
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 2,656,605 – 2,656,712 |
| Length | 107 |
| Max. P | 0.517516 |

| Location | 2,656,605 – 2,656,712 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 78.83 |
| Mean single sequence MFE | -36.60 |
| Consensus MFE | -22.61 |
| Energy contribution | -22.62 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.62 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.517516 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
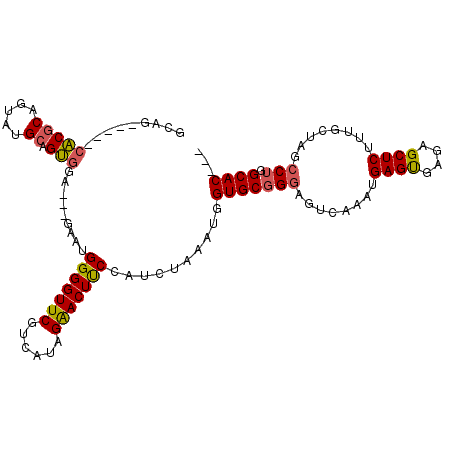
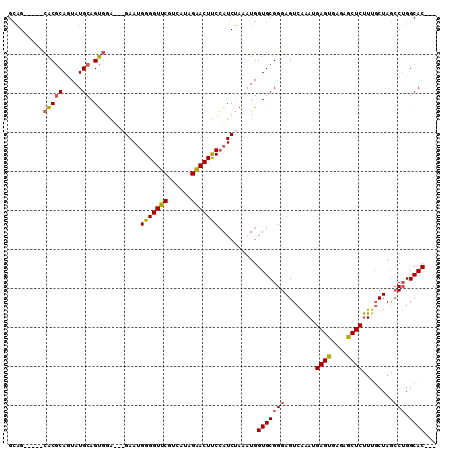
>2R_DroMel_CAF1 2656605 107 - 20766785 GCAGGAUGCCACGCAGUAUGCAGUGGG---GAAUGGGGUUCGUCUUAGGACUUCCAUCUAAAUGGUGCGGGAGUCAAAUGAGUGAGAGCUCUUUGCUAGCCUGGCACUCG .((((...((((((.....)).))))(---(...(((((((..((((.((((((((((.....)))..)))))))...))))...)))))))...))..))))....... ( -37.50) >DroSec_CAF1 89842 102 - 1 GGAG-----CACGCAGUAUGCAGUGGAUAAGAAUGGGGUUCGUCAUAGAACUUCCAUCUAAAUGGUGCGGGAGUCAAAUGAGCGAAAGCUCUUGGCUAAACUGGCAC--- ....-----(((((.....)).)))....(((.((((((((......)))).))))))).....((((.((((((((..((((....))))))))))...)).))))--- ( -34.20) >DroSim_CAF1 87762 102 - 1 GCAG-----CACGCAGUAUGCAGUGGAUAAGAAUGGGGUUCGUCAUAGAACUUCCAUCUAAGUGGUGCGGGAGUCAAAUGAGUGAGAGCUCUUUGCUAGCCUGGCAC--- (((.-----(((((.....))........(((.((((((((......)))).)))))))..))).)))((.((.((((.((((....))))))))))..))......--- ( -33.00) >DroEre_CAF1 88147 97 - 1 GCUG-----UGCCCACUAUGCAGCUCG---GAA-GGGGUUCCUUAUGGGACUCCGAUCGCGAUGGUGCGGGAGUCGGAUGAGUGAGAGCUCCUUUCUAGCC-GGCAC--- ((((-----(((.......)))(((.(---(((-((((((((((((..((((((...((((....))))))))))..)))))...))))))))))).))))-)))..--- ( -41.70) >consensus GCAG_____CACGCAGUAUGCAGUGGA___GAAUGGGGUUCGUCAUAGAACUUCCAUCUAAAUGGUGCGGGAGUCAAAUGAGUGAGAGCUCUUUGCUAGCCUGGCAC___ .........(((((.....)).))).........(((((((......)))))))..........(((((((........((((....))))........))).))))... (-22.61 = -22.62 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:39:20 2006