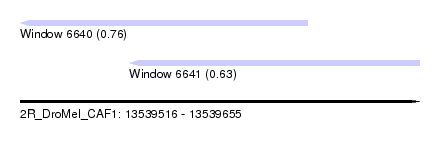

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 13,539,516 – 13,539,655 |
| Length | 139 |
| Max. P | 0.763121 |

| Location | 13,539,516 – 13,539,616 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 89.08 |
| Mean single sequence MFE | -19.47 |
| Consensus MFE | -16.65 |
| Energy contribution | -16.09 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.763121 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
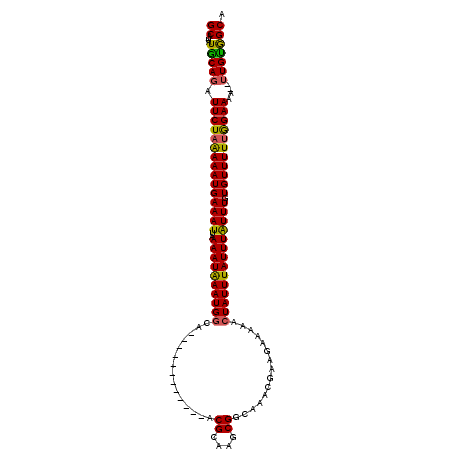
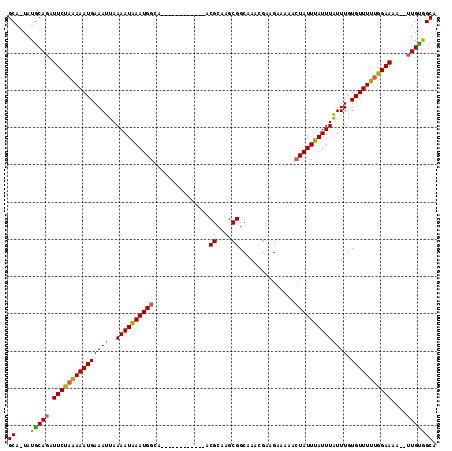
>2R_DroMel_CAF1 13539516 100 - 20766785 GCA-UAUGCAGAUUCUAAAAAUGAAAUUAAAAUAAAUGGCA------------ACGCAAGCGGCAAACGAAGAAAAACUAUUUAUUUAUUUGUGUUUUUGGAAAA--UUGUGGCA ((.-...((((.(((((((((((((((..((((((((((..------------.(....)((.....))........)))))))))))))).)))))))))))..--)))).)). ( -18.70) >DroSec_CAF1 87190 100 - 1 GCA-UAUGCAGAUUCUAAAAAUGAAAUUAAAAUAAAUGGCA------------ACGCAAGCGGCAAACGAAGAAAAACUAUUUAUUUAUUUGUGUUUUUAGAAAA--UUGUGGCA ((.-...((((.(((((((((((((((..((((((((((..------------.(....)((.....))........)))))))))))))).)))))))))))..--)))).)). ( -19.00) >DroSim_CAF1 91123 100 - 1 GCA-UAUGCAGAUUCUAAAAAUGAAAUUAAAAUAAAUGGCA------------ACGCAAGCGGCAAACGAAGAAAAACUAUUUAUUUAUUUGUGUUUUUGGAAAA--UUGUGGCA ((.-...((((.(((((((((((((((..((((((((((..------------.(....)((.....))........)))))))))))))).)))))))))))..--)))).)). ( -18.70) >DroEre_CAF1 88345 100 - 1 GCA-UAUGCAGAUUCUAAAAAUGAAAUUAAAAUAAAUGGAA------------ACGCAAGCGGCAAAUGAAGAAAAACUAUUUAUUUAUUUGUGUUUUUGGAAUA--UUGCGGCA ((.-...(((((((((((((((...............(...------------.).......(((((((((.(((.....))).)))))))))))))))))))).--)))).)). ( -22.40) >DroYak_CAF1 92029 102 - 1 GCAAUAUACAGAUUCUAGAAAUGAAAUUAAAAUAAAUGGAA------------ACGCA-GCGGCAAAUGAAGAAAAACUAUUUAUUUAUUUGUGUUUUUGGAAAAUUUUGUGGCA ((....((((((((((((((((...............(...------------.)...-...(((((((((.(((.....))).)))))))))))))))))))...)))))))). ( -18.90) >DroAna_CAF1 89228 108 - 1 GCA-UAUGCAGAUUCUAAAAAUGAAAUUAAAAUGAAUGGCAAACGCAACGCAAACGCAAGCGGCAAA---AGAAAAAGUAUUUAUUUGUUUGUGUUUUCAGAAAA---UGUAGCU ((.-..((((..((((.((((((......(((((((((((....))...((...(....)..))...---........))))))))).....)))))).))))..---)))))). ( -19.10) >consensus GCA_UAUGCAGAUUCUAAAAAUGAAAUUAAAAUAAAUGGCA____________ACGCAAGCGGCAAACGAAGAAAAACUAUUUAUUUAUUUGUGUUUUUGGAAAA__UUGUGGCA ((....(((((.(((((((((((((((..((((((((((...............((....))...............)))))))))))))).)))))))))))....))))))). (-16.65 = -16.09 + -0.55)

| Location | 13,539,554 – 13,539,655 |
|---|---|
| Length | 101 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 92.83 |
| Mean single sequence MFE | -19.10 |
| Consensus MFE | -18.19 |
| Energy contribution | -17.97 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.626201 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
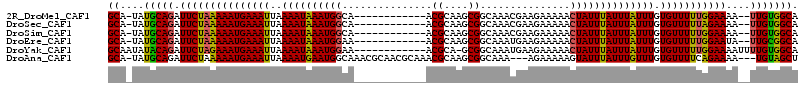
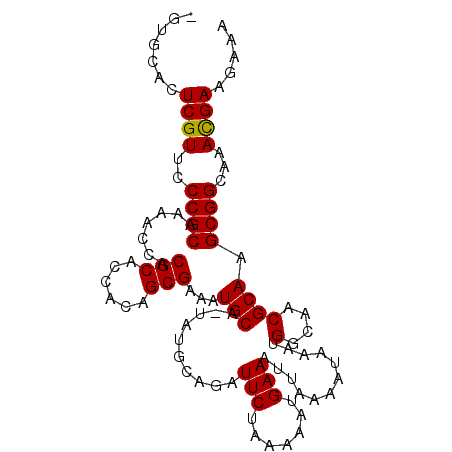
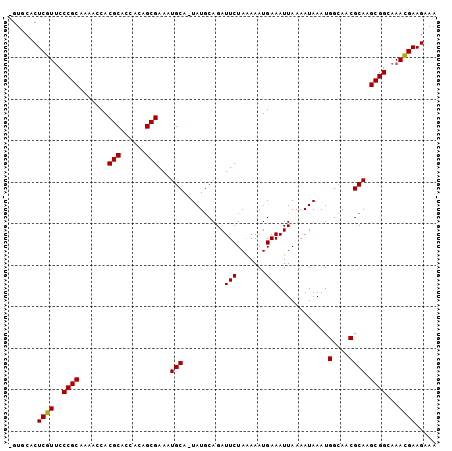
>2R_DroMel_CAF1 13539554 101 - 20766785 -GUGCACUCGUUCCCGCAAAACCACGCACCACAGCGAAAUGCA-UAUGCAGAUUCUAAAAAUGAAAUUAAAAUAAAUGGCAACGCAAGCGGCAAACGAAGAAA -......(((((.((((.......(((......))).......-..(((...(((.......)))............(....)))).))))..)))))..... ( -19.70) >DroSim_CAF1 91161 101 - 1 -GUGCACUCGUCCCCGCAAAACCACGCACCGCAGCGAAAUGCA-UAUGCAGAUUCUAAAAAUGAAAUUAAAAUAAAUGGCAACGCAAGCGGCAAACGAAGAAA -......((((..((((.......(((......))).......-..(((...(((.......)))............(....)))).))))...))))..... ( -18.80) >DroYak_CAF1 92069 102 - 1 CGUCCACUCGUUCCCGCAAAACCACGCACCACAGCGAACUGCAAUAUACAGAUUCUAGAAAUGAAAUUAAAAUAAAUGGAAACGCA-GCGGCAAAUGAAGAAA .......(((((.((((.......(((......)))...(((........((((.((....)).)))).........(....))))-))))..)))))..... ( -18.80) >consensus _GUGCACUCGUUCCCGCAAAACCACGCACCACAGCGAAAUGCA_UAUGCAGAUUCUAAAAAUGAAAUUAAAAUAAAUGGCAACGCAAGCGGCAAACGAAGAAA .......((((..((((.......(((......)))...(((..........(((.......)))............(....)))).))))...))))..... (-18.19 = -17.97 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:23 2006