
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 2,569,694 – 2,569,800 |
| Length | 106 |
| Max. P | 0.999987 |

| Location | 2,569,694 – 2,569,800 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 72.21 |
| Mean single sequence MFE | -37.80 |
| Consensus MFE | -25.77 |
| Energy contribution | -27.08 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.22 |
| Mean z-score | -4.67 |
| Structure conservation index | 0.68 |
| SVM decision value | 5.35 |
| SVM RNA-class probability | 0.999984 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 2569694 106 + 20766785 UUUGGUAAAGACUGGUUCCACUGUCACACAUAUGUUGCUGCUGCUCAUGUUUACGCGCAGUUAAAUAUUAAAUUGCGACAGUGAAAUCAGGCUUUGUUAGACAAC--------A----- (((..(((((.((((((.(((((((.((...(((((...(((((.(........).)))))..))))).....)).))))))).)))))).)))))..)))....--------.----- ( -34.60) >DroSec_CAF1 2978 86 + 1 UUUGGUAAAGCCUGGUUUCACUGCCACGCAUAUUUUGCUGCUACUCA--------------------UUAAAUUGGGACAGUGAAAUCAGGCUAUGGUAGACAGC--------A----- ((((.((.(((((((((((((((....(((.....))).....((((--------------------......)))).))))))))))))))).)).))))....--------.----- ( -33.00) >DroSim_CAF1 2989 86 + 1 UUUGGUAAAGCCUGGUUUCACUGCCACGCAUAUUUUGCUGCUGCUCA--------------------UUAAAUUGGGACAGUGAAAUCAGGCUAUGAUAGACAGC--------A----- ........(((((((((((((((((..(((.....))).......((--------------------......)))).)))))))))))))))............--------.----- ( -30.80) >DroEre_CAF1 2829 119 + 1 UUUGGUAUGGCUUGGUUCCACUGCCACACAUAUUUUGCUACUGCUUGUGUUUUCGCGCAGUUAAAAGUAAAAUUGCGGCAGUGAAAUCAGGCUAUGCUAGACAGCUGGACAGCAGGUCU ..(((((((((((((((.(((((((.((...(((((((((((((..(((....))))))))....)))))))))).))))))).)))))))))))))))(((.(((....)))..))). ( -52.80) >consensus UUUGGUAAAGCCUGGUUCCACUGCCACACAUAUUUUGCUGCUGCUCA____________________UUAAAUUGCGACAGUGAAAUCAGGCUAUGAUAGACAGC________A_____ (((((((((((((((((((((((((.((.............................................)).))))))))))))))))))))))))).................. (-25.77 = -27.08 + 1.31)
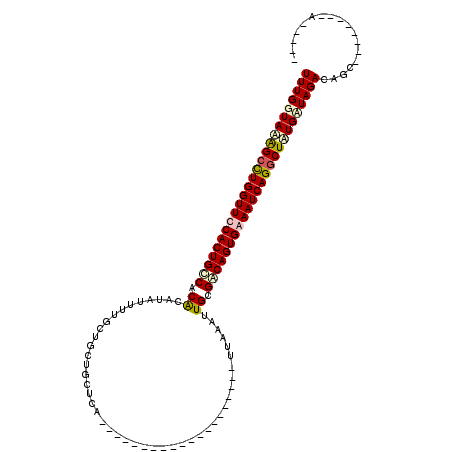


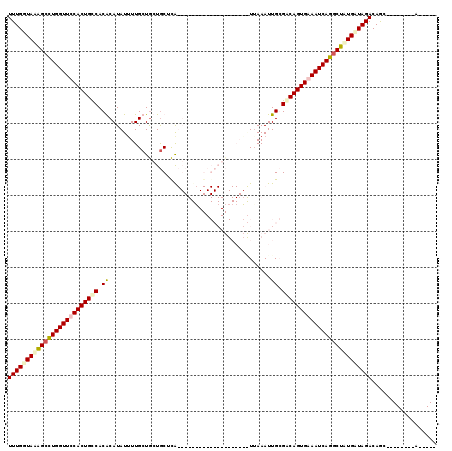
| Location | 2,569,694 – 2,569,800 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 72.21 |
| Mean single sequence MFE | -30.38 |
| Consensus MFE | -23.58 |
| Energy contribution | -24.20 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.24 |
| Mean z-score | -3.47 |
| Structure conservation index | 0.78 |
| SVM decision value | 5.46 |
| SVM RNA-class probability | 0.999987 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
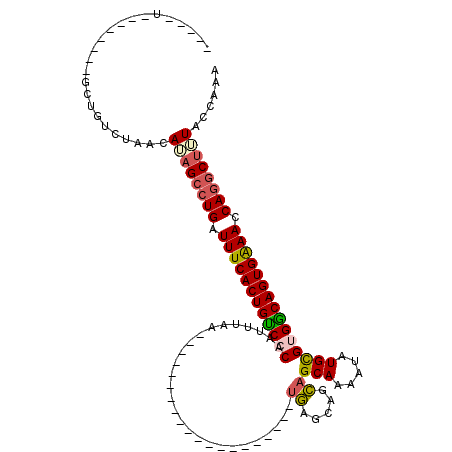
>2R_DroMel_CAF1 2569694 106 - 20766785 -----U--------GUUGUCUAACAAAGCCUGAUUUCACUGUCGCAAUUUAAUAUUUAACUGCGCGUAAACAUGAGCAGCAGCAACAUAUGUGUGACAGUGGAACCAGUCUUUACCAAA -----.--------..........((((.(((.(((((((((((((.....((((....((((((..........)).))))....)))).))))))))))))).))).))))...... ( -32.50) >DroSec_CAF1 2978 86 - 1 -----U--------GCUGUCUACCAUAGCCUGAUUUCACUGUCCCAAUUUAA--------------------UGAGUAGCAGCAAAAUAUGCGUGGCAGUGAAACCAGGCUUUACCAAA -----.--------............((((((.((((((((((.((......--------------------))....(((........)))..)))))))))).))))))........ ( -25.90) >DroSim_CAF1 2989 86 - 1 -----U--------GCUGUCUAUCAUAGCCUGAUUUCACUGUCCCAAUUUAA--------------------UGAGCAGCAGCAAAAUAUGCGUGGCAGUGAAACCAGGCUUUACCAAA -----.--------............((((((.((((((((((.((......--------------------))....(((........)))..)))))))))).))))))........ ( -25.90) >DroEre_CAF1 2829 119 - 1 AGACCUGCUGUCCAGCUGUCUAGCAUAGCCUGAUUUCACUGCCGCAAUUUUACUUUUAACUGCGCGAAAACACAAGCAGUAGCAAAAUAUGUGUGGCAGUGGAACCAAGCCAUACCAAA ((((..(((....))).))))......((.((.((((((((((((((((((.((....(((((............))))))).)))))...))))))))))))).)).))......... ( -37.20) >consensus _____U________GCUGUCUAACAUAGCCUGAUUUCACUGUCCCAAUUUAA____________________UGAGCAGCAGCAAAAUAUGCGUGGCAGUGAAACCAGGCUUUACCAAA ........................((((((((.((((((((((((...........................((.....))(((.....))))))))))))))).))))))))...... (-23.58 = -24.20 + 0.62)

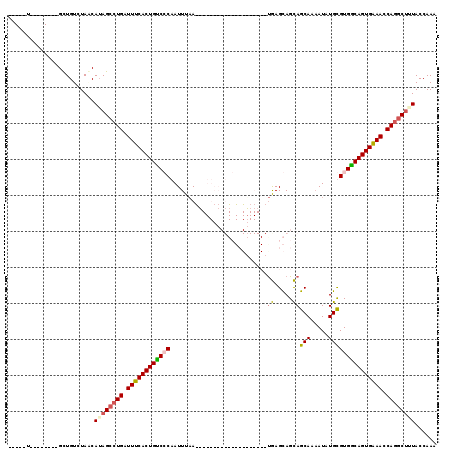
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:38:48 2006