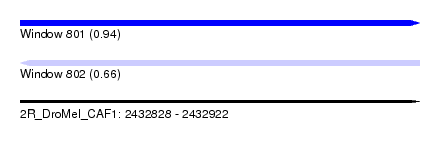

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 2,432,828 – 2,432,922 |
| Length | 94 |
| Max. P | 0.944693 |

| Location | 2,432,828 – 2,432,922 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 78.90 |
| Mean single sequence MFE | -29.66 |
| Consensus MFE | -16.92 |
| Energy contribution | -18.52 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.57 |
| SVM decision value | 1.35 |
| SVM RNA-class probability | 0.944693 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
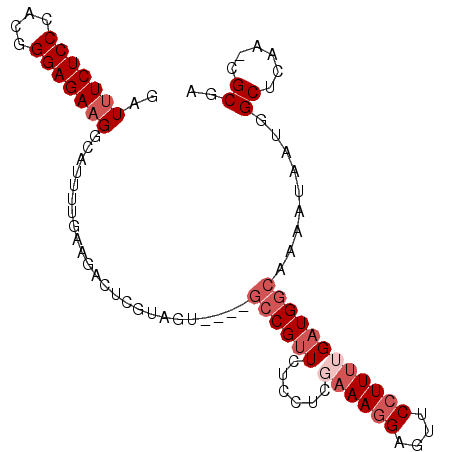
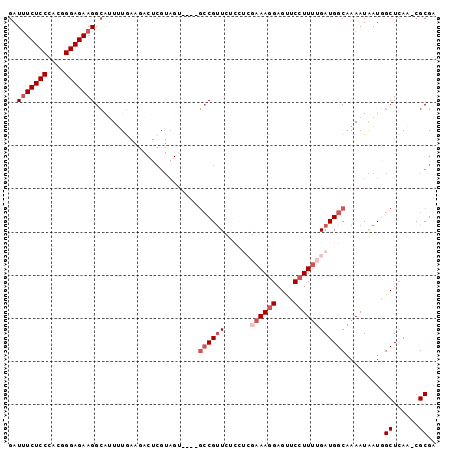
>2R_DroMel_CAF1 2432828 94 + 20766785 GAUGUCUCCCACGGGAGAAGGCAUUUUGAAGACUCGUAGU----GCCGUUCUCCUCGAAAGGAGUUCCUUUUGAUGGCAAAAUAAUGGCUCAA-CGCGA ..((((.......((((((((((((.(((....))).)))----))).))))))((((((((....)))))))).))))........((....-.)).. ( -31.60) >DroSec_CAF1 88726 93 + 1 GAUUUCUCCCACGGGAGAAGGCAUU-UGAAGACUCGUAGU----GCCGUUCUCCUCGAAAGGAGUUCCUUUUGAUGGCAAAAUAAUAGCUCAA-CGCGA ...((((((....))))))((((((-(((....))).)))----))).....((((((((((....)))))))).))..........((....-.)).. ( -29.80) >DroSim_CAF1 46203 94 + 1 GAUUUCUCCCACGGGAGAAGGCAUUUUGAAGACUCGUAGU----GCCGUUCUCCUCGAAAGGAGUUCCUUUUGAUGGCAAAAUAAUGGCUAAA-CGCGA .........(.((((((((((((((.(((....))).)))----))).))))))((((((((....))))))))((((.........))))..-)).). ( -30.50) >DroEre_CAF1 44554 77 + 1 GAUUUCUCCCAUGGGAGAAGGCGUUUUG--------------GAGCCGUU-------CAAGGAGCUC-UUCUGAUGGCAAAAUGAUGGCUCAGUCGCGU ..(((((((....)))))))(((.....--------------(((((((.-------((....(((.-.......)))....)))))))))...))).. ( -27.20) >DroYak_CAF1 45059 92 + 1 GAUUUCUCCCAUGGGAGAAGGGAGUUUAAAGACUCGUAGAAGGACGCGAU-------AAAGGAGUUCCUUUUGAUGGCAAAAUAAUAGCUCAGUCGCGU ..(((((((....))))))).(((((....)))))........(((((((-------....(((((..(((((....)))))....))))).))))))) ( -29.20) >consensus GAUUUCUCCCACGGGAGAAGGCAUUUUGAAGACUCGUAGU____GCCGUUCUCCUCGAAAGGAGUUCCUUUUGAUGGCAAAAUAAUGGCUCAA_CGCGA ..(((((((....)))))))........................((((((......((((((....)))))))))))).........((......)).. (-16.92 = -18.52 + 1.60)

| Location | 2,432,828 – 2,432,922 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 78.90 |
| Mean single sequence MFE | -24.84 |
| Consensus MFE | -11.75 |
| Energy contribution | -12.23 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.47 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.660775 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
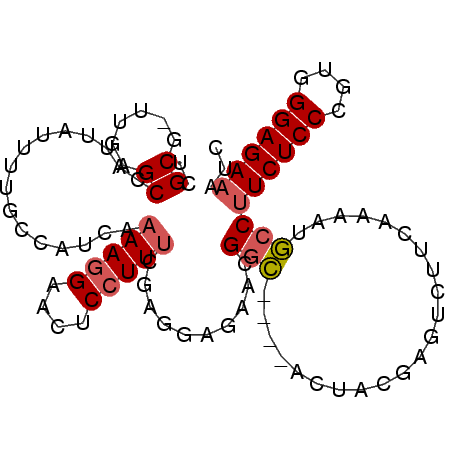
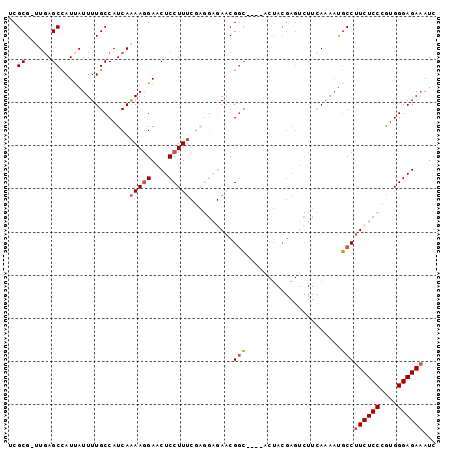
>2R_DroMel_CAF1 2432828 94 - 20766785 UCGCG-UUGAGCCAUUAUUUUGCCAUCAAAAGGAACUCCUUUCGAGGAGAACGGC----ACUACGAGUCUUCAAAAUGCCUUCUCCCGUGGGAGACAUC ((...-((((..((......))...))))...)).(((((..((.((((((.(((----(....(......)....)))))))))))).)))))..... ( -25.70) >DroSec_CAF1 88726 93 - 1 UCGCG-UUGAGCUAUUAUUUUGCCAUCAAAAGGAACUCCUUUCGAGGAGAACGGC----ACUACGAGUCUUCA-AAUGCCUUCUCCCGUGGGAGAAAUC ..(((-(((((...(..(..((((.((.....)).(((((....)))))...)))----)..)..)...))).-))))).((((((....))))))... ( -26.10) >DroSim_CAF1 46203 94 - 1 UCGCG-UUUAGCCAUUAUUUUGCCAUCAAAAGGAACUCCUUUCGAGGAGAACGGC----ACUACGAGUCUUCAAAAUGCCUUCUCCCGUGGGAGAAAUC ..(((-(((................((.(((((....))))).))(((((.((..----....))..))))).)))))).((((((....))))))... ( -25.60) >DroEre_CAF1 44554 77 - 1 ACGCGACUGAGCCAUCAUUUUGCCAUCAGAA-GAGCUCCUUG-------AACGGCUC--------------CAAAACGCCUUCUCCCAUGGGAGAAAUC ..(((.((((..((......))...))))..-(((((.....-------...)))))--------------.....))).((((((....))))))... ( -22.30) >DroYak_CAF1 45059 92 - 1 ACGCGACUGAGCUAUUAUUUUGCCAUCAAAAGGAACUCCUUU-------AUCGCGUCCUUCUACGAGUCUUUAAACUCCCUUCUCCCAUGGGAGAAAUC ((((((.((.((.........))))...(((((....)))))-------.))))))........((((......))))..((((((....))))))... ( -24.50) >consensus UCGCG_UUGAGCCAUUAUUUUGCCAUCAAAAGGAACUCCUUUCGAGGAGAACGGC____ACUACGAGUCUUCAAAAUGCCUUCUCCCGUGGGAGAAAUC ..((......))................(((((....)))))..........(((......................)))((((((....))))))... (-11.75 = -12.23 + 0.48)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:37:54 2006