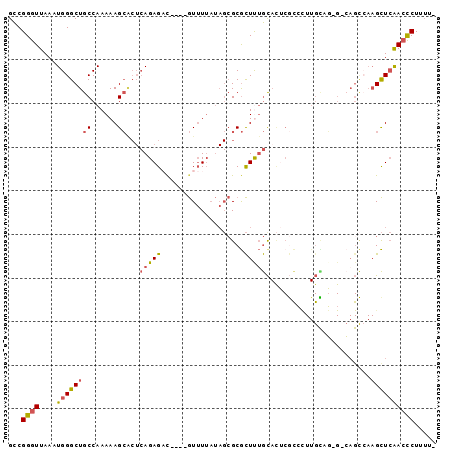
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:28:34 2006