
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 14,865,291 – 14,865,382 |
| Length | 91 |
| Max. P | 0.976162 |

| Location | 14,865,291 – 14,865,382 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 90.23 |
| Mean single sequence MFE | -32.28 |
| Consensus MFE | -27.06 |
| Energy contribution | -26.75 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.60 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.76 |
| SVM RNA-class probability | 0.976162 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
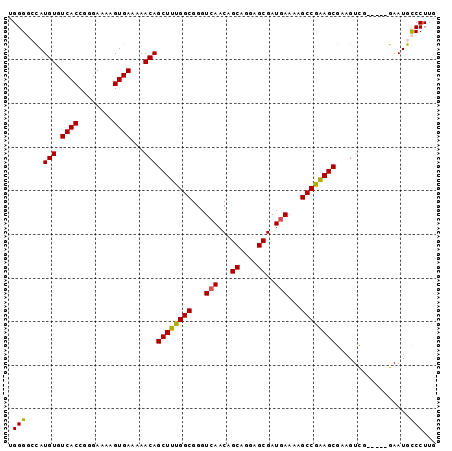
>2R_DroMel_CAF1 14865291 91 + 20766785 UGGG--CAUGUGUCACCGGGAAAAGUGAAAAACAGCUUUGGCGGGUCAACAGCAGGAGCGAUGAAAAGCCGAAGCGAAGUCG-----GAAUGCCCUUG .(((--(((((.((((........))))...)).((((((((...(((...((....))..)))...)))))))).......-----..))))))... ( -33.50) >DroSec_CAF1 37261 93 + 1 UGGGGCCAUGUGUCACCGGGAAAAGUGAAAAACAGCUUUGGCGGGUCAACAGCUGGAGCGAUAAAAAGCCGGAGCGAAGUCG-----GAAUGCCCUUG .(((((..(((.((((........))))...)))((((((((.........((....))........)))))))).......-----....))))).. ( -28.63) >DroSim_CAF1 39374 93 + 1 UGGGGCCAUGUGUCACCGGGAAAAGUGAAAAACAGCUUUGGCGGGUCAACAGCUGGAGCGAUGAAAAGCCGAAGCGAAGUCG-----GAAUGCCCUUG .(((((..(((.((((........))))...)))((((((((...(((...((....))..)))...)))))))).......-----....))))).. ( -32.50) >DroYak_CAF1 39021 98 + 1 UGGGCAUAUGUGUCACCCGGAAAAGUGAAAAACAGCUUUGGCGGGUCAACAGCAGCAGCGAUGAAAAGCCAAAGCGACGUUGAACACGAAUGUCCUUG .((((((.(((((((..((.....((.....)).((((((((...(((...((....))..)))...))))))))..)).)).))))).))))))... ( -34.50) >consensus UGGGGCCAUGUGUCACCGGGAAAAGUGAAAAACAGCUUUGGCGGGUCAACAGCAGGAGCGAUGAAAAGCCGAAGCGAAGUCG_____GAAUGCCCUUG .(((....(((.((((........))))...)))((((((((...(((...((....))..)))...)))))))).................)))... (-27.06 = -26.75 + -0.31)


| Location | 14,865,291 – 14,865,382 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 90.23 |
| Mean single sequence MFE | -25.42 |
| Consensus MFE | -18.99 |
| Energy contribution | -19.30 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.678229 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
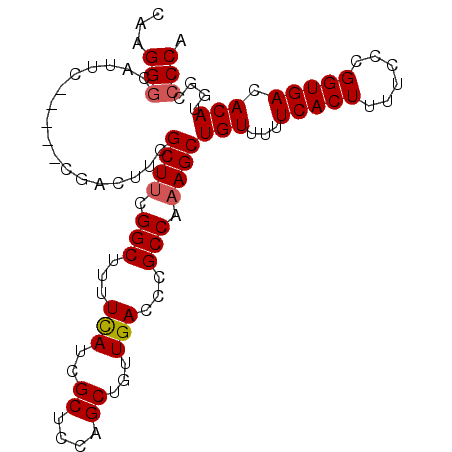
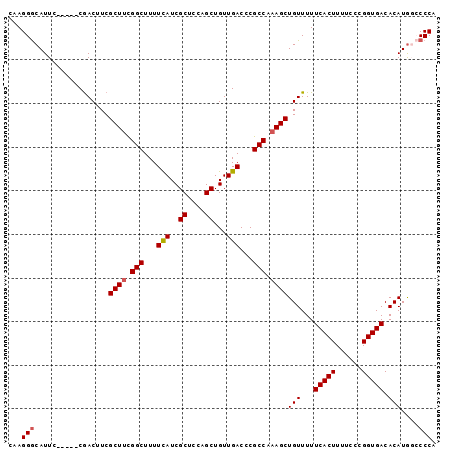
>2R_DroMel_CAF1 14865291 91 - 20766785 CAAGGGCAUUC-----CGACUUCGCUUCGGCUUUUCAUCGCUCCUGCUGUUGACCCGCCAAAGCUGUUUUUCACUUUUCCCGGUGACACAUG--CCCA ...((((((..-----.......((((.(((...(((..((....))...)))...))).))))......(((((......)))))...)))--))). ( -26.90) >DroSec_CAF1 37261 93 - 1 CAAGGGCAUUC-----CGACUUCGCUCCGGCUUUUUAUCGCUCCAGCUGUUGACCCGCCAAAGCUGUUUUUCACUUUUCCCGGUGACACAUGGCCCCA ...(((.....-----((((...(((..(((........)))..))).))))....((((....(((...(((((......))))).)))))))))). ( -21.90) >DroSim_CAF1 39374 93 - 1 CAAGGGCAUUC-----CGACUUCGCUUCGGCUUUUCAUCGCUCCAGCUGUUGACCCGCCAAAGCUGUUUUUCACUUUUCCCGGUGACACAUGGCCCCA ...((((....-----.......((((.(((...(((..((....))...)))...))).))))(((...(((((......))))).)))..)))).. ( -24.50) >DroYak_CAF1 39021 98 - 1 CAAGGACAUUCGUGUUCAACGUCGCUUUGGCUUUUCAUCGCUGCUGCUGUUGACCCGCCAAAGCUGUUUUUCACUUUUCCGGGUGACACAUAUGCCCA ...((.(((..(((.........((((((((...(((..((....))...)))...))))))))......(((((......))))))))..))).)). ( -28.40) >consensus CAAGGGCAUUC_____CGACUUCGCUUCGGCUUUUCAUCGCUCCAGCUGUUGACCCGCCAAAGCUGUUUUUCACUUUUCCCGGUGACACAUGGCCCCA ...(((.................((((.(((...(((..((....))...)))...))).))))(((...(((((......))))).)))....))). (-18.99 = -19.30 + 0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:22:44 2006