| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 15,405,937 – 15,406,056 |
| Length | 119 |
| Max. P | 0.993429 |

| Location | 15,405,937 – 15,406,056 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.36 |
| Mean single sequence MFE | -24.85 |
| Consensus MFE | -17.43 |
| Energy contribution | -17.79 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.11 |
| Mean z-score | -3.36 |
| Structure conservation index | 0.70 |
| SVM decision value | 2.40 |
| SVM RNA-class probability | 0.993429 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 15405937 119 + 20766785 AAUCGAAUUACAAGCCAUCCGAAGUGAGUGAAAUGAAAAUUAAUUGAAGACAAUGGAAUGCAAAAUGCAAUUUCAAUUAAUUUUCAUUCUUUGCACACUGCGACCGCCGCACGUACAUA- ........(((...........((((.(..(((((((((((((((((((.(....)..(((.....))).)))))))))))))))))..))..).))))(((.....)))..)))....- ( -28.20) >DroPse_CAF1 240757 95 + 1 AAUCGAAUUACAGCCAAUUCAGAGUGAUUGAAAUGAAAAUUAAUUGAAGACAAUGGAAUGCAGCAUGCAUUUUCAAUUAAUUUUCAUUCUCUGCA------------------------- ....(((((......)))))((((.......((((((((((((((((........((((((.....))))))))))))))))))))))))))...------------------------- ( -23.41) >DroWil_CAF1 419367 97 + 1 AAACGUAUUACAAGCCAUUCGCAGUGAUUGAAAUGAAAAUUAAUUGAAGACAAUGGCAUGCAAAAUGGAUUUUCAAUUAAUUUUCAUUCUUUU---------------------AC-UU- .............((.....))(((((..(((.(((((((((((((((((..((.(....)...))...))))))))))))))))))))..))---------------------))-).- ( -22.40) >DroMoj_CAF1 307090 110 + 1 ACUCAAUUUGCUUGCUAUUGGAAAUGAUUGAAAUGAAAAUUAAUUGAAGACAAUGGCAUGCAAAAUGCAUUUUCAAUUAAUUUUCAUUCUUUAU-------GACA---ACAUAUUCAUAU ..((((((...((.(....).))..))))))(((((((((((((((((((.....((((.....)))).)))))))))))))))))))...(((-------((..---......))))). ( -26.70) >DroAna_CAF1 189092 107 + 1 AAUCGAAUUACAAGCUAUAUGAAGCGAACGAAAUGAAAAUUAAUUGAAGACAAUGGAAUGCAAAAUGCAUUUUCAAUUAAUUUUCAUUCUUUGCAGAAGA---------AACGUGG---- .......................(((((.(((.((((((((((((((........((((((.....))))))))))))))))))))))))))))......---------.......---- ( -25.00) >DroPer_CAF1 200777 95 + 1 AAUCGAAUUACAGCCAAUUCAGAGUGAUUGAAAUGAAAAUUAAUUGAAGACAAUGGAAUGCAGCAUGCAUUUUCAAUUAAUUUUCAUUCUCUGCA------------------------- ....(((((......)))))((((.......((((((((((((((((........((((((.....))))))))))))))))))))))))))...------------------------- ( -23.41) >consensus AAUCGAAUUACAAGCAAUUCGAAGUGAUUGAAAUGAAAAUUAAUUGAAGACAAUGGAAUGCAAAAUGCAUUUUCAAUUAAUUUUCAUUCUUUGCA____________________C____ ...................(((((.......(((((((((((((((((((........(((.....)))))))))))))))))))))))))))........................... (-17.43 = -17.79 + 0.36)


| Location | 15,405,937 – 15,406,056 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.36 |
| Mean single sequence MFE | -24.65 |
| Consensus MFE | -17.78 |
| Energy contribution | -18.45 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.11 |
| Structure conservation index | 0.72 |
| SVM decision value | 2.33 |
| SVM RNA-class probability | 0.992470 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 15405937 119 - 20766785 -UAUGUACGUGCGGCGGUCGCAGUGUGCAAAGAAUGAAAAUUAAUUGAAAUUGCAUUUUGCAUUCCAUUGUCUUCAAUUAAUUUUCAUUUCACUCACUUCGGAUGGCUUGUAAUUCGAUU -..((((((((((.....)))).))))))..((((((((((((((((((..(((.....)))..........))))))))))))))))))........((((((........)))))).. ( -33.00) >DroPse_CAF1 240757 95 - 1 -------------------------UGCAGAGAAUGAAAAUUAAUUGAAAAUGCAUGCUGCAUUCCAUUGUCUUCAAUUAAUUUUCAUUUCAAUCACUCUGAAUUGGCUGUAAUUCGAUU -------------------------(((((.(((((((((((((((((((((((.....)))))........))))))))))))))))))(((((.....).)))).)))))........ ( -25.00) >DroWil_CAF1 419367 97 - 1 -AA-GU---------------------AAAAGAAUGAAAAUUAAUUGAAAAUCCAUUUUGCAUGCCAUUGUCUUCAAUUAAUUUUCAUUUCAAUCACUGCGAAUGGCUUGUAAUACGUUU -..-((---------------------(...((((((((((((((((((..........(((......))).)))))))))))))))))).......(((((.....))))).))).... ( -18.20) >DroMoj_CAF1 307090 110 - 1 AUAUGAAUAUGU---UGUC-------AUAAAGAAUGAAAAUUAAUUGAAAAUGCAUUUUGCAUGCCAUUGUCUUCAAUUAAUUUUCAUUUCAAUCAUUUCCAAUAGCAAGCAAAUUGAGU ..((((.((((.---...)-------)))..(((((((((((((((((((((((((.....))).))))...))))))))))))))))))...))))....................... ( -21.60) >DroAna_CAF1 189092 107 - 1 ----CCACGUU---------UCUUCUGCAAAGAAUGAAAAUUAAUUGAAAAUGCAUUUUGCAUUCCAUUGUCUUCAAUUAAUUUUCAUUUCGUUCGCUUCAUAUAGCUUGUAAUUCGAUU ----.......---------.....(((((.(((((((((((((((((((((((.....)))))........)))))))))))))))))).....(((......))))))))........ ( -25.10) >DroPer_CAF1 200777 95 - 1 -------------------------UGCAGAGAAUGAAAAUUAAUUGAAAAUGCAUGCUGCAUUCCAUUGUCUUCAAUUAAUUUUCAUUUCAAUCACUCUGAAUUGGCUGUAAUUCGAUU -------------------------(((((.(((((((((((((((((((((((.....)))))........))))))))))))))))))(((((.....).)))).)))))........ ( -25.00) >consensus ____G____________________UGCAAAGAAUGAAAAUUAAUUGAAAAUGCAUUUUGCAUUCCAUUGUCUUCAAUUAAUUUUCAUUUCAAUCACUUCGAAUAGCUUGUAAUUCGAUU ...............................(((((((((((((((((((((((.....)))))........)))))))))))))))))).............................. (-17.78 = -18.45 + 0.67)
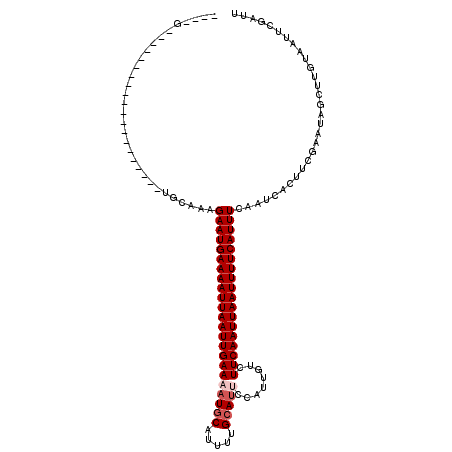



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:27:15 2006