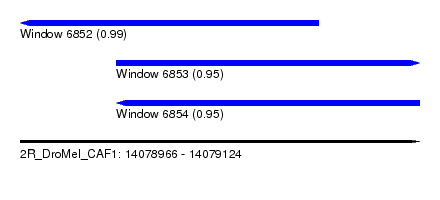

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 14,078,966 – 14,079,124 |
| Length | 158 |
| Max. P | 0.990820 |

| Location | 14,078,966 – 14,079,084 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.53 |
| Mean single sequence MFE | -36.03 |
| Consensus MFE | -34.82 |
| Energy contribution | -35.27 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.16 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.23 |
| SVM RNA-class probability | 0.990820 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 14078966 118 - 20766785 GGAGCUGGGACAUCGAUGAUGGCCACUAAGCGCAUUAAUUUAAUGCUCUCAACAGCUGGCCACGCCGUGCAUAAAUAAAAUGUCAUCGCAUACCAAACCGUUGCGUCUCAUUCCAU--CC ((((.((((((...(((((((((((((.((.((((((...)))))).))....)).)))))..((...))...........))))))(((.((......)))))))))))))))..--.. ( -36.80) >DroSim_CAF1 3492 118 - 1 GGAGCUGGGACAUCGAUGAUGGCCACUAAGCGCAUUAAUUUAAUGCUCUCAACAGCUGGCCACGCCGUGCAUAAAUAAAAUGUCAUCGCAUACCAAUCCAUUGCGUCCCAUUCCAU--CC ((((.((((((..(((((.((((((((.((.((((((...)))))).))....)).)))))).((.((((((.......))).))).)).........))))).))))))))))..--.. ( -39.60) >DroYak_CAF1 4181 120 - 1 GGACCUGGGACAUCGAUGAUGGCCACUAAGCGCAUUAAUUUAAUGCUCUCAACAGCUGGCCACGCCGUGCAUAAAUAAAAUGUCAUCGCAUACCAAUCCAUUGCAUCUCAUUCCGUCUCC (((..(((((...(((((.((((((((.((.((((((...)))))).))....)).)))))).((.((((((.......))).))).)).........)))))..))))).)))...... ( -31.70) >consensus GGAGCUGGGACAUCGAUGAUGGCCACUAAGCGCAUUAAUUUAAUGCUCUCAACAGCUGGCCACGCCGUGCAUAAAUAAAAUGUCAUCGCAUACCAAUCCAUUGCGUCUCAUUCCAU__CC ((((.((((((...(((((((((((((.((.((((((...)))))).))....)).)))))..((...))...........))))))(((...........)))))))))))))...... (-34.82 = -35.27 + 0.45)


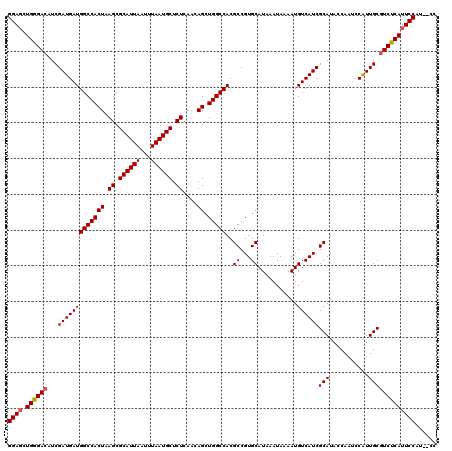
| Location | 14,079,004 – 14,079,124 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 99.44 |
| Mean single sequence MFE | -37.73 |
| Consensus MFE | -34.93 |
| Energy contribution | -35.27 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.97 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952584 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 14079004 120 + 20766785 UUUUAUUUAUGCACGGCGUGGCCAGCUGUUGAGAGCAUUAAAUUAAUGCGCUUAGUGGCCAUCAUCGAUGUCCCAGCUCCGCGUCUGGCAAUUGAUAUUUUAAUCGCCAUUUCAUUUUGU ..............(((((((..((((((((((.((((((...)))))).))))).((.((((...)))).))))))))))))))((((.(((((....))))).))))........... ( -37.10) >DroSim_CAF1 3530 120 + 1 UUUUAUUUAUGCACGGCGUGGCCAGCUGUUGAGAGCAUUAAAUUAAUGCGCUUAGUGGCCAUCAUCGAUGUCCCAGCUCCGCGUCUGGCAAUUGAUAUUUUAAUCGCCAUUUCAUUUUGU ..............(((((((..((((((((((.((((((...)))))).))))).((.((((...)))).))))))))))))))((((.(((((....))))).))))........... ( -37.10) >DroYak_CAF1 4221 120 + 1 UUUUAUUUAUGCACGGCGUGGCCAGCUGUUGAGAGCAUUAAAUUAAUGCGCUUAGUGGCCAUCAUCGAUGUCCCAGGUCCGCGUCUGGCAAUUGAUAUUUUAAUCGCCAUUUCAUUUUGU ..............(((((((((.((..(((((.((((((...)))))).)))))..))((((...)))).....)).)))))))((((.(((((....))))).))))........... ( -39.00) >consensus UUUUAUUUAUGCACGGCGUGGCCAGCUGUUGAGAGCAUUAAAUUAAUGCGCUUAGUGGCCAUCAUCGAUGUCCCAGCUCCGCGUCUGGCAAUUGAUAUUUUAAUCGCCAUUUCAUUUUGU ..............(((((((..((((((((((.((((((...)))))).))))).((.((((...)))).))))))))))))))((((.(((((....))))).))))........... (-34.93 = -35.27 + 0.33)

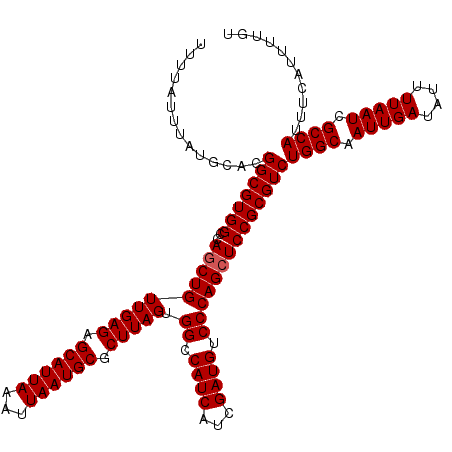
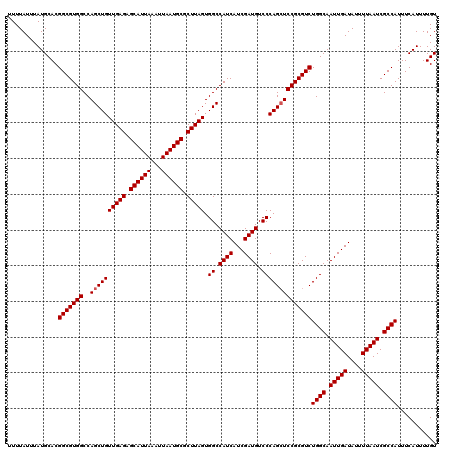
| Location | 14,079,004 – 14,079,124 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 99.44 |
| Mean single sequence MFE | -31.97 |
| Consensus MFE | -31.57 |
| Energy contribution | -31.57 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.36 |
| SVM RNA-class probability | 0.945788 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 14079004 120 - 20766785 ACAAAAUGAAAUGGCGAUUAAAAUAUCAAUUGCCAGACGCGGAGCUGGGACAUCGAUGAUGGCCACUAAGCGCAUUAAUUUAAUGCUCUCAACAGCUGGCCACGCCGUGCAUAAAUAAAA .....(((..((((((((......)))....(((((.(((..((.(((..((((...)))).)))))..))((((((...))))))........))))))...))))).)))........ ( -32.10) >DroSim_CAF1 3530 120 - 1 ACAAAAUGAAAUGGCGAUUAAAAUAUCAAUUGCCAGACGCGGAGCUGGGACAUCGAUGAUGGCCACUAAGCGCAUUAAUUUAAUGCUCUCAACAGCUGGCCACGCCGUGCAUAAAUAAAA .....(((..((((((((......)))....(((((.(((..((.(((..((((...)))).)))))..))((((((...))))))........))))))...))))).)))........ ( -32.10) >DroYak_CAF1 4221 120 - 1 ACAAAAUGAAAUGGCGAUUAAAAUAUCAAUUGCCAGACGCGGACCUGGGACAUCGAUGAUGGCCACUAAGCGCAUUAAUUUAAUGCUCUCAACAGCUGGCCACGCCGUGCAUAAAUAAAA .....(((..((((((.......((((.(((.((((........)))).).)).)))).((((((((.((.((((((...)))))).))....)).)))))))))))).)))........ ( -31.70) >consensus ACAAAAUGAAAUGGCGAUUAAAAUAUCAAUUGCCAGACGCGGAGCUGGGACAUCGAUGAUGGCCACUAAGCGCAUUAAUUUAAUGCUCUCAACAGCUGGCCACGCCGUGCAUAAAUAAAA .....(((..((((((.......((((.(((.((((........)))).).)).)))).((((((((.((.((((((...)))))).))....)).)))))))))))).)))........ (-31.57 = -31.57 + -0.00)

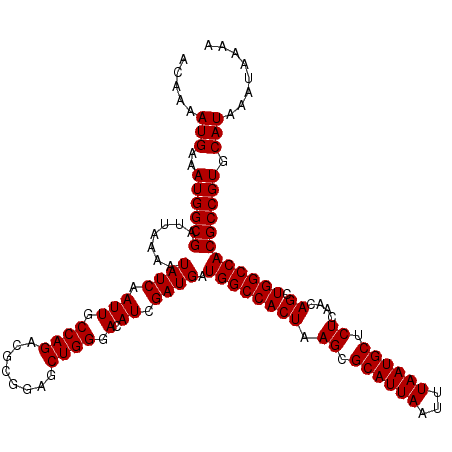
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:15:47 2006