
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 15,022,905 – 15,023,001 |
| Length | 96 |
| Max. P | 0.820724 |

| Location | 15,022,905 – 15,023,001 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 90.60 |
| Mean single sequence MFE | -29.54 |
| Consensus MFE | -23.98 |
| Energy contribution | -23.62 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.801137 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
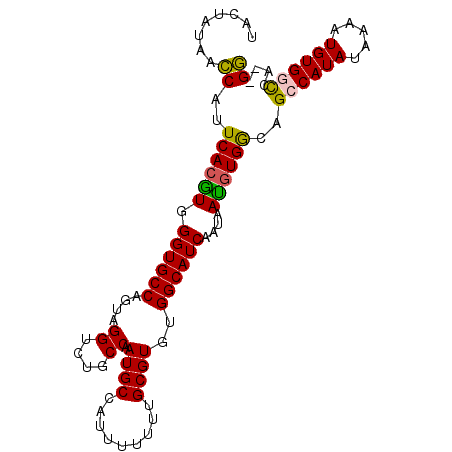
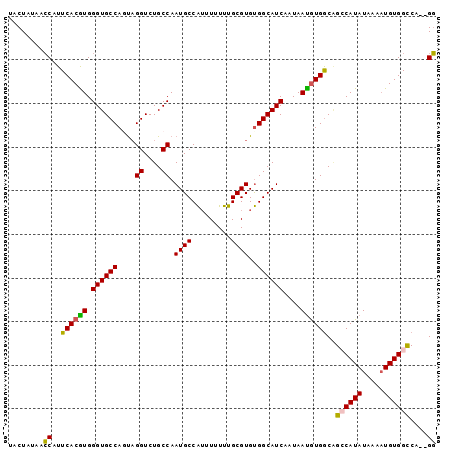
>2R_DroMel_CAF1 15022905 96 + 20766785 UACUAUAACCAUUCACGUGGGUGCCAGUUGGUCUGCCAAUGCCAGUUUUUUGCGUGUGGCAUCAAUAAUAUGACAGCCAUAUAAAAUGUGGCCA--GG ........((..(((..(.((((((((((((....)))))(((((....))).)).)))))))....)..)))..(((((((...)))))))..--)) ( -26.10) >DroSec_CAF1 10780 96 + 1 UACUAUAACCAUUCACGUGGGUGCCAGUAGGUCUGCCAAUGCCGUUUUUUUGCGUGCGGCAUCAAUAAUGUGGCAGCCAUAUAAAAUGUGGCCA--GG ........((..((((((.((((((....((....))...(((((......))).))))))))....))))))..(((((((...)))))))..--)) ( -31.00) >DroSim_CAF1 8767 96 + 1 UACUAUAACCAUUCACGUGGGUGCCAGUAGGUCUGCCAAUGCCAUUUUUUUGCGUGUGGCAUCAAUAAUGUGGCAGCCAUAUAAAAUGUGGCCA--GG ........((..((((((.(((((((...((....)).((((.........)))).)))))))....))))))..(((((((...)))))))..--)) ( -27.80) >DroEre_CAF1 10137 95 + 1 UACUAUAGCCAUUCACGUGGGUGCCACCAGGUCUGCCAAUGCCAUUU-UUUGCGUGUGGCAUCAAUAACGUGGCCGGCAUAUAAAUUGUGGCCA--GG ...(((((((..((((((.((((((((..((....)).((((.....-...))))))))))))....))))))..))).))))...........--.. ( -31.50) >DroYak_CAF1 10119 98 + 1 UACUAUAAGCAUUCACAUGGGUGCCACUAGGUCUGCCAAUGCCAUUUUUUUGCGUGUGGCAUCAAUAAUGUGGCGGGCAUAUAAAAUGUGGUCCUGGC ..(((...((((((....))))))...)))....((((..(((((((((.(((.(((.((((.....)))).))).)))...)))).)))))..)))) ( -31.30) >consensus UACUAUAACCAUUCACGUGGGUGCCAGUAGGUCUGCCAAUGCCAUUUUUUUGCGUGUGGCAUCAAUAAUGUGGCAGCCAUAUAAAAUGUGGCCA__GG ........((..((((((.((((((....((....)).((((.........))))..))))))....))))))..((((((.....))))))....)) (-23.98 = -23.62 + -0.36)

| Location | 15,022,905 – 15,023,001 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 90.60 |
| Mean single sequence MFE | -25.90 |
| Consensus MFE | -22.28 |
| Energy contribution | -22.68 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820724 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
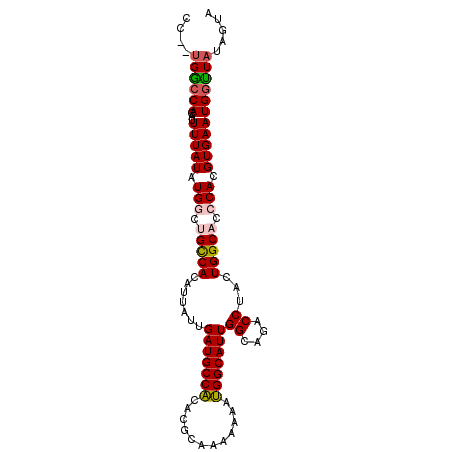
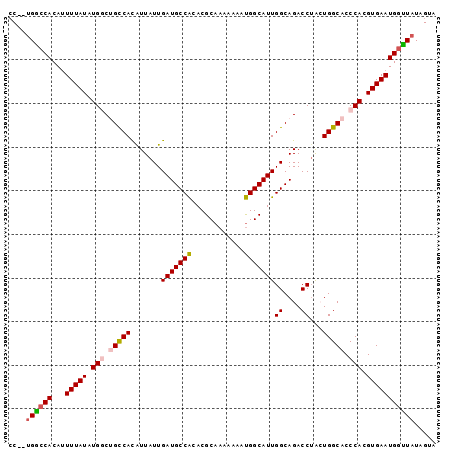
>2R_DroMel_CAF1 15022905 96 - 20766785 CC--UGGCCACAUUUUAUAUGGCUGUCAUAUUAUUGAUGCCACACGCAAAAAACUGGCAUUGGCAGACCAACUGGCACCCACGUGAAUGGUUAUAGUA ..--((((((...((((..(((.(((((.......(((((((............)))))))((....))...))))).)))..))))))))))..... ( -24.30) >DroSec_CAF1 10780 96 - 1 CC--UGGCCACAUUUUAUAUGGCUGCCACAUUAUUGAUGCCGCACGCAAAAAAACGGCAUUGGCAGACCUACUGGCACCCACGUGAAUGGUUAUAGUA ..--((((((...((((..(((.(((((.......(((((((............)))))))((....))...))))).)))..))))))))))..... ( -27.60) >DroSim_CAF1 8767 96 - 1 CC--UGGCCACAUUUUAUAUGGCUGCCACAUUAUUGAUGCCACACGCAAAAAAAUGGCAUUGGCAGACCUACUGGCACCCACGUGAAUGGUUAUAGUA ..--((((((...((((..(((.(((((.......(((((((............)))))))((....))...))))).)))..))))))))))..... ( -26.30) >DroEre_CAF1 10137 95 - 1 CC--UGGCCACAAUUUAUAUGCCGGCCACGUUAUUGAUGCCACACGCAAA-AAAUGGCAUUGGCAGACCUGGUGGCACCCACGUGAAUGGCUAUAGUA ..--((((((((.......(((((.(((.(((.(..((((((........-...))))))..)..))).))))))))......))..))))))..... ( -28.82) >DroYak_CAF1 10119 98 - 1 GCCAGGACCACAUUUUAUAUGCCCGCCACAUUAUUGAUGCCACACGCAAAAAAAUGGCAUUGGCAGACCUAGUGGCACCCAUGUGAAUGCUUAUAGUA .............((((((((...(((((......(((((((............)))))))((....))..)))))...))))))))........... ( -22.50) >consensus CC__UGGCCACAUUUUAUAUGGCUGCCACAUUAUUGAUGCCACACGCAAAAAAAUGGCAUUGGCAGACCUACUGGCACCCACGUGAAUGGUUAUAGUA ....((((((...(((((.(((.(((((.......(((((((............)))))))((....))...))))).))).)))))))))))..... (-22.28 = -22.68 + 0.40)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:24:38 2006