
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 14,768,772 – 14,768,864 |
| Length | 92 |
| Max. P | 0.873504 |

| Location | 14,768,772 – 14,768,864 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 81.61 |
| Mean single sequence MFE | -23.02 |
| Consensus MFE | -12.46 |
| Energy contribution | -14.66 |
| Covariance contribution | 2.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.772292 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
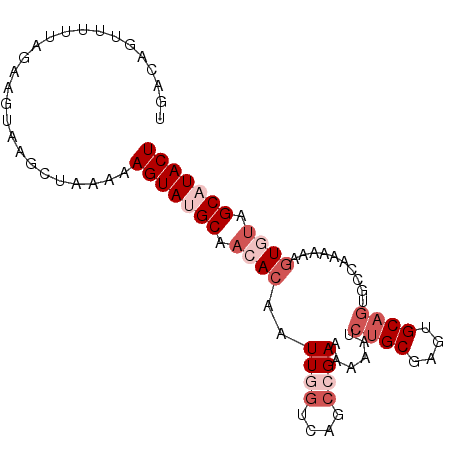
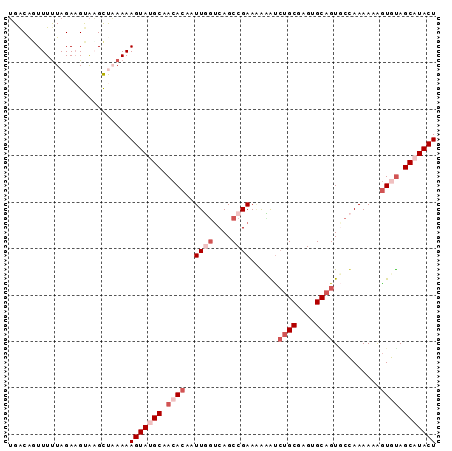
>2R_DroMel_CAF1 14768772 92 + 20766785 UGACAGUUUCUAGAAGUAAGCUAAAAAGUAUGCAACACAAUUGGUCAGCCGAAGAUAUAAGCGAGAGCAGUGCCAAAAAAGUGUAGCAUACU ....((((((.....).)))))....(((((((.((((..((((....))))........((....))............)))).))))))) ( -23.10) >DroSec_CAF1 3635 91 + 1 UGACGUUUUUUAGAAGUAAGCUAAAAAGUAUGCAACACAAUUGGACAGCCGAAAGCAUCUGCGAGUGCAGUGCCAAAA-AAUGCAGCAUACU ......(((((((.......)))))))((((((.......((((....))))..(((.((((....))))))).....-......)))))). ( -22.00) >DroSim_CAF1 10131 91 + 1 UGACGGUUUUUAGAAGUAAGCUAAAAAGUAUGCAACACAAUUGGUCAGCCGAAAGCAUCUGCGAGUGCAGUGCCAAAA-AGUGUAGCAUACU ......(((((((.......)))))))((((((.((((..((((....))))..(((.((((....))))))).....-.)))).)))))). ( -28.30) >DroEre_CAF1 2498 92 + 1 GGACAGUUUUUAGAAGCGCGUCCAAAAGUAGGCAACACAAUUUGUCAGCCGAAAAAGCCUGCAUGUGCAGUACUAAAAAGGUAUAGCAUACU ((((.((((....))))..))))....((((((.......................)))))).(((((.(((((.....))))).))))).. ( -25.20) >DroYak_CAF1 9728 92 + 1 UGGCAGUUUUUAGAUGCAUGCCCAAAAGUAGGCAACACAAUUUUUCAGCCGAAUAAAUCUGCAUGUGCAGUCCUAAAAAAGUAUAGCAUACU ..((..(((((((.((((((((........)))..............((.((.....)).))..)))))...)))))))......))..... ( -16.50) >consensus UGACAGUUUUUAGAAGUAAGCUAAAAAGUAUGCAACACAAUUGGUCAGCCGAAAAAAUCUGCGAGUGCAGUGCCAAAAAAGUGUAGCAUACU ..........................(((((((.((((..((((....))))......((((....))))..........)))).))))))) (-12.46 = -14.66 + 2.20)

| Location | 14,768,772 – 14,768,864 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 81.61 |
| Mean single sequence MFE | -22.17 |
| Consensus MFE | -14.72 |
| Energy contribution | -15.00 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873504 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
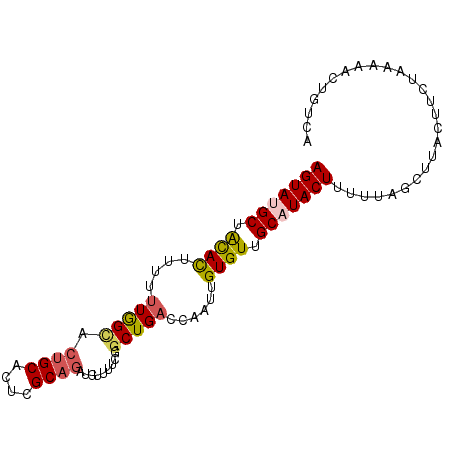
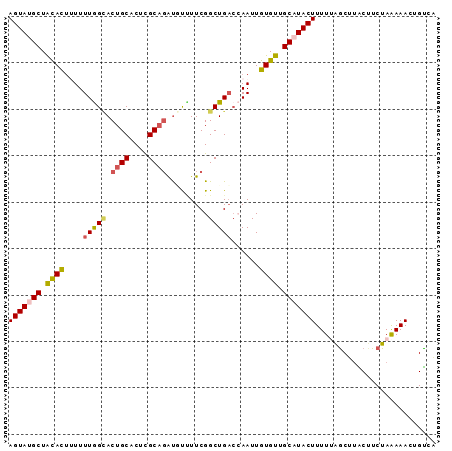
>2R_DroMel_CAF1 14768772 92 - 20766785 AGUAUGCUACACUUUUUUGGCACUGCUCUCGCUUAUAUCUUCGGCUGACCAAUUGUGUUGCAUACUUUUUAGCUUACUUCUAGAAACUGUCA .((((((.((((....((((((..((....))......(....).)).))))..)))).))))))(((((((.......)))))))...... ( -20.00) >DroSec_CAF1 3635 91 - 1 AGUAUGCUGCAUU-UUUUGGCACUGCACUCGCAGAUGCUUUCGGCUGUCCAAUUGUGUUGCAUACUUUUUAGCUUACUUCUAAAAAACGUCA .((((((.((((.-..((((((((((....))))..((.....)))).))))..)))).))))))(((((((.......)))))))...... ( -25.50) >DroSim_CAF1 10131 91 - 1 AGUAUGCUACACU-UUUUGGCACUGCACUCGCAGAUGCUUUCGGCUGACCAAUUGUGUUGCAUACUUUUUAGCUUACUUCUAAAAACCGUCA .((((((.((((.-..((((((((((....))))..((.....)))).))))..)))).))))))(((((((.......)))))))...... ( -26.40) >DroEre_CAF1 2498 92 - 1 AGUAUGCUAUACCUUUUUAGUACUGCACAUGCAGGCUUUUUCGGCUGACAAAUUGUGUUGCCUACUUUUGGACGCGCUUCUAAAAACUGUCC .((((((((........))))).)))....((((........(((.((((.....)))))))...(((((((......))))))).)))).. ( -18.30) >DroYak_CAF1 9728 92 - 1 AGUAUGCUAUACUUUUUUAGGACUGCACAUGCAGAUUUAUUCGGCUGAAAAAUUGUGUUGCCUACUUUUGGGCAUGCAUCUAAAAACUGCCA ......(((........)))..((((....))))........(((.........((((((((((....)))))).)))).........))). ( -20.67) >consensus AGUAUGCUACACUUUUUUGGCACUGCACUCGCAGAUGUUUUCGGCUGACCAAUUGUGUUGCAUACUUUUUAGCUUACUUCUAAAAACUGUCA (((((((.((((....(((((.((((....)))).........)))))......)))).))))))).......................... (-14.72 = -15.00 + 0.28)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:21:17 2006