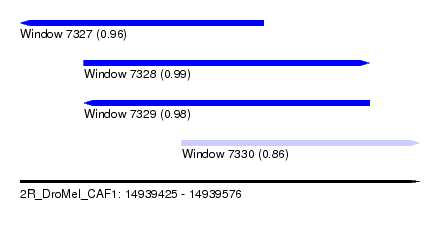

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 14,939,425 – 14,939,576 |
| Length | 151 |
| Max. P | 0.993441 |

| Location | 14,939,425 – 14,939,517 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 68.12 |
| Mean single sequence MFE | -25.36 |
| Consensus MFE | -9.58 |
| Energy contribution | -8.88 |
| Covariance contribution | -0.70 |
| Combinations/Pair | 1.58 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.38 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961722 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 14939425 92 - 20766785 ACACACAC---------GCACGGACACGAAUGUGUGUAUAUGGCGCGGCGCUUGUGUGUG--AAAGUGUAUUCGUA-UAAAUAAUAUGACAUAAUCGACGUGUG---------------- .(((((((---------(((((........))))))).((((((((..(((......)))--...))))..(((((-(.....))))))))))......)))))---------------- ( -26.50) >DroSim_CAF1 25450 92 - 1 ACACACAC---------GCACGGACACGAAUGUGUGUAUAUGGCGCGGCGCUUGUGUGUG--AAAGUGUAUUCGUA-UAAAUAAUAUGACAUAAUCGACGUGUG---------------- .(((((((---------(((((........))))))).((((((((..(((......)))--...))))..(((((-(.....))))))))))......)))))---------------- ( -26.50) >DroWil_CAF1 36269 119 - 1 ACAUAUAUAAAAUAGAUGUACAUACAUACGUGUAUGUGUGUGUUGUG-UGUAUGUGUAUUAAAAAGUGUGUAUAAAUUAAAUAAUAUGACAUAAUCGACGUGUGUGCAUCUGUGUGAGUG ...........((((((((((((((...((..(((((.((((((((.-..((((..((((....))))..))))......)))))))))))))..))..))))))))))))))....... ( -32.70) >DroYak_CAF1 25637 92 - 1 ACACACAC---------GCACGGACACGAAUGUGUGUAUAUGGCGCGACGCUUGUGUGCG--AAAGUGUAUUCGUA-UAAAUAAUAUGACAUAAUCGACGUGUG---------------- ..((((((---------(..((....))..))))))).....(((((.((.((((((((.--...)).....((((-(.....))))))))))).)).))))).---------------- ( -27.30) >DroAna_CAF1 24061 70 - 1 ACAUACAC-----------------ACGAA-AUGUGUACAUGUUGCGACCC---------------UGUAUUUGUA-UAAAUAAUAUGACAUAAUCGACGUGUG---------------- .....(((-----------------(((..-.(((((.((((((((((...---------------.....)))))-......)))))))))).....))))))---------------- ( -13.80) >consensus ACACACAC_________GCACGGACACGAAUGUGUGUAUAUGGCGCGACGCUUGUGUGUG__AAAGUGUAUUCGUA_UAAAUAAUAUGACAUAAUCGACGUGUG________________ .(((((.................(((((....))))).....(((..((((((..........))))))...)))........................)))))................ ( -9.58 = -8.88 + -0.70)



| Location | 14,939,449 – 14,939,557 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.17 |
| Mean single sequence MFE | -32.59 |
| Consensus MFE | -22.01 |
| Energy contribution | -23.82 |
| Covariance contribution | 1.81 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.68 |
| SVM decision value | 2.40 |
| SVM RNA-class probability | 0.993441 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 14939449 108 + 20766785 UUA-UACGAAUACACUUU--CACACACAAGCGCCGCGCCAUAUACACACAUUCGUGUCCGUGC---------GUGUGUGUGUAUGAACGGCGUCGCAUUCUCGUCACCGGCAAUGCCGUG ...-...(((......))--)...(((..((((((...((((((((((((..(((......))---------)))))))))))))..)))))).(((((.(((....))).))))).))) ( -37.70) >DroSim_CAF1 25474 108 + 1 UUA-UACGAAUACACUUU--CACACACAAGCGCCGCGCCAUAUACACACAUUCGUGUCCGUGC---------GUGUGUGUGUAUGAACGGCGUCGCAUUCUCGUCACCGGCAAUGCCGUG ...-...(((......))--)...(((..((((((...((((((((((((..(((......))---------)))))))))))))..)))))).(((((.(((....))).))))).))) ( -37.70) >DroWil_CAF1 36309 114 + 1 UUAAUUUAUACACACUUUUUAAUACACAUACA-CACAACACACACAUACACGUAUGUAUGUACAUCUAUUUUAUAUAUGUAAAUUAACGGCGUCUCGUU-----AACCGGCAACGGCGUG ..........(((.........((((.(((..-..........(((((((....)))))))............))).))))..(((((((....)))))-----))(((....))).))) ( -19.35) >DroYak_CAF1 25661 108 + 1 UUA-UACGAAUACACUUU--CGCACACAAGCGUCGCGCCAUAUACACACAUUCGUGUCCGUGC---------GUGUGUGUGUAUCAACGGCGUCGCAUUCUCGUCACCGGCAAUGCCGUG ...-..((((......))--))..(((..(((..(((((..((((((((((.(((......))---------).))))))))))....))))))))............(((...)))))) ( -35.60) >consensus UUA_UACGAAUACACUUU__CACACACAAGCGCCGCGCCAUAUACACACAUUCGUGUCCGUGC_________GUGUGUGUGUAUGAACGGCGUCGCAUUCUCGUCACCGGCAAUGCCGUG ........................(((..((((((...(((((((((((((.....................)))))))))))))..)))))).(((((.(((....))).))))).))) (-22.01 = -23.82 + 1.81)
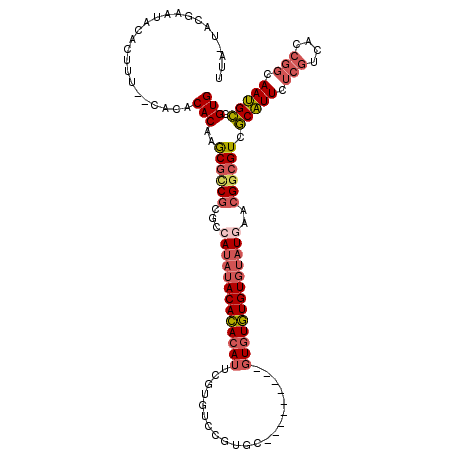



| Location | 14,939,449 – 14,939,557 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.17 |
| Mean single sequence MFE | -37.80 |
| Consensus MFE | -21.85 |
| Energy contribution | -22.60 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.58 |
| SVM decision value | 1.91 |
| SVM RNA-class probability | 0.982110 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 14939449 108 - 20766785 CACGGCAUUGCCGGUGACGAGAAUGCGACGCCGUUCAUACACACACAC---------GCACGGACACGAAUGUGUGUAUAUGGCGCGGCGCUUGUGUGUG--AAAGUGUAUUCGUA-UAA .((((((((..((....))..))))).((((..(((((((((((((((---------(..((....))..)))))))....((((...)))).)))))))--)).))))...))).-... ( -39.80) >DroSim_CAF1 25474 108 - 1 CACGGCAUUGCCGGUGACGAGAAUGCGACGCCGUUCAUACACACACAC---------GCACGGACACGAAUGUGUGUAUAUGGCGCGGCGCUUGUGUGUG--AAAGUGUAUUCGUA-UAA .((((((((..((....))..))))).((((..(((((((((((((((---------(..((....))..)))))))....((((...)))).)))))))--)).))))...))).-... ( -39.80) >DroWil_CAF1 36309 114 - 1 CACGCCGUUGCCGGUU-----AACGAGACGCCGUUAAUUUACAUAUAUAAAAUAGAUGUACAUACAUACGUGUAUGUGUGUGUUGUG-UGUAUGUGUAUUAAAAAGUGUGUAUAAAUUAA (((((((((..((...-----..)).))))...(((((.((((((((((....(.(..((((((((....))))))))..).)..))-)))))))).)))))...))))).......... ( -32.00) >DroYak_CAF1 25661 108 - 1 CACGGCAUUGCCGGUGACGAGAAUGCGACGCCGUUGAUACACACACAC---------GCACGGACACGAAUGUGUGUAUAUGGCGCGACGCUUGUGUGCG--AAAGUGUAUUCGUA-UAA ..((((...))))...(((((....((.((((((..(((((((((..(---------(........))..)))))))))))))))))...)))))(((((--((......))))))-).. ( -39.60) >consensus CACGGCAUUGCCGGUGACGAGAAUGCGACGCCGUUCAUACACACACAC_________GCACGGACACGAAUGUGUGUAUAUGGCGCGGCGCUUGUGUGUG__AAAGUGUAUUCGUA_UAA (((.(((((..((....))..)))))..(((((((((((.(((((((.......................))))))).))))).))))))...............)))............ (-21.85 = -22.60 + 0.75)

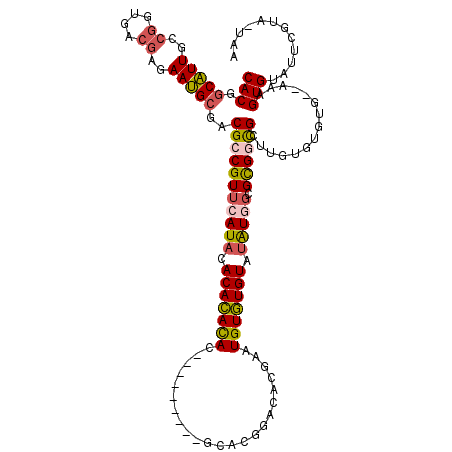
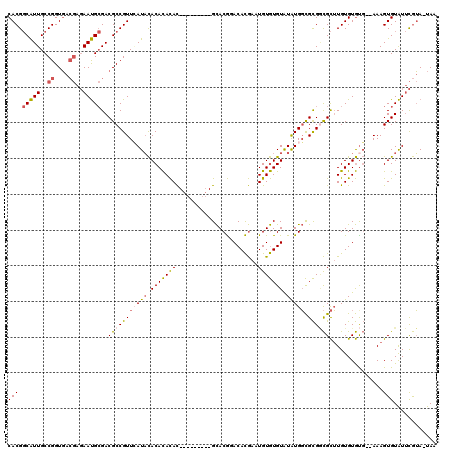
| Location | 14,939,486 – 14,939,576 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 67.23 |
| Mean single sequence MFE | -27.56 |
| Consensus MFE | -13.92 |
| Energy contribution | -15.52 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.855456 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 14939486 90 + 20766785 UAUACACACAUUCGUGUCCGUGC---------GUGUGUGUGUAUGAACGGCGUCGCAUUCUCGUCACCGGCAAUGCCGUGAAAAAAUCGAUGGU---CAAUU--------- (((((((((((.(((......))---------).)))))))))))...(((((((.(((....((((.(((...)))))))...))))))).))---)....--------- ( -30.80) >DroSim_CAF1 25511 90 + 1 UAUACACACAUUCGUGUCCGUGC---------GUGUGUGUGUAUGAACGGCGUCGCAUUCUCGUCACCGGCAAUGCCGUGAAAAAAUCGAUGGU---CAAUU--------- (((((((((((.(((......))---------).)))))))))))...(((((((.(((....((((.(((...)))))))...))))))).))---)....--------- ( -30.80) >DroWil_CAF1 36348 105 + 1 CACACAUACACGUAUGUAUGUACAUCUAUUUUAUAUAUGUAAAUUAACGGCGUCUCGUU-----AACCGGCAACGGCGUGAAA-AAUCGAUAUGGCUGCGCUUUUACAGUU ...(((((((....)))))))..(((.(((((.(((.......(((((((....)))))-----))(((....))).))).))-))).)))..(((((........))))) ( -21.70) >DroYak_CAF1 25698 90 + 1 UAUACACACAUUCGUGUCCGUGC---------GUGUGUGUGUAUCAACGGCGUCGCAUUCUCGUCACCGGCAAUGCCGUGAAAAAAUCGAUGGG---CAAUA--------- .((((((((((.(((......))---------).))))))))))......(((((.(((....((((.(((...)))))))...))))))))..---.....--------- ( -28.60) >DroAna_CAF1 24109 84 + 1 UGUACACAU-UUCGU-----------------GUGUAUGUGUAUAGACGGCGUC------CCGUCACCGGCAAUGCCGUGAAAAAAUCGAUGCG--UCCUCUUUCACAGA- .((((((((-...))-----------------))))))(((...(((.(((((.------.((((((.(((...)))))))......))..)))--)).)))..)))...- ( -25.90) >consensus UAUACACACAUUCGUGUCCGUGC_________GUGUGUGUGUAUGAACGGCGUCGCAUUCUCGUCACCGGCAAUGCCGUGAAAAAAUCGAUGGG___CAAUU_________ (((((((((((.......................))))))))))).(((((((.((.............)).)))))))................................ (-13.92 = -15.52 + 1.60)

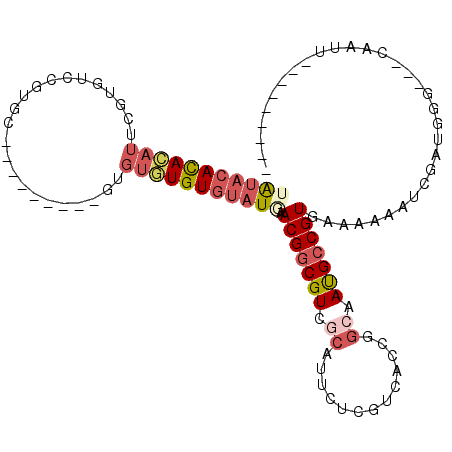
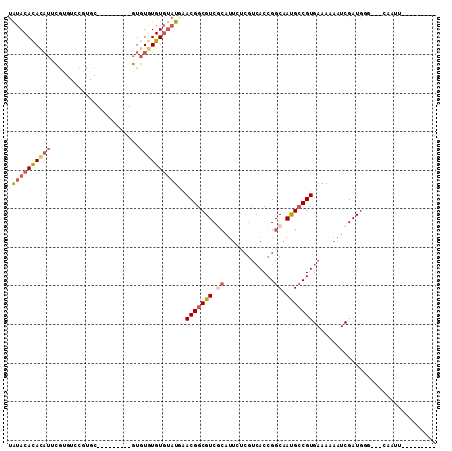
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:23:25 2006