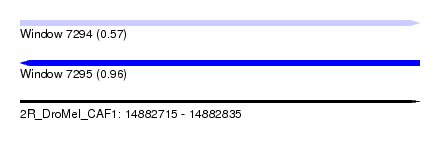

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 14,882,715 – 14,882,835 |
| Length | 120 |
| Max. P | 0.956219 |

| Location | 14,882,715 – 14,882,835 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 73.94 |
| Mean single sequence MFE | -35.37 |
| Consensus MFE | -19.71 |
| Energy contribution | -19.47 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.61 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.567134 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
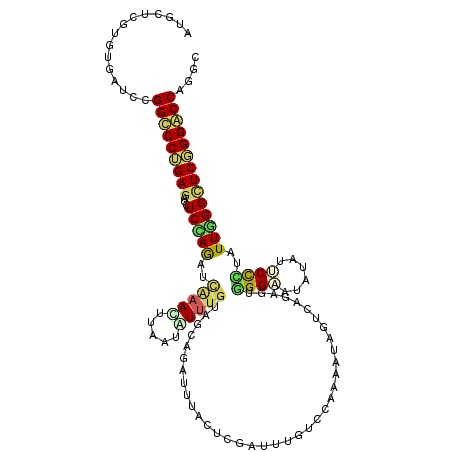
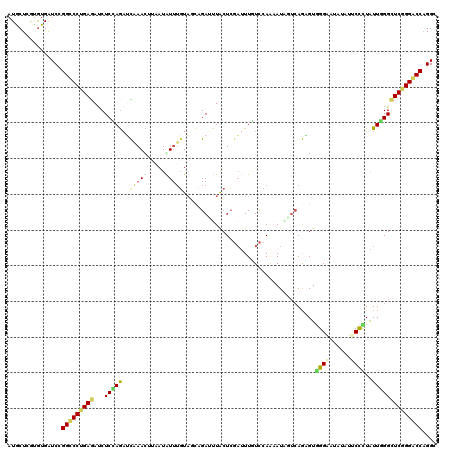
>2R_DroMel_CAF1 14882715 120 + 20766785 AUGCUCGUGUGAUCCGGUCCUGAGAUCUCAAGAUCAAACCUAAUAUUUGUAUCUGAAUUACUCGAUUUUUCCAAAAUAGUCAGAGUGGGAAUAUAUUCCCUAUUUGGCUCGGGACCAGGA ............(((((((((((((((....))))........(((((......(((((....))))).....)))))((((((..(((((....)))))..)))))))))))))).))) ( -34.10) >DroPse_CAF1 61793 120 + 1 AUACUCGUGUGGUCGGGCCCCGAGAUCUCUAGUGUGAACUUUAUGUUUUUAGCAGAGCUGCUGGACUUGUCGAUUAUGCUCAAUGUGGGCAAGUAGUCCUUGUUGGGCUCUGGGCCCGGG .(((....))).(((((((((((.(..((((((..((((.....))))..(((...)))))))))..).))).....((((((((.((((.....)))).))))))))...)))))))). ( -43.70) >DroGri_CAF1 60787 120 + 1 AUGCUUGUAUGAUCCGGUCCUGAAACCUCCAAAGUGAAUAUAAUAUUUGUCAUCGAAGUGCUCUCUUUGUUUAAUAUUUUCAACGUUGGCAUAUAGGCUUUGUUGGGCUCGGGACCAGGC .............((((((((((..((..((((((..........((((....))))(((((..(.(((...........))).)..)))))....))))))..))..)))))))).)). ( -32.80) >DroSim_CAF1 56597 120 + 1 AUGCUCGUGUGAUCCGGCCCUGAGAUCUCAAGAUCAAACUUAAUAUUUGUAGCUGAUUUACUCGAUUUUUCCAAAAUCGUCAGAGUGGGAAUGUAUUCCCUAUUUGGCUCGGGGCCAGGU .............((((((((((((((....))))............................(((((.....)))))((((((..(((((....)))))..)))))))))))))).)). ( -35.40) >DroYak_CAF1 56723 120 + 1 AUGCUCGUGUGAUCUGGCCCUGAGAUCUCCAGAUCAAAUUUAAUAUUUGUAUCCGAUUUGCUCGAUUUAUCCAAAAUAGUCAGUGUGGGAAUAUACUCCCUAUUUGGCUCGGGACCAGGC ............(((((.((((((.......((.((((((..(((....)))..)))))).)).......(((((.(((..((((((....))))))..)))))))))))))).))))). ( -32.00) >DroAna_CAF1 63280 120 + 1 AUGCUGAUGUGAUCAGGCCCAGAGAUCUCCAGAUCGAAUUGUAUGUGGGUAGCAGAUUUACUCGACUUGUCCAGAAUUUGAAGAGUAGGAAUAUAUCCCUUGUUGGGUUCUGGGCCAGGC ....(((.((((((..(((((.(((((....))))........).)))))....)).))))))).((.((((((((((..(.(((..(((.....)))))).)..)))))))))).)).. ( -34.20) >consensus AUGCUCGUGUGAUCCGGCCCUGAGAUCUCCAGAUCAAACUUAAUAUUUGUAGCAGAUUUACUCGAUUUGUCCAAAAUAGUCAGAGUGGGAAUAUAUUCCCUAUUGGGCUCGGGACCAGGC ...............(((((((((...(((((..(((((.....))))).....................................((((......))))..)))))))))))))).... (-19.71 = -19.47 + -0.24)
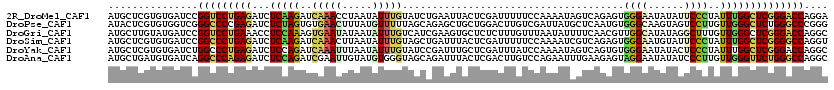

| Location | 14,882,715 – 14,882,835 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 73.94 |
| Mean single sequence MFE | -31.44 |
| Consensus MFE | -18.43 |
| Energy contribution | -18.60 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.67 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.59 |
| SVM decision value | 1.46 |
| SVM RNA-class probability | 0.956219 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
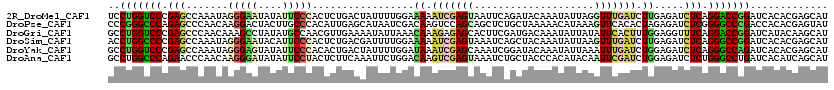
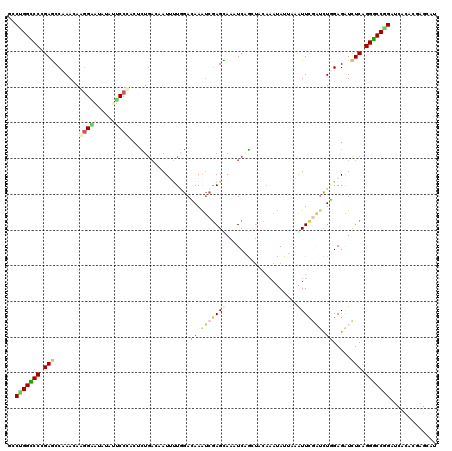
>2R_DroMel_CAF1 14882715 120 - 20766785 UCCUGGUCCCGAGCCAAAUAGGGAAUAUAUUCCCACUCUGACUAUUUUGGAAAAAUCGAGUAAUUCAGAUACAAAUAUUAGGUUUGAUCUUGAGAUCUCAGGACCGGAUCACACGAGCAU ..(((((((.(((((.(((((((((....)))))..(((((....(((((.....)))))....)))))......)))).)))))((((....))))...)))))))............. ( -33.80) >DroPse_CAF1 61793 120 - 1 CCCGGGCCCAGAGCCCAACAAGGACUACUUGCCCACAUUGAGCAUAAUCGACAAGUCCAGCAGCUCUGCUAAAAACAUAAAGUUCACACUAGAGAUCUCGGGGCCCGACCACACGAGUAU ..(((((((.(((..(..((((.....)))).......(((((......((....)).((((....))))...........))))).......)..))).)))))))............. ( -31.80) >DroGri_CAF1 60787 120 - 1 GCCUGGUCCCGAGCCCAACAAAGCCUAUAUGCCAACGUUGAAAAUAUUAAACAAAGAGAGCACUUCGAUGACAAAUAUUAUAUUCACUUUGGAGGUUUCAGGACCGGAUCAUACAAGCAU ..(((((((.(((.((..(((((...(((((....(((((((.....................)))))))........)))))...)))))..)).))).)))))))............. ( -26.30) >DroSim_CAF1 56597 120 - 1 ACCUGGCCCCGAGCCAAAUAGGGAAUACAUUCCCACUCUGACGAUUUUGGAAAAAUCGAGUAAAUCAGCUACAAAUAUUAAGUUUGAUCUUGAGAUCUCAGGGCCGGAUCACACGAGCAU ..(((((((.(((.......(((((....))))).(((.((((((((.....)))))).............(((((.....))))).))..)))..))).)))))))............. ( -33.50) >DroYak_CAF1 56723 120 - 1 GCCUGGUCCCGAGCCAAAUAGGGAGUAUAUUCCCACACUGACUAUUUUGGAUAAAUCGAGCAAAUCGGAUACAAAUAUUAAAUUUGAUCUGGAGAUCUCAGGGCCAGAUCACACGAGCAU (((((((((.(((.......(((((....)))))..........(((..(((...((((.....))))...(((((.....))))))))..)))..))).))))))).((....)))).. ( -32.20) >DroAna_CAF1 63280 120 - 1 GCCUGGCCCAGAACCCAACAAGGGAUAUAUUCCUACUCUUCAAAUUCUGGACAAGUCGAGUAAAUCUGCUACCCACAUACAAUUCGAUCUGGAGAUCUCUGGGCCUGAUCACAUCAGCAU ((..((((((((.(((.....)))...........(((((((.....))))((..((((((....................))))))..)))))...))))))))((((...)))))).. ( -31.05) >consensus GCCUGGCCCCGAGCCAAACAAGGAAUAUAUUCCCACUCUGACAAUUUUGGACAAAUCGAGCAAAUCAGCUACAAAUAUUAAAUUCGAUCUGGAGAUCUCAGGGCCGGAUCACACGAGCAU ..(((((((.(((.......(((((....))))).................((.(((((((....................))))))).)).....))).)))))))............. (-18.43 = -18.60 + 0.17)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:22:51 2006