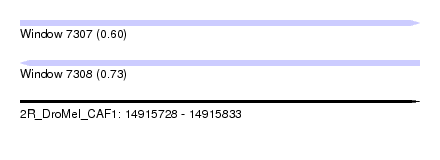

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 14,915,728 – 14,915,833 |
| Length | 105 |
| Max. P | 0.725592 |

| Location | 14,915,728 – 14,915,833 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 79.71 |
| Mean single sequence MFE | -33.48 |
| Consensus MFE | -21.47 |
| Energy contribution | -23.13 |
| Covariance contribution | 1.67 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.597125 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 14915728 105 + 20766785 ---------GAUGUCACCACAUUGAGCAGCUUGCGGUUGUCCAAGCUCAAUGUUGUGAGAUCUAUGAAGGACACAACCUUCUCGCGCUGCUCGGCCACCCGUUGCCAAAUGGAU ---------(((.((((.(((((((((.(....)((....))..))))))))).)))).)))...(((((......)))))..(.(((....))))..(((((....))))).. ( -36.60) >DroSec_CAF1 2165 105 + 1 ---------GAUGUCACCACAUUGAGCAGCUUGCGAUUGUCCAUGCUCAAUGUGGUGAGAUCUAUGAAGGACACAACCUUAUCGCGCUGCUCGGCCACCCGUUGCCAUAUGGAU ---------(((.(((((((((((((((....((....))...))))))))))))))).)))....((((......))))...(.(((....))))..((((......)))).. ( -38.70) >DroEre_CAF1 2322 105 + 1 ---------GAUGUCACCACAUUGAGCAGCUUGCGGUUGUCCAAGCUCAACGUCGUGAGAUCUAUGAAGGACACGACCUUCUCGCGCUGCUCGGCCACCGUUUGCCAAAUGGAU ---------.......(((....(((((((..(((((((........))))(((....)))....(((((......))))).))))))))))(((........)))...))).. ( -34.90) >DroYak_CAF1 2131 105 + 1 ---------GAUGUCACCACAUUCAGCAGCUUGCGGUUGUCCAUGCUCAGUGUCGUGAGAUCUAUGAAGGAUACAACCUUCUCGCGUUGCUCGGCCACAGUUUGCCAAAUGGAU ---------..........((((..((((.(((.(((((..(((((..((.(((....)))))..(((((......)))))..))).))..))))).))).))))..))))... ( -30.10) >DroMoj_CAF1 2498 112 + 1 CGAACUG--GCACUCAAAACAUUGAGCAUCUUGCGCUUGGCAAAGCUCAGACUGGUCAGAUCAAUGAAGGAGACAACUCUCUCGCGCUGCUCAGCCACCGUUUGCCAAAUGCUU .....((--(((..(......((((((...((((.....)))).))))))..((((..((.((.((..(((((....)))))..)).)).)).))))..)..)))))....... ( -29.60) >DroAna_CAF1 2131 111 + 1 ---GUCCCAGCCGACAUCACACUGAGCAACUUCCGAUUCUCCAUACUCAGCGUGGCGAGAUUAAUGAAGGACACGACCUUCUCCCGCUGCUCGGCAACUGUCCGCCAAAUGGAC ---......(((((((.(((.(((((...................))))).)))(((.((.....(((((......))))))).))))).)))))....(((((.....))))) ( -31.01) >consensus _________GAUGUCACCACAUUGAGCAGCUUGCGGUUGUCCAAGCUCAAUGUGGUGAGAUCUAUGAAGGACACAACCUUCUCGCGCUGCUCGGCCACCGUUUGCCAAAUGGAU ................(((....(((((((..((((.........((((......))))......(((((......))))))))))))))))(((........)))...))).. (-21.47 = -23.13 + 1.67)

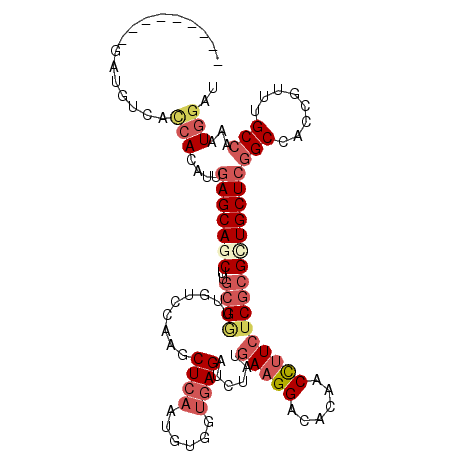
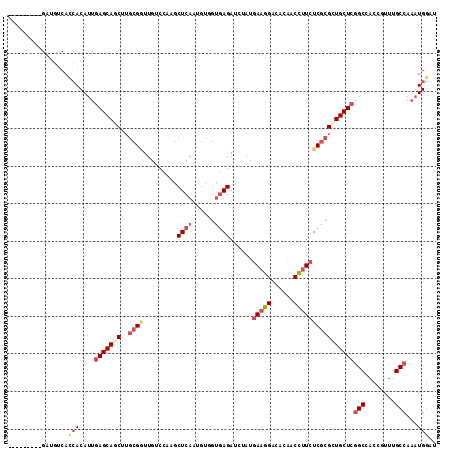
| Location | 14,915,728 – 14,915,833 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 79.71 |
| Mean single sequence MFE | -37.13 |
| Consensus MFE | -24.34 |
| Energy contribution | -24.40 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.725592 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 14915728 105 - 20766785 AUCCAUUUGGCAACGGGUGGCCGAGCAGCGCGAGAAGGUUGUGUCCUUCAUAGAUCUCACAACAUUGAGCUUGGACAACCGCAAGCUGCUCAAUGUGGUGACAUC--------- ..(((((((....)))))))..((((((((((.(((((......)))))...((.((((......)))).)).......)))..)))))))...(((....))).--------- ( -38.20) >DroSec_CAF1 2165 105 - 1 AUCCAUAUGGCAACGGGUGGCCGAGCAGCGCGAUAAGGUUGUGUCCUUCAUAGAUCUCACCACAUUGAGCAUGGACAAUCGCAAGCUGCUCAAUGUGGUGACAUC--------- ..((((.((....)).))))..(((..(((((((...)))))))..)))...(((.((((((((((((((((((.....).))...))))))))))))))).)))--------- ( -40.00) >DroEre_CAF1 2322 105 - 1 AUCCAUUUGGCAAACGGUGGCCGAGCAGCGCGAGAAGGUCGUGUCCUUCAUAGAUCUCACGACGUUGAGCUUGGACAACCGCAAGCUGCUCAAUGUGGUGACAUC--------- ........(((........)))((((((((((.....((((((..((....))....))))))((((........)))))))..)))))))...(((....))).--------- ( -34.40) >DroYak_CAF1 2131 105 - 1 AUCCAUUUGGCAAACUGUGGCCGAGCAACGCGAGAAGGUUGUAUCCUUCAUAGAUCUCACGACACUGAGCAUGGACAACCGCAAGCUGCUGAAUGUGGUGACAUC--------- ...((((..(((...((((((((.((.......(((((......))))).......(((......))))).)))....)))))...)))..))))((....))..--------- ( -29.20) >DroMoj_CAF1 2498 112 - 1 AAGCAUUUGGCAAACGGUGGCUGAGCAGCGCGAGAGAGUUGUCUCCUUCAUUGAUCUGACCAGUCUGAGCUUUGCCAAGCGCAAGAUGCUCAAUGUUUUGAGUGC--CAGUUCG ..((.(((((((((((((((..(((((((.(....).)))).)))..)))))).((.((....)).))..))))))))).))..(..((((((....))))))..--)...... ( -36.60) >DroAna_CAF1 2131 111 - 1 GUCCAUUUGGCGGACAGUUGCCGAGCAGCGGGAGAAGGUCGUGUCCUUCAUUAAUCUCGCCACGCUGAGUAUGGAGAAUCGGAAGUUGCUCAGUGUGAUGUCGGCUGGGAC--- ((((....(((.((((...........(((((((((((......))))).....))))))(((((((((((........(....).))))))))))).)))).))).))))--- ( -44.40) >consensus AUCCAUUUGGCAAACGGUGGCCGAGCAGCGCGAGAAGGUUGUGUCCUUCAUAGAUCUCACCACACUGAGCAUGGACAACCGCAAGCUGCUCAAUGUGGUGACAUC_________ (((((((.(((........)))((((((((((.(((((......)))))......((((......))))..........)))..))))))))))).)))............... (-24.34 = -24.40 + 0.06)
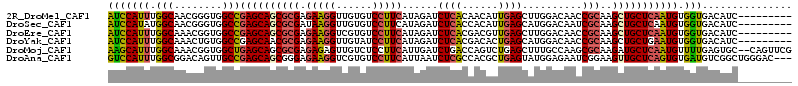
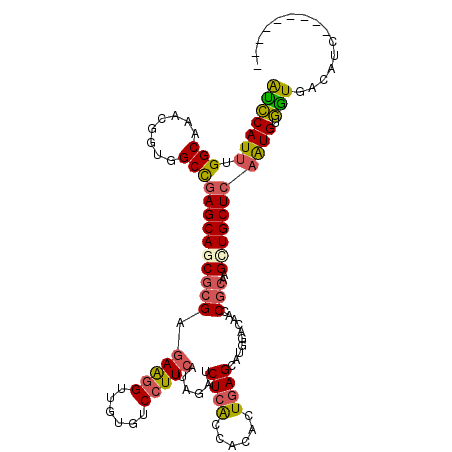
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:23:05 2006